Abstract
Prolonged activation of interferon–STAT1 signaling is closely related to inflammatory autoimmune disorders, and therefore the identification of negative regulators of these pathways is important. Through high-content screening of 115 mouse RING-domain E3 ligases, we identified the E3 ubiquitin ligase RNF2 as a potent inhibitor of interferon-dependent antiviral responses. RNF2 deficiency substantially enhanced interferon-stimulated gene (ISG) expression and antiviral responses. Mechanistically, nuclear RNF2 directly bound to STAT1 after interferon stimulation and increased K33-linked polyubiquitination of the DNA-binding domain of STAT1 at position K379, in addition to promoting the disassociation of STAT1/STAT2 from DNA and consequently suppressing ISG transcription. Our study provides insight into the regulation of interferon-dependent responses via a previously unrecognized post-translational modification of STAT1 in the nucleus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Stark, G. R. & Darnell, J. E. Jr. The JAK-STAT pathway at twenty. Immunity 36, 503–514 (2012).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15, 87–103 (2015).
Wilson, E. B. et al. Blockade of chronic type I interferon signaling to control persistent LCMV infection. Science 340, 202–207 (2013).
Teijaro, J. R. et al. Persistent LCMV infection is controlled by blockade of type I interferon signaling. Science 340, 207–211 (2013).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Porritt, R. A. & Hertzog, P. J. Dynamic control of type I IFN signalling by an integrated network of negative regulators. Trends Immunol. 36, 150–160 (2015).
Bancerek, J. et al. CDK8 kinase phosphorylates transcription factor STAT1 to selectively regulate the interferon response. Immunity 38, 250–262 (2013).
Krämer, O. H. et al. A phosphorylation-acetylation switch regulates STAT1 signaling. Genes Dev. 23, 223–235 (2009).
Tanaka, T., Soriano, M. A. & Grusby, M. J. SLIM is a nuclear ubiquitin E3 ligase that negatively regulates STAT signaling. Immunity 22, 729–736 (2005).
Chen, K. et al. Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell 170, 492–506 (2017).
Jiang, X. & Chen, Z. J. The role of ubiquitylation in immune defence and pathogen evasion. Nat. Rev. Immunol. 12, 35–48 (2011).
Lawrence, D. W. & Kornbluth, J. E3 ubiquitin ligase NKLAM ubiquitinates STAT1 and positively regulates STAT1-mediated transcriptional activity. Cell. Signal. 28, 1833–1841 (2016).
Yuan, C., Qi, J., Zhao, X. & Gao, C. Smurf1 protein negatively regulates interferon-γ signaling through promoting STAT1 protein ubiquitination and degradation. J. Biol. Chem. 287, 17006–17015 (2012).
Versteeg, G. A. et al. The E3-ligase TRIM family of proteins regulates signaling pathways triggered by innate immune pattern-recognition receptors. Immunity 38, 384–398 (2013).
Simon, J. A. & Kingston, R. E. Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol. Cell 49, 808–824 (2013).
Kondo, T. et al. Polycomb potentiates Meis2 activation in midbrain by mediating interaction of the promoter with a tissue-specific enhancer. Dev. Cell 28, 94–101 (2014).
Rai, K. et al. Dual roles of RNF2 in melanoma progression. Cancer Discov 5, 1314–1327 (2015).
Arimoto, K. et al. Negative regulation of the RIG-I signaling by the ubiquitin ligase RNF125. Proc. Natl. Acad. Sci. USA 104, 7500–7505 (2007).
Wang, W. et al. RNF122 suppresses antiviral type I interferon production by targeting RIG-I CARDs to mediate RIG-I degradation. Proc. Natl. Acad. Sci. USA 113, 9581–9586 (2016).
Voncken, J. W. et al. Rnf2 (Ring1b) deficiency causes gastrulation arrest and cell cycle inhibition. Proc. Natl. Acad. Sci. USA 100, 2468–2473 (2003).
Suzuki, M. et al. Involvement of the Polycomb-group gene Ring1B in the specification of the anterior-posterior axis in mice. Development 129, 4171–4183 (2002).
McBride, K. M., Banninger, G., McDonald, C. & Reich, N. C. Regulated nuclear import of the STAT1 transcription factor by direct binding of importin-α. EMBO J. 21, 1754–1763 (2002).
Elderkin, S. et al. A phosphorylated form of Mel-18 targets the Ring1B histone H2A ubiquitin ligase to chromatin. Mol. Cell 28, 107–120 (2007).
Meyer, T., Marg, A., Lemke, P., Wiesner, B. & Vinkemeier, U. DNA binding controls inactivation and nuclear accumulation of the transcription factor Stat1. Genes Dev. 17, 1992–2005 (2003).
Banninger, G. & Reich, N. C. STAT2 nuclear trafficking. J. Biol. Chem. 279, 39199–39206 (2004).
Endoh, M. et al. Histone H2A mono-ubiquitination is a crucial step to mediate PRC1-dependent repression of developmental genes to maintain ES cell identity. PLoS Genet. 8, e1002774 (2012).
Eskeland, R. et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 38, 452–464 (2010).
Zhou, W. et al. Histone H2A monoubiquitination represses transcription by inhibiting RNA polymerase II transcriptional elongation. Mol. Cell 29, 69–80 (2008).
Wauman, J., De Ceuninck, L., Vanderroost, N., Lievens, S. & Tavernier, J. RNF41 (Nrdp1) controls type 1 cytokine receptor degradation and ectodomain shedding. J. Cell Sci. 124, 921–932 (2011).
Grinde, B., Hetland, G. & Johnson, E. Effects on gene expression and viral load of a medicinal extract from Agaricus blazei in patients with chronic hepatitis C infection. Int. Immunopharmacol. 6, 1311–1314 (2006).
Luesch, H. et al. A functional genomics approach to the mode of action of apratoxin A. Nat. Chem. Biol. 2, 158–167 (2006).
Toubiana, J. et al. Heterozygous STAT1 gain-of-function mutations underlie an unexpectedly broad clinical phenotype. Blood 127, 3154–3164 (2016).
Mizoguchi, Y. et al. Simple diagnosis of STAT1 gain-of-function alleles in patients with chronic mucocutaneous candidiasis. J. Leukoc. Biol. 95, 667–676 (2014).
Takezaki, S. et al. Chronic mucocutaneous candidiasis caused by a gain-of-function mutation in the STAT1 DNA-binding domain. J. Immunol. 189, 1521–1526 (2012).
Rosas-Murrieta, N. H., Herrera-Camacho, I., Palma-Ocampo, H., Santos-López, G. & Reyes-Leyva, J. Interaction of mumps virus V protein variants with STAT1-STAT2 heterodimer: experimental and theoretical studies. Virol. J. 7, 263 (2010).
Huang, H. et al. K33-linked polyubiquitination of T cell receptor-ζ regulates proteolysis-independent T cell signaling. Immunity 33, 60–70 (2010).
Yang, M. et al. K33-linked polyubiquitination of Zap70 by Nrdp1 controls CD8+ T cell activation. Nat. Immunol. 16, 1253–1262 (2015).
Yuan, W. C. et al. K33-linked polyubiquitination of Coronin 7 by Cul3-KLHL20 ubiquitin E3 ligase regulates protein trafficking. Mol. Cell 54, 586–600 (2014).
Meng, J. et al. Rb selectively inhibits innate IFN-b production by enhancing deacetylation of IFN-β promoter through HDAC1 and HDAC8. J. Autoimmun. 73, 42–53 (2016).
Xia, M. et al. Histone methyltransferase Ash1l suppresses interleukin-6 production and inflammatory autoimmune diseases by inducing the ubiquitin-editing enzyme A20. Immunity 39, 470–481 (2013).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (grants 81788104 and 31390431 to X.C.), the National Key Basic Research Program of China (grant 2013CB530503 to X.C.; grant 2013CB944903 to M.J.), the National 135 Mega Program of China (grant 2017ZX10102032-001 to X.C.) and the CAMS Innovation Fund for Medical Sciences (grant 2016-12M-1-003 to X.C.).
Author information
Authors and Affiliations
Contributions
S.L., W.W., W.L., X.S., Z.M., S.Z., L.L. and Y.L. performed the experiments; S.L., M.J. and X.C. analyzed data and wrote the manuscript; and X.C. was responsible for research supervision, coordination and strategy.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 RNF2 inhibits type I interferon antiviral responses in vitro
(a) Q-PCR analysis of mRNA expression of Rnf2, Usp18 and Foxo3 in mouse peritoneal macrophages transfected with si-RNF2, si-Usp18, si-Foxo3 or si-Ctrl for 48 h. (b) Q-PCR analysis of VSV replicates, mRNA expression of Ifit1, Mx1 and Isg15 in macrophages transfected with si-RNF2, si-Usp18, si-Foxo3 or si-Ctrl for 48 h, infected with VSV for 12 h. (c) VSV titer detection of cell culture supernatant of macrophages transfected with si-RNF2, si-Usp18, si-Foxo3 or si-Ctrl for 48 h, infected with VSV for indicated hours. (d) Immunoblot analysis of protein level of RNF2 in macrophages infected with VSV for indicated hours. (e) Immunoblot analysis of protein level of RNF2 in Ifnar1 +/+ or Ifnar1 -/- macrophages infected with VSV for 12 h. (f) Schematic illustration of the target region of Rnf2 -/- RAW264.7 cells. (g) CCK-8 (Cell counting kit) assay of cell proliferation of RAW264.7 and Rnf2 -/- RAW264.7 cells. (h) CCK-8 assay of cell proliferation of RAW264.7 and RNF2-overexpressing cells. Data are representative of three independent experiments with n = 3 technical replicates (a-c, g, h), three independent experiments (d, e), each symbol represents an individual technical replicate (a-c) (shown as mean and SEM in a, b, c, g, h), * P < 0.05, ** P < 0.01, *** P < 0.001, two-tailed unpaired Student’s t-test
Supplementary Figure 2 RNF2 does not affect percentages of myeloid cells in mouse spleen
(a) Schematic illustration of the knock-out region of Rnf2 fl/fl Lyz2-cre mice. Immunoblot analysis of protein level of RNF2 in different cells from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice. (b-g) Flow cytometry analysis of the percentages of T cells (b), B cells (c), dentritic cells (DCs) (d), neutrophils (e), F4/80+ CD11b+ macrophages (f), and natural killer (NK) cells (g) in splenocytes from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice. Data are representative of three independent experiments (b-g)
Supplementary Figure 3 RNF2 inhibits IFN-γ-induced ISG transcription
(a) Q-PCR analysis of mRNA expression of Irf1, Irf8 and Fgl2 in peritoneal macrophages from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice stimulated with mouse IFN-γ (500 pg/ml) for 6 h. (b) Dual luciferase reporter assay of GAS activity in L929 cells co-transfected with GAS-luciferase reporter vector and RNF2-expressing vector, stimulated with mouse IFN-γ (1 ng/ml) for 12 h. Data are representative of three independent experiments with n = 3 (a), n = 4 (b) technical replicates, each symbol represents an individual technical replicate (a, b) (shown as mean and SEM in a, b), * P < 0.05, two-tailed unpaired Student’s t-test
Supplementary Figure 4 RNF2 inhibits type I interferon antiviral responses in vivo
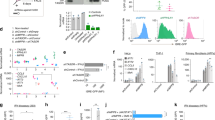
(a) Q-PCR analysis of EMCV replicates in the heart, brain, and pancreas of Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice of 8 weeks age infected with EMCV (1 × 105 PFU per gram body weight) by intraperitoneal injection for 48 h. (c) Immunoblot analysis of protein levels of IFIT1 and ISG15 in the heart and brain from mice described in (b). (d) Q-PCR analysis of mRNA expression of Ifit1, Oas2, and Mx1 in the heart, brain, and pancreas from mice described in (b). (e) Hematoxylin-and eosin staining of heart sections from mice described in (b). Scale bar, 20 μm. (f) Q-PCR analysis of IAV replicates and mRNA expression of Ifit1, Oas2, and Mx1 in the lung of Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice of 8 weeks age infected with IAV (100 PFU) by intranasal delivery for 24 h. (g) Immunoblot analysis of protein levels of IFIT1 and ISG15 in the lung from mice described in (f). (h) Hematoxylin-and eosin staining of lung sections from mice described in (f). Scale bar, 20 μm. Data are representative of three independent experiments with n = 3 (a, c, e) technical replicates, three independent experiments (b, d, f, g), each symbol represents an individual technical replicate (a, c, e) (shown as mean and SEM in a, c, e), * P < 0.05, ** P < 0.01, *** P < 0.001, two-tailed unpaired Student’s t-test
Supplementary Figure 5 RNF2 promotes K33-linked polyubiquitination of STAT1 in the nucleus
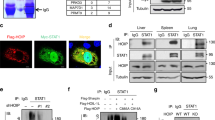
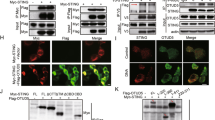
(a) Immunofluorescence analysis of the nuclear localization of RNF2 in RNF2-overexpressing cells, RAW264.7 and Rnf2 -/- RAW264.7 cells. Scale bar, 10 μm. (b) Schematic illustration of the target region of Stat1 -/- L929 cells. Immunoblot analysis of STAT1 in L929 and Stat1 -/- L929 cells stimulated with mouse IFN-β (1 ng/ml) for indicated hours. (c) Immunoprecipitation analysis of the interaction between STAT1 or L407A and RNF2 in Stat1 -/- L929 cells transfected with Flag-STAT1 or Flag-L407A, stimulated with mouse IFN-β (1 ng/ml) for indicated hours. (d) Immunoprecipitation analysis of the polyubiquitination of STAT1 or L407A mediated by RNF2 in Stat1 -/- L929 cells co-transfected with Flag-STAT1 or Flag-L407A, HA-Ub, with or without V5-RNF2, stimulated with mouse IFN-β (1 ng/ml) for 3 h. (e) Immunoprecipitation analysis of polyubiquitination of STAT1 mediated by RNF2 in HEK293T cells co-transfected with Flag-STAT1, HA-Ub mutants, with or without V5-RNF2, stimulated with human IFN-β (1 ng/ml) for 3 h. (f) Immunoprecipitation analysis of polyubiquitination of STAT1 mediated by RNF2 in HEK293T cells co-transfected with Flag-STAT1, HA-Ub mutants, with or without V5-RNF2, stimulated with human IFN-β (1 ng/ml) for 3 h. Data are representative of three independent experiments (a-f)
Supplementary Figure 6 RNF2 promotes dephosphorylation of STAT1
(a) Immunoprecipitation analysis of the association between STAT1 and JAK1 in L929 cells transfected with Flag-STAT1, with or without HA-RNF2, stimulated with mouse IFN-β (1 ng/ml) for 1 h. (b) Immunoprecipitation analysis of the association of STAT1-STAT2 or STAT1-RNF2 in L929 cells co-transfected with HA-Ub-K33O or not, with or without V5-RNF2, stimulated with mouse IFN-β (1 ng/ml) for 3 h. (c) Immunoblot analysis of pY-STAT1 and pY-STAT2 in RAW264.7 and RNF2-overexpressing cells stimulated with mouse IFN-β (500 pg/ml) for indicated times. (d) Immunoblot analysis of pY-STAT1 in macrophages from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice stimulated with mouse IFN-γ (500 pg/ml) for indicated times. (e) Immunoblot analysis of protein level of STAT1 and RNF2 in si-RNA-transfected macrophages from Ifnar1 -/- mice treated with cycloheximide (CHX) (100 μg/ml) for indicated hours 3 h post IFN-β stimulation (500 pg/ml). (f) Immunoblot analysis of pY-STAT1 and pY-STAT2 at indicated times upon addition of staurosporine (100 nM) in RAW264.7 and RNF2-overexpressing cells 30 min post mouse IFN-β stimulation (500 pg/ml). The image intensity of pY-STAT1 and pY-STAT2 was quantified in below. Data are representative of three independent experiments (a-f), pooled from three independent experiments (f) (shown as mean and SEM in f), * P < 0.05, ** P < 0.01, two-tailed unpaired Student’s t-test
Supplementary Figure 7 RNF2 inhibits ISG transcription independently of chromatin structure and monoubiquitination of H2A
(a) ChIP analysis of STAT1 DNA-binding in promoters of Irf1, Irf8 and Fgl2 in macrophages from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice stimulated with mouse IFN-γ (500 pg/ml) for 3 h. (b) ChIP analysis of RNF2 enrichment in promoters of Ifit1 and Mx1 in macrophages stimulated with mouse IFN-β (500 pg/ml) for indicated hours. (c) ChIP analysis of HA-RNF2 enrichment in promoters of Ifit1 and Mx1 in RNF2-overexpressing cells stimulated with mouse IFN-β (500 pg/ml) for indicated hours. (d) ChIP analysis of monoubiquitination of H2A enrichment in promoters of Ifit1, Oas2, Mx1 and Isg15 in peritoneal macrophages from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice stimulated with mouse IFN-β (500 pg/ml) for indicated hours. (e) DNase I sensitivity assay of promoters of Ifit1, Oas2, Mx1 and Isg15 in macrophages from Rnf2 fl/fl and Rnf2 fl/fl Lyz2-cre mice stimulated with mouse IFN-β (500 pg/ml) for indicated hours. (f-g) Immunoprecipitation analysis of the polyubiquitination of STAT1 (f), ChIP analysis of STAT1-STAT2 enrichment in promoters of Ifit1, Oas2, Mx1 (g) in Rnf2 -/- or RNF2-overexpressing RAW264.7 cells transfected with Flag-STAT1, HA-Ub-K33O or HA-Ub-K33R, stimulated with mouse IFN-β (1 ng/ml) for 3 h. Data are representative of three independent experiments with n = 3 technical replicates (a-e, g), three independent experiments (f), each symbol represents an individual technical replicate (a-c, g) (shown as mean and SEM in a, b, c, d, e, g), * P < 0.05, ** P < 0.01, *** P < 0.001, two-tailed unpaired Student’s t-test
Supplementary Figure 8 RNF2 functions via K33-linked polyubiquitination of STAT1 at K379
(a) Dual luciferase reporter assay of ISRE activity and immunoblot analysis of the protein level of STAT1 or STAT1 mutants in Stat1 -/- L929 cells co-transfected with ISRE-luciferase reporter vector, STAT1 or STAT1 mutants, stimulated with mouse IFN-β (1 ng/ml) for 12 h. (b) Dual luciferase reporter assay of ISRE activity in Stat1 -/- L929 cells co-transfected with ISRE-luciferase reporter vector, STAT1 or K379R, with or without RNF2, stimulated with mouse IFN-β (1 ng/ml) for 12 h. (c) Immunoprecipitation analysis of K33-linked polyubiquitination of STAT1 or K379R in HEK293T cells stimulated with human IFN-β (1 ng/ml) for 3 h. (d) Immunoprecipitation analysis of K33-linked polyubiquitination of STAT1 or K379R in HEK293T cells stimulated with human IFN-γ (1 ng/ml) for 3 h. (e) Q-PCR analysis of mRNA expression of Ifit1, Oas2, Mx1 and Isg15 in Stat1 -/- L929 cells transfected with STAT1 or K379R, stimulated with mouse IFN-β (1 ng/ml) for 10 h. (f) Immunoblot analysis of protein level of STAT1 or K379R in Stat1 -/- L929 cells co-transfected with STAT1 or K379R, treated with CHX (100 μg/ml) for indicated hours 3 h post mouse IFN-β stimulation (1 ng/ml). (g) Immunoprecipitation analysis of interaction of STAT1-STAT2 or STAT1-RNF2 in Stat1 -/- L929 cells co-transfected with STAT1 or K379R, stimulated with mouse IFN-β (1 ng/ml) for 3 h. (h) EMSA analysis of ISRE-binding activity of STAT1 or K379R in Stat1 -/- L929 cells co-transfected with STAT1 or K379R, stimulated with mouse IFN-β (1 ng/ml) for 1 h. Data are representative of three independent experiments with n = 3 technical replicates (a, b, e), three independent experiments (c, d, f-h), each symbol represents an individual technical replicate (a, b, e) (shown as mean and SEM in a, b, e), * P < 0.05, ** P < 0.01, *** P < 0.001, two-tailed unpaired Student’s t-test
Supplementary information
Supplementary Figures
Supplementary Figures 1-8
Supplementary Tables
Supplementary Tables 1-6
Rights and permissions
About this article
Cite this article
Liu, S., Jiang, M., Wang, W. et al. Nuclear RNF2 inhibits interferon function by promoting K33-linked STAT1 disassociation from DNA. Nat Immunol 19, 41–52 (2018). https://doi.org/10.1038/s41590-017-0003-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0003-0
This article is cited by
-
USP33 promotes pancreatic cancer malignant phenotype through the regulation of TGFBR2/TGFβ signaling pathway
Cell Death & Disease (2023)
-
The RING finger protein family in health and disease
Signal Transduction and Targeted Therapy (2022)
-
Beyond K48 and K63: non-canonical protein ubiquitination
Cellular & Molecular Biology Letters (2021)
-
E3 ubiquitin ligases: styles, structures and functions
Molecular Biomedicine (2021)
-
A major quantitative trait locus affecting resistance to Tilapia lake virus in farmed Nile tilapia (Oreochromis niloticus)
Heredity (2021)