Abstract
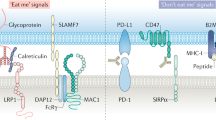
Exciting progress in the field of cancer immunotherapy has renewed the urgency of the need for basic studies of immunoregulation in both adaptive cell lineages and innate cell lineages. Here we found a central role for major histocompatibility complex (MHC) class I in controlling the phagocytic function of macrophages. Our results demonstrated that expression of the common MHC class I component β2-microglobulin (β2M) by cancer cells directly protected them from phagocytosis. We further showed that this protection was mediated by the inhibitory receptor LILRB1, whose expression was upregulated on the surface of macrophages, including tumor-associated macrophages. Disruption of either MHC class I or LILRB1 potentiated phagocytosis of tumor cells both in vitro and in vivo, which defines the MHC class I–LILRB1 signaling axis as an important regulator of the effector function of innate immune cells, a potential biomarker for therapeutic response to agents directed against the signal-regulatory protein CD47 and a potential target of anti-cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Weiner, L. M., Dhodapkar, M. V. & Ferrone, S. Monoclonal antibodies for cancer immunotherapy. Lancet 373, 1033–1040 (2009).
Weiskopf, K. & Weissman, I. L. Macrophages are critical effectors of antibody therapies for cancer. MAbs 7, 303–310 (2015).
Jaiswal, S. et al. CD47 is upregulated on circulating hematopoietic stem cells and leukemia cells to avoid phagocytosis. Cell 138, 271–285 (2009).
Oldenborg, P. A. et al. Role of CD47 as a marker of self on red blood cells. Science 288, 2051–2054 (2000).
Majeti, R. et al. CD47 is an adverse prognostic factor and therapeutic antibody target on human acute myeloid leukemia stem cells. Cell 138, 286–299 (2009).
Chao, M. P. et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-Hodgkin lymphoma. Cell 142, 699–713 (2010).
Willingham, S. et al. The CD47-SIRPa interaction is a therapeutic target for human solid tumors. Proc. Natl. Acad. Sci. USA 109, 6662–6667 (2012).
Weiskopf, K. et al. Engineered SIRPα variants as immunotherapeutic adjuvants to anticancer antibodies. Science 341, 88–91 (2013).
Weiskopf, K. Cancer immunotherapy targeting the CD47/SIRPα axis. Eur. J. Cancer 76, 100–109 (2017).
Weiskopf, K. et al. CD47-blocking immunotherapies stimulate macrophage-mediated destruction of small-cell lung cancer. J. Clin. Invest 126, 2610–2620 (2016).
Liu, J. et al. Pre-clinical development of a humanized anti-CD47 antibody with anti-cancer therapeutic potential. PLoS One 10, e0137345 (2015).
Krampitz, G. W. et al. Identification of tumorigenic cells and therapeutic targets in pancreatic neuroendocrine tumors. Proc. Natl. Acad. Sci. USA 113, 4464–4469 (2016).
Bjorkman, P. J. et al. Structure of the human class I histocompatibility antigen, HLA-A2. Nature 329, 506–512 (1987).
Klein, J. & Sato, A. The HLA system. First of two parts. N. Engl. J. Med. 343, 702–709 (2000).
Klein, J. & Sato, A. The HLA system. Second of two parts. N. Engl. J. Med. 343, 782–786 (2000).
Bicknell, D. C., Rowan, A. & Bodmer, W. F. β2-microglobulin gene mutations: a study of established colorectal cell lines and fresh tumors. Proc. Natl. Acad. Sci. USA 91, 4751–4755 (1994).
Borges, L., Hsu, M. L., Fanger, N., Kubin, M. & Cosman, D. A family of human lymphoid and myeloid Ig-like receptors, some of which bind to MHC class I molecules. J. Immunol. 159, 5192–5196 (1997).
Barnstable, C. J. et al. Production of monoclonal antibodies to group A erythrocytes, HLA and other human cell surface antigens-new tools for genetic analysis. Cell 14, 9–20 (1978).
Fanger, N. A. et al. The MHC class I binding proteins LIR-1 and LIR-2 inhibit Fc receptor-mediated signaling in monocytes. Eur. J. Immunol. 28, 3423–3434 (1998).
Willcox, B. E., Thomas, L. M. & Bjorkman, P. J. Crystal structure of HLA-A2 bound to LIR-1, a host and viral major histocompatibility complex receptor. Nat. Immunol. 4, 913–919 (2003).
Cheng, H. et al. Crystal structure of leukocyte Ig-like receptor LILRB4 (ILT3/LIR-5/CD85k): a myeloid inhibitory receptor involved in immune tolerance. J. Biol. Chem. 286, 18013–18025 (2011).
Pulford, K., Micklem, K., Thomas, J., Jones, M. & Mason, D. Y. A 72-kD B cell-associated surface glycoprotein expressed at high levels in hairy cell leukaemia and plasma cell neoplasms. Clin. Exp. Immunol 85, 429–435 (1991).
Allan, D. S. J. et al. Tetrameric complexes of human histocompatibility leukocyte antigen (HLA)-G bind to peripheral blood myelomonocytic cells. J. Exp. Med. 189, 1149–1156 (1999).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res 42, e168 (2014).
Garcia, K. C. et al. An alphabeta T cell receptor structure at 2.5 A and its orientation in the TCR-MHC complex. Science 274, 209–219 (1996).
Garboczi, D. N. et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Boyington, J. C., Motyka, S. A., Schuck, P., Brooks, A. G. & Sun, P. D. Crystal structure of an NK cell immunoglobulin-like receptor in complex with its class I MHC ligand. Nature 405, 537–543 (2000).
Sanderson, A. R. HLA “help” for human B2-microglobulin across species barriers. Nature 269, 414–417 (1977).
Shultz, L. D. et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2Rγ null mice engrafted with mobilized human hemopoietic stem cells. J. Immunol. 174, 6477–6489 (2005).
Mantovani, A. et al. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 25, 677–686 (2004).
Cosman, D. et al. A novel immunoglobulin superfamily receptor for cellular and viral MHC class I molecules. Immunity 7, 273–282 (1997).
Prod’homme, V. et al. The human cytomegalovirus MHC class I homolog UL18 inhibits LIR-1+ but activates LIR-1– NK cells. J. Immunol. 178, 4473–4481 (2007).
Baía, D. et al. Interaction of the LILRB1 inhibitory receptor with HLA class Ia dimers. Eur. J. Immunol. 46, 1681–1690 (2016).
Kim, T. et al. Human LilrB2 is a β-amyloid receptor and its murine homolog PirB regulates synaptic plasticity in an Alzheimer’s model. Science 341, 1399–1404 (2013).
Campbell, K. S. & Purdy, A. K. Structure/function of human killer cell immunoglobulin-like receptors: lessons from polymorphisms, evolution, crystal structures and mutations. Immunology 132, 315–325 (2011).
Lv, Z. et al. Loss of cell surface CD47 clustering formation and binding avidity to SIRPα facilitate apoptotic cell clearance by macrophages. J. Immunol. 195, 661–671 (2015).
Garcia-Lora, A., Algarra, I. & Garrido, F. MHC class I antigens, immune surveillance, and tumor immune escape. J. Cell. Physiol. 195, 346–355 (2003).
Challa-Malladi, M. et al. Combined genetic inactivation of β2-microglobulin and CD58 reveals frequent escape from immune recognition in diffuse large B cell lymphoma. Cancer Cell 20, 728–740 (2011).
Zijlstra, M. et al. Beta 2-microglobulin deficient mice lack CD4–8+ cytolytic T cells. Nature 344, 742–746 (1990).
Koller, B. H., Marrack, P., Kappler, J. W. & Smithies, O. Normal development of mice deficient in β2M, MHC class I proteins, and CD8+ T cells. Science 248, 1227–1230 (1990).
Tseng, D. et al. Anti-CD47 antibody-mediated phagocytosis of cancer by macrophages primes an effective antitumor T-cell response. Proc. Natl. Acad. Sci. USA 110, 11103–11108 (2013).
Acknowledgements
We thank the members of the Weissman laboratory and the Stanford Stem Cell Center community for discussion and assistance; J.P. Volkmer, R. Majeti, K. Loh, N. Fernhoff, M. McCracken and J. Zee for input and advice; T. Storm, T. Naik, A. McCarty, F. Khameneh, J. Ho and P. Lovelace for logistical support; and G. Krampitz (Stanford University) for the APL1 cell line. Supported by the D.K. Ludwig Fund for Cancer Research (NIH R01CA086017 and NIHG R01GM100315 to I.L.W.), the Cancer Research Institute (Irvington Fellowship to R.L.M.), the Human Frontier Science Program Organization (postdoctoral fellowship to B.R.), the NIH (hematology training grant T32 HL120824-03 to B.R.), the University of Wisconsin Medical Scientists Training Program (GM008692 to L.J.B.), the National Research Service Award 1F30DK108561, the Paul and Daisy Soros Fellowship for New Americans (to J.M.T.) and the Stanford Medical Scientist Training Program (NIH GM07365 to A.A.B., K.W., A.M.R., J.M.T., B.M.G. and J.Y.C.).
Author information
Authors and Affiliations
Contributions
A.A.B., K.W., I.L.W. and R.L.M. wrote the manuscript; A.A.B., K.W., A.M.R., I.L.W. and R.L.M. conceived of and designed all experiments; K.W. and R.L.M. performed LegendScreen analysis of tumor cell lines; K.S.K., S.R.G., B.M.G. and R.Z.C. assisted with mouse experiments; B.R. and Y.Y.Y. performed receptor-Fc staining experiments; M.M. assisted with tumor measurements and injections and provided sample preparation for primary human macrophages; N.G.R. generated human macrophages and tested the in vitro growth kinetics of DLD1 sublines; J.M.T. performed histology and immunofluorescence staining; K.M.M. generated NSG macrophages; K.M.M., P.Y.H. and R.L.M. tested CD47 expression in tumor cell lines; J.Y.C. assisted with generating knockout cell lines; L.J.B. assisted with statistical analysis; A.M.R. generated Fab fragments of W6/32; and I.L.W. supervised the research and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.W., A.M.R., I.L.W. and R.L.M. and are co-inventors on patent application PCT/US2015/057233, which is related to this work, and own stock of FortySeven, which is pursing clinical approval of antibody Hu5F9-G4, directed against human CD47.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1. Validation of flow cytometry–based in vitro phagocytosis assay
(a) Representative flow cytometry plots of in vitro phagocytosis assay. Macrophages are defined as CD45+ events, and tumor cells as GFP+ events. CD45+, GFP+ events correspond to macrophages that have phagocytosed target cells. (b) Representative images from an automated microscopy-based approach for quantifying in vitro phagocytosis. Using a custom program on the GE Cytell, we defined human macrophages based on staining with anti-CD45-PE antibody (yellow), and tumor cells based on GFP expression (green). Phagocytic events are defined based on overlap of these two signals (right panel, white arrowheads). (c) Comparison of FACS-based quantification (left) or automated microscopy-based quantification (right) of DLD1 phagocytosis under conditions of PBS treatment, treatment with the anti-CD47 antibody Hu5F9-G4, or treatment with the anti-EGFR antibody Cetuximab.
Supplementary Figure 2. Analysis of CD47 and MHC class I component expression in cell lines
(a) Log-transformed scatterplot of normalized phagocytic efficiency upon treatment with the anti-CD47 antibody Hu5F9-G4 (Y axis) plotted against surface expression of CD47, as measured by FACS analysis with the anti-human mouse monoclonal antibody B6H12 (X axis). There is no significant relationship between these two parameters, R2=0.094, p=0.217. (b) Histogram plots of HLA-A, B, C (top) and β2M expression (bottom) for two colon cancer cell lines: DLD1 (red) and HCT116 (magenta); two small cell lung cancer lines, NCI-H82 (blue), and NCI-H196 (purple); and a pancreatic neuroendocrine tumor APL1 (green), as measured by the BioLegend LegendScreen FACS-based antibody array system. Isotype control stain is indicated in blue, while specific stain is indicated in red. (c) Log-scale scatterplot of phagocytic efficiency in a panel of cell lines upon treatment with the anti-CD47 antibody Hu5F9-G4 (Y axis), plotted against surface expression of HLA-A, B, C as measured by FACS analysis with a pan-HLA-A, B, C binding antibody (X axis). The data is a subset of that shown in Figure 1b, in order to highlight the cell lines used for the experiment shown in Figure 2g.
Supplementary Figure 3. Analysis of the expression and function of LILRB1 and LILRB2 in macrophages
(a) Flow cytometry-based measurement of the percentage of LILRB1 (blue) or LILRB2 (red) positive, non-immune cells from six samples of primary ascites fluid macrophages, and four samples of primary human colorectal carcinoma samples, as determined by fluorescence minus one controls. n.s. not significant, 2-way ANOVA with multiple comparisons correction. (b) Flow cytometry-based measurement of the percentage of LILRB1 (blue) or LILRB2 (red) positive tumor associated macrophages from four independent primary human colorectal carcinoma samples, as determined by fluorescence minus one controls. n.s. not significant, 2-way ANOVA with multiple comparisons correction. (c) Representative histogram plots from FACS analysis of LILRB1 and LILRB2 expression on primary human CD14+ peripheral blood monocytes (left), and day 7 ex vivo cultured macrophages derived from the same donor (right). Specific staining is indicated in blue; IgG control is indicated in red. (d) LILRB1 and LILRB2 expression levels, as measured by mean fluorescence intensity (MFI, left panel) or percent positive cells (right panel), in 4 pairs of primary monocytes (red) and ex vivo cultured macrophages from the same donors (blue). n.s., not significant. *** p < 0.001, 2-way ANOVA with multiple comparisons correction. (e) Flow cytometry-based measurement of phagocytosis by donor-derived macrophages of MHC I– DLD1 (black) and MHC I+ DLD1 (red). Values are normalized to the highest level of phagocytosis observed in a given experimental replicate. Error bars represent s.d. of assays performed with n = 8 biological replicates. * p < 0.01, ** p < 0.001, 2-way ANOVA with multiple comparisons correction.
Supplementary Figure 4. Mouse β2M does not efficiently form stable MHC class I complexes with human HLA α-chains
FACS histogram of parental DLD1 cells (black), DLD1 cells reconstituted with a human β2M transgene (red), or with a mouse β2M transgene (blue). While fully human β2M expression facilitates robust surface MHC class I expression, as detected by the pan-HLA antibody W6/32, expression of mouse β2M facilitates only low levels of surface MHC class I.
Supplementary Figure 5. Growth kinetics and immunofluorescence of engineered cell lines
(a) Growth curve of MHC I– DLD1 (pink) and chimeric MHC I+ DLD1 (blue), as measured by cell counting at 1, 2, and 4 days. Error bars represent are s.d. of n = 3 replicates. (b) Flow cytometry analysis of MHC class I expression in a 50-50 in vitro co-culture of MHC I– APL1 and MHC I- APL1. Error bars represent s.d. of n = 3 replicates. (c) Flow cytometry-based quantification of the percentage of fully human MHC class I expression in a 50-50 in vitro co-culture of MHC I– APL1 and MHC I- APL1 cells pre-engraftment and at 28 days post engraftment into NSG mice. Error bars are s.d. of n = 5 mice. (d) Fluorescence microscopy of sections taken from mixed MHC I+ and MHC I– APL1 tumors injected subcutaneously into the flanks of NSG mice. Chimeric MHC class I+ cells are marked by GFP expression, while MHC I– cells are marked by RFP expression. Mouse macrophages are visualized by F4/80 staining (blue). A high degree of macrophage infiltration into the tumor is evident, and macrophages can be observed in the act of phagocytosis (white arrowhead). Scale bar represents 200 μm.
Supplementary Figure 6. Dynamics of tumor-infiltrating macrophages
(a) Tumor volume of B16-F10 variants 15 days post-engraftment into immunocompetent C57Bl/6 wild type mice (black) or CCR2-/- mice (red). n.s. not significant. * p < 0.05, *** p < 0.001, Student’s t test. (b) FACS-based measurement of LILRB1 expression in primary human macrophages electroporated with either off-target sgRNA (black) or LILRB1-targeting sgRNA, as determined by MFI. MFI values are normalized to the highest value for each experimental replicate. Error bars represent the standard deviation of assays performed with n = 2 macrophage donors. (c) Frequency of allelic deletion of LILRB1 in primary human macrophages, as measured by TIDE analysis software, n = 2 macrophage donors. (d) Percentage of total macrophages with an M1-like phenotype in macrophages modified with off-target sgRNA (black) or LILRB1-targeting sgRNA (red) in n = 2 macrophage donors. (e) Gating strategy to define infiltrating immune cells within dissociated B16-F10 mouse tumors.
Supplementary information
Rights and permissions
About this article
Cite this article
Barkal, A.A., Weiskopf, K., Kao, K.S. et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat Immunol 19, 76–84 (2018). https://doi.org/10.1038/s41590-017-0004-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0004-z
This article is cited by
-
Nanoparticles in tumor microenvironment remodeling and cancer immunotherapy
Journal of Hematology & Oncology (2024)
-
The Hippo-YAP signaling pathway drives CD24-mediated immune evasion in esophageal squamous cell carcinoma via macrophage phagocytosis
Oncogene (2024)
-
Sirpα on tumor-associated myeloid cells restrains antitumor immunity in colorectal cancer independent of its interaction with CD47
Nature Cancer (2024)
-
Fcγ receptors and immunomodulatory antibodies in cancer
Nature Reviews Cancer (2024)
-
Mechanism of Efferocytosis in Determining Ischaemic Stroke Resolution—Diving into Microglia/Macrophage Functions and Therapeutic Modality
Molecular Neurobiology (2024)