Abstract
Carbon dioxide is an omnipresent gas that drives adaptive responses within organisms from all domains of life. The molecular mechanisms by which proteins serve as sensors of CO2 are, accordingly, of great interest. Because CO2 is electrophilic, one way it can modulate protein biochemistry is by carboxylation of the amine group of lysine residues. However, the resulting CO2-carboxylated lysines spontaneously decompose, giving off CO2, which makes studying this modification difficult. Here we describe a method to stably mimic CO2-carboxylated lysine residues in proteins. We leverage this method to develop a quantitative approach to identify CO2-carboxylated lysines of proteins and explore the lysine ‘carboxylome’ of the CO2-responsive cyanobacterium Synechocystis sp. We uncover one CO2-carboxylated lysine within the effector binding pocket of the metabolic signaling protein PII. CO2-carboxylatation of this lysine markedly lowers the affinity of PII for its regulatory effector ligand ATP, illuminating a negative molecular control mechanism mediated by CO2.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All raw proteomics data were deposited to the ProteomeXchange Consortium via the PRIDE partner repository, with the dataset identifier (PXD031976). The hCit OXA-48 crystal structure has been deposited in the PDB with accession code 7LXG. The PDB codes for previously determined X-ray crystallographic structures used in the preparation of this manuscript are as follows: OXA-48 (4S2P), apo Synechocystis sp. PII (1UL3), ATP/2-OG bound S. elongatus PII (2XUL) and A. thaliana PII (2O66). The NCBI accession codes used in generating recombinant proteins for this study are as follows: K. pneumoniae OXA-48 (API82700), Synechocystis sp. PII (CAA66127.1) and A. thaliana PII (OAP00825.1). UniProt reference proteomes used in LysCarComp–MS analysis are as follows: E. coli K12 (UP000000625), Synechocystis sp. 6803 (UP000001425) and K. pneumoniae OXA-48 (Q6XEC0). All other data needed to evaluate the conclusions in the manuscript are available within the main text or supplementary materials. Source data are provided with this paper.
Code availability
No custom code was generated.
References
Cummins, E. P., Strowitzki, M. J. & Taylor, C. T. Mechanisms and consequences of oxygen and carbon dioxide sensing in mammals. Physiol. Rev. 100, 463–488 (2020).
Cummins, E. P., Selfridge, A. C., Sporn, P. H., Sznajder, J. I. & Taylor, C. T. Carbon dioxide-sensing in organisms and its implications for human disease. Cell. Mol. Life Sci. 71, 831–845 (2014).
Griffiths, H., Meyer, M. T. & Rickaby, R. E. M. Overcoming adversity through diversity: aquatic carbon concentrating mechanisms. J. Exp. Bot. 68, 3689–3695 (2017).
Mackinder, L. C. M. et al. A spatial interactome reveals the protein organization of the algal CO2-concentrating mechanism. Cell 171, 133–147 (2017).
Supuran, C. T. Structure and function of carbonic anhydrases. Biochem. J. 473, 2023–2032 (2016).
Chen, Y. et al. Soluble adenylyl cyclase as an evolutionarily conserved bicarbonate sensor. Science 289, 625–628 (2000).
Li, J. et al. Lysine carboxylation in proteins: OXA-10 beta-lactamase. Proteins 61, 246–257 (2005).
Ewing, S. P., Lockshon, D. & Jencks, W. P. Mechanism of cleavage of carbamate anions. J. Am. Chem. Soc. 102, 3072–3084 (1980).
Golemi, D., Maveyraud, L., Vakulenko, S., Samama, J. P. & Mobashery, S. Critical involvement of a carbamylated lysine in catalytic function of class D beta-lactamases. Proc. Natl. Acad. Sci. USA 98, 14280–14285 (2001).
Lorimer, G. H. & Miziorko, H. M. Carbamate formation on the epsilon-amino group of a lysyl residue as the basis for the activation of ribulosebisphosphate carboxylase by CO2 and Mg2+. Biochemistry 19, 5321–5328 (1980).
Jabri, E., Carr, M. B., Hausinger, R. P. & Karplus, P. A. The crystal structure of urease from Klebsiella aerogenes. Science 268, 998–1004 (1995).
Hong, S., Kuo, J. M., Mullins, L. S. & Raushel, F. M. CO2 is required for the assembly of the binuclear metal center of phosphotriesterase. J. Am. Chem. Soc. 117, 7580–7581 (1995).
Meigh, L. CO2 carbamylation of proteins as a mechanism in physiology. Biochem. Soc. Trans. 43, 460–464 (2015).
Li, Y. et al. Reversible post-translational carboxylation modulates the enzymatic activity of N-acetyl-l-ornithine transcarbamylase. Biochemistry 49, 6887–6895 (2010).
Stec, B. Structural mechanism of RuBisCO activation by carbamylation of the active site lysine. Proc. Natl. Acad. Sci. USA 109, 18785–18790 (2012).
Hacker, S. M. et al. Global profiling of lysine reactivity and ligandability in the human proteome. Nat. Chem. 9, 1181–1190 (2017).
Terrier, P. & Douglas, D. J. Carbamino group formation with peptides and proteins studied by mass spectrometry. J. Am. Soc. Mass. Spectrom. 21, 1500–1505 (2010).
Linthwaite, V. L. et al. The identification of carbon dioxide mediated protein post-translational modifications. Nat. Commun. 9, 1–11 (2018).
Forchhammer, K. & Lüddecke, J. Sensory properties of the PII signalling protein family. FEBS J. 283, 425–437 (2016).
Evers, J. et al. Molecular structure of isocyanic acid, HNCO, the imide of carbon dioxide. J. Phys. Chem. A. 122, 3287–3292 (2018).
Lippincott, J. & Apostol, I. Carbamylation of cysteine: a potential artifact in peptide mapping of hemoglobins in the presence of urea. Anal. Biochem. 267, 57–64 (1999).
Stark, G. R. Modification of proteins with cyanate. Enzym. Struct., Part B. 25, 590–594 (1967).
Stark, G. R., Stein, W. H. & Moore, S. Reactions of the cyanate present in aqueous urea with amino acids and proteins. J. Biol. Chem. 235, 3177–3181 (1960).
Steckel, A., Borbely, A., Uray, K. & Schlosser, G. Quantification of the effect of citrulline and homocitrulline residues on the collision-induced fragmentation of peptides. J. Am. Soc. Mass. Spectrom. 31, 1744–1750 (2020).
Lorimer, G. H. & Pierce, J. Carbonyl sulfide: an alternate substrate for but not an activator of ribulose-1,5-bisphosphate carboxylase. J. Biol. Chem. 264, 2764–2772 (1989).
Yamaguchi, K. & Hausinger, R. P. Subsitution of the urease active site carbamate by dithiocarbamate and vanadate. Biochemistry 36, 15118–15122 (1997).
Lohans, C. T. et al. (13)C-carbamylation as a mechanistic probe for the inhibition of class D beta-lactamases by avibactam and halide ions. Org. Biomol. Chem. 15, 6024–6032 (2017).
Mangan, N. M., Flamholz, A., Hood, R. D., Milo, R. & Savage, D. F. pH determines the energetic efficiency of the cyanobacterial CO2 concentrating mechanism. Proc. Natl. Acad. Sci. USA 113, E5354–E5362 (2016).
Walsh, M. A., Otwinowski, Z., Perrakis, A., Anderson, P. M. & Joachimiak, A. Structure of cyanase reveals that a novel dimeric and decameric arrangement of subunits is required for formation of the enzyme active site. Structure 8, 505–514 (2000).
Muroski, J. M., Fu, J. Y., Nguyen, H. H., Ogorzalek Loo, R. R. & Loo, J. A. Leveraging immonium ions for targeting acyl-lysine modifications in proteomic datasets. Proteomics 21, e2000111 (2021).
Sanders, J. D., Greer, S. M. & Brodbelt, J. S. Integrating carbamylation and ultraviolet photodissociation mass spectrometry for middle-down proteomics. Anal. Chem. 89, 11772–11778 (2017).
Price, G. D., Badger, M. R., Woodger, F. J. & Long, B. M. Advances in understanding the cyanobacterial CO2-concentrating-mechanism (CCM): functional components, Ci transporters, diversity, genetic regulation and prospects for engineering into plants. J. Exp. Bot. 59, 1441–1461 (2008).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Laing, W. A. & Christeller, J. T. A model for the kinetics of activation and catalysis of ribulose 1,5-bisphosphate carboxylase. Biochem. J. 159, 563–570 (1976).
McNevin, D., von Caemmerer, S. & Farquhar, G. Determining RuBisCO activation kinetics and other rate and equilibrium constants by simultaneous multiple non-linear regression of a kinetic model. J. Exp. Bot. 57, 3883–3900 (2006).
Watzer, B. et al. The signal transduction Protein PII controls ammonium, nitrate and urea uptake in cyanobacteria. Front. Microbiol. 10, 1428 (2019).
Zhao, M. X. et al. Crystal structure of the cyanobacterial signal transduction protein PII in complex with PipX. J. Mol. Biol. 402, 552–559 (2010).
Heinrich, A., Maheswaran, M., Ruppert, U. & Forchhammer, K. The Synechococcus elongatus PII signal transduction protein controls arginine synthesis by complex formation with N-acetyl-l-glutamate kinase. Mol. Microbiol. 52, 1303–1314 (2004).
Hauf, W., Schmid, K., Gerhardt, E. C., Huergo, L. F. & Forchhammer, K. Interaction of the nitrogen regulatory protein GlnB (PII) with biotin carboxyl carrier protein (BCCP) controls acetyl-CoA levels in the cyanobacterium Synechocystis sp. PCC 6803. Front. Microbiol. 7, 1700 (2016).
Orthwein, T. et al. The novel PII-interactor PirC identifies phosphoglycerate mutase as key control point of carbon storage metabolism in cyanobacteria. Proc. Natl. Acad. Sci. USA 118, e2019988118 (2021).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Xu, Y. et al. The structures of the PII proteins from the cyanobacteria Synechococcus sp. PCC 7942 and Synechocystis sp. PCC 6803. Acta Crystallogr. D. Biol. Crystallogr. D59, 2183–2190 (2003).
Fokina, O., Chellamuthu, V. R., Forchhammer, K. & Zeth, K. Mechanism of 2-oxoglutarate signaling by the Synechococcus elongatus PII signal transduction protein. Proc. Natl. Acad. Sci. USA 107, 19760–19765 (2010).
Osanai, T., Sato, S., Tabata, S. & Tanaka, K. Identification of PamA as a PII-binding membrane protein important in nitrogen-related and sugar-catabolic gene expression in Synechocystis sp. PCC 6803. J. Biol. Chem. 280, 34684–34690 (2005).
Wang, S. et al. Advanced activity-based protein profiling application strategies for drug development. Front. Pharm. 9, 353 (2018).
Pagels, F., Guedes, A. C., Amaro, H. M., Kijjoa, A. & Vasconcelos, V. Phycobiliproteins from cyanobacteria: chemistry and biotechnological applications. Biotechnol. Adv. 37, 422–443 (2019).
De Porcellinis, A. J. et al. Overexpression of bifunctional fructose-1,6-bisphosphatase/sedoheptulose-1,7-bisphosphatase leads to enhanced photosynthesis and global reprogramming of carbon metabolism in Synechococcus sp. PCC 7002. Metab. Eng. 47, 170–183 (2018).
Selim, K. A., Ermilova, E. & Forchhammer, K. From cyanobacteria to Archaeplastida: new evolutionary insights into PII signalling in the plant kingdom. N. Phytol. 227, 722–731 (2020).
Buck, J. & Levin, L. R. Physiological sensing of carbon dioxide/bicarbonate/pH via cyclic nucleotide signaling. Sensors 11, 2112–2128 (2011).
Rossetti, T., Jackvony, S., Buck, J. & Levin, L. R. Bicarbonate, carbon dioxide and pH sensing via mammalian bicarbonate-regulated soluble adenylyl cyclase. Interface Focus 11, 20200034 (2021).
King, D. T., King, A. M., Lal, S. M., Wright, G. D. & Strynadka, N. C. J. Molecular mechanism of avibactam-mediated beta-lactamase inhibition. ACS Infect. Dis. 1, 175–184 (2015).
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D. Biol. Crystallogr. 67, 271–281 (2011).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D. Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D. Biol. Crystallogr. 68, 352–367 (2012).
Speicher, K. D., Gorman, N. & Speicher, D. W. N-terminal sequence analysis of proteins and peptides. Curr. Protoc. Protein Sci. Chapter: Unit-11.10 (NIH, 2001).
Neilson, K. A. et al. The influence of signals from chilled roots on the proteome of shoot tissues in rice seedlings. Proteomics 13, 1922–1933 (2013).
Rossi, A. M. & Taylor, C. W. Analysis of protein-ligand interactions by fluorescence polarization. Nat. Protoc. 6, 365–387 (2011).
Butler, J. N. Carbon Dioxide Equilibria and their Applications (Lewis Publishers Inc., 1991).
Acknowledgements
We are grateful to the Natural Sciences and Engineering Research Council of Canada (NSERC) (grant no. RGPIN298406) and the Human Frontiers Science Program (grant no. RGP0058/2020) for supporting this research. D.J.V. thanks the Canada Research Chairs program for support as a Tier I Canada Research Chair in Chemical Biology. D.T.K. is supported by a postdoctoral fellowship from the Canadian Institutes of Health Research (CIHR) and a Trainee Award from the Michael Smith Foundation for Health Research and Pacific Alzheimer Research Foundation. We acknowledge the UVic-Genome BC Proteomics Centre, Victoria, Canada for performing all MS experiments. X-ray crystallography was performed at the Canadian Light Source synchrotron facility, a national research facility of the University of Saskatchewan, which is supported by the Canada Foundation for Innovation (CFI), NSERC, the National Research Council (NRC), CIHR, the Government of Saskatchewan and the University of Saskatchewan. We are grateful to L. Craig, S. Cecioni, Y. Zhu and M. Alteen for their expert input. Parts of the schematics were generated using BioRender.com.
Author information
Authors and Affiliations
Contributions
D.J.V. and D.T.K. conceived and designed experiments. D.T.K., Z.M. and S.K. performed plasmid construction and protein purification. D.T.K. performed biochemistry experiments, MS preparations and data analysis. D.T.K and J.E.S.N. performed X-ray crystallography and data analysis, D.B.H. performed MS, and D.T.K. and D.B.H. analyzed the MS data. S.Z. performed the NMR experiments and analyzed the data. D.J.V. and D.T.K. analyzed other data and wrote the manuscript with input from all.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Cong-Zhao Zhou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Homocitrullination of OXA-48 Lys73 blocks catalytic activity.
a-b, Mechanistic diagram of WT (Lys73-CO2) and homocitrullinated (hCit73) OXA-48 showing how hCit73 is unable to act as a general base and activate the nucleophilic serine.
Extended Data Fig. 2 In vitro OCNH-mediated homocitrullination of OXA-48, related to Fig. 1.
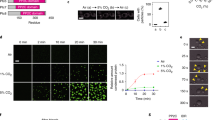
a, IC50 experiments comparing OXA-48 inhibition by CO2 analogues. Data represent mean values ± SD from n = 4 independent experiments. b, Far-UV CD spectra of OXA-48 pre-incubated in the presence of 50 mM NaCl or 50 mM KOCN. Data represent mean values ± SD from n = 4 independent experiments. c, Activity assays on samples from (b) taken immediately before (0 hrs) and after CD (6 hrs). OXA-48 remains inactivated throughout the CD experiment. Data represent means ± SD from n = 3 independent experiments. d, Jump-dilution kinetics experiments to detect time-dependant decarbamylation of OXA-48. Data represent mean values ± SD from n = 6 independent experiments. e, Anti-His immunoblot for OXA-48 samples from (d) incubated in the presence of either 50 mM NaCl or 50 mM KOCN at 0- and 48-hour timepoints. Samples were centrifuged to remove insoluble aggregates prior to immunoblotting. This immunoblot was performed once. f, Immunoblot competition assays for WT and K73A OXA-48. The image shown in f is a representative image from three independent experiments. g, Densitometry analysis of select bands from (f). The chart displays mean ± SD from two experimental replicates with P values: two tailed student’s t-test assuming unequal variance; **P < 0.01. The indicated significant P values in (g) are as follows: (50 mM KOCN) vs. (50 mM KOCN, 50 mM NaHCO3) = 0.00346, (50 mM KOCN, 50 mM NaHCO3) vs. (50 mM KOCN, 50 mM NaCl) = 0.00337. h, Crystallographic evidence for hCit73 OXA-48. The hCit73 omit Fo-Fc electron density map contoured at 3.5 σ is shown overlaid on the final refined coordinates. The protein backbone is displayed in magenta cartoon with key active site residues displayed as white sticks with heteroatoms coloured by type. The hCit73 residue is displayed as green sticks. Select hydrogen bonds are shown as blue dashes.
Extended Data Fig. 3 CO2 competes with OCNH on OXA-48 in a concentration dependant manner, related to Fig. 2.
a, Immunoblot competition assay performed on purified OXA-48 with detection using a polyclonal anti-hCit antibody. b, Densitometry analysis of relative OXA-48 band intensities in a. Data points are normalized to the Anti-His control band and then again to the zero NaHCO3 control. Data are fit to a three-parameter sigmoidal dose-response curve in GraphPad Prism.
Extended Data Fig. 4 Lys-CO2 modification of RuBisCO large subunit identified by proteome wide LysCarComp-MS performed on Synechocystis sp., related to Fig. 4.
a, LC-MS/MS spectra showing unambiguous evidence for hCit196 residue within TMTduplex-labelled RuBisCO large subunit peptide [TMT-GGLDFThCit(K196 + 43.01 Da)DDENINSQPFMR, MH + [Da] = 2452.16]. b, Close-up of TMTduplex reporter ions and immonium ions derived from (a). The mean RCO2 with SD from three independent experimental replicates is given.
Extended Data Fig. 6 CO2 blocks BODIPY FL ATP-γ-S binding to PII from Synechocystis sp., related to Fig. 5.
a, FP equilibrium dissociation binding of BODIPY FL ATP-γ-S to WT and K90A SsPII. b-c, FP equilibrium competition binding experiments for WT (b) and K90A SsPII (c) in the presence of varying concentrations of small molecules (2-OG, GTP, UDP, ADP, ATP). d, Time-dependant effect of KOCN on FP equilibrium probe binding to WT and K90A SsPII. e, FP equilibrium binding of BODIPY FL ATP-γ-S following pre-incubation of SsPII in the presence and absence of KOCN followed by jump dilution and desalting to remove residual KOCN. In (a-e), all data is presented as mean values ± SD from n = 2 independent experiments. f, FP equilibrium competition binding experiments for WT and K90A SsPII in the presence of varying concentrations of HCO3−. Data in (f) are presented as mean values ± SD from n = 4 independent experiments. In (b) and (f), KD’s for competitor compounds were calculated from IC50 values using a standard FP equilibrium binding formula as previously described59. KD(CO2) was determined from KD using standard equations for carbonate equilibria as previously described60.
Extended Data Fig. 7 Biolayer interferometry confirms that CO2 blocks ATP binding to SsPII, related to Fig. 5.
a, Chemical structure of the biotin-conjugated ATP analogue used in BLI. b-c, Representative independent duplicate BLI association/dissociation curves for WT SsPII run in the absence (b) and presence (c) of 50 mM HCO3−. d, Representative BLI association/dissociation curves for K90A SsPII run in the absence of HCO3−. SsPII protein concentrations and curves are exactly as shown in b-c. e, Equilibrium response ratio for SsPII binding to the ATP-biotin probe in the presence and absence of 50 mM HCO3−. Req values correspond to the response value at the 240 s timepoint in b and c. Data are presented as mean values ± SD from three independent experiments.
Extended Data Fig. 8 Lys164-CO2 modification blocks ATP binding to PII from Arabidopsis thaliana.
a, Active site overlay of apo SsPII and apo Arabidopsis thaliana PII (AtPII, 53.6% sequence identity, PDB IDs: 1UL3 and 2O66). The SsPII and AtPII protein chains are displayed as green and yellow cartoons with select residues shown as sticks with atoms coloured by type. b, 13C NMR using purified WT and K164A AtPII in the presence and absence of 50 mM NaH13CO3. c, FP equilibrium dissociation binding of BODIPY FL ATP-γ-S to WT and K164A AtPII. d, FP equilibrium competition binding experiments for WT and K164A AtPII in the presence of varying concentrations of HCO3−. In c-d, data are presented as mean values ± SD from n = 2 independent experiments. In d, KD’s for competitor compounds were calculated from IC50 values using a standard FP equilibrium binding formula as previously described59. KD(CO2) was determined from KD using standard equations for carbonate equilibria as previously described60.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9, Tables 1–5 and Uncropped gel images.
Supplementary Data 1
Excel-based data table summarizing all LysCarComp–MS proteomics data. The table includes the amino acid sequence, accession numbers, modification site(s), missed cleavages, PSMs, confidence scores (XCorr) and relative reporter ion intensities across all replicates for every peptide identified.
Supplementary Data 2
Excel-based data table with source data for Supplementary Fig. 2. Includes an LC–MS/MS analysis of various OCNH adducts on tryptic peptides derived from OXA-48.
Supplementary Data 3
Excel-based data table with source data for Supplementary Fig. 5. Includes a kinetic analysis of OXA-48 activity following addition of various amounts of N3-PEG4-NCO.
Supplementary Data 4
Excel-based data table with source data for Supplementary Fig. 6. Includes a kinetic analysis of OXA-48 activity determined at different pH values.
Supplementary Data 5
Excel-based data table with source data for Supplementary Fig. 7. Includes data for aggregation assays performed using cellular lysates in the presence and absence of both OCNH and HCO3−. Also, the table includes a kinetic analysis of OXA-48 activity determined in the presence and absence of supplementary carbonic anhydrase.
Supplementary Data 6
Excel-based data table with source data for Supplementary Fig. 8. Includes LC–MS/MS data used in mapping the synthetic hCit OXA-48 peptide.
Source data
Source Data Fig. 1
Source data for Fig. 1c–e.
Source Data Fig. 2
Source data for Fig. 2a,b,d,f.
Source Data Fig. 2
Unprocessed western blots for Fig. 2c,e.
Source Data Fig. 3
Source data for Fig. 3b,d.
Source Data Fig. 4
Source data for Fig. 4a,c,d.
Source Data Fig. 4
Unprocessed western blots for Fig. 4b.
Source Data Fig. 5
Source data for Fig. 5a–c,f,g.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2a–d,g.
Source Data Extended Data Fig. 2
Unprocessed western blots for Extended Data Fig. 2e,f.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3b.
Source Data Extended Data Fig. 3
Unprocessed western blots for Extended Data Fig. 3a.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4a,b.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6a–f
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7b–e.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8b–d.
Rights and permissions
About this article
Cite this article
King, D.T., Zhu, S., Hardie, D.B. et al. Chemoproteomic identification of CO2-dependent lysine carboxylation in proteins. Nat Chem Biol 18, 782–791 (2022). https://doi.org/10.1038/s41589-022-01043-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01043-1
This article is cited by
-
Carbon dioxide and MAPK signalling: towards therapy for inflammation
Cell Communication and Signaling (2023)
-
Catch your breath
Nature Chemical Biology (2022)