Abstract
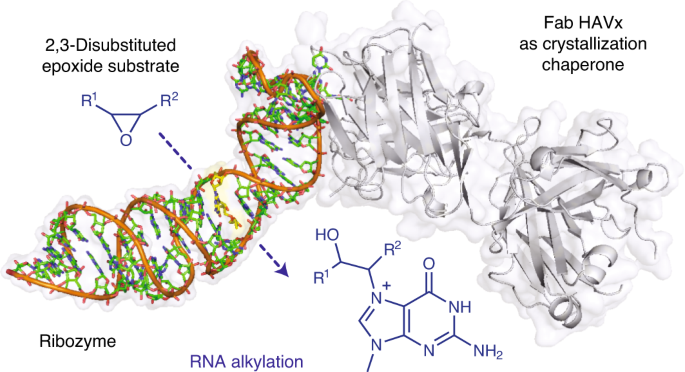
Ribozymes that react with small-molecule probes have important applications in transcriptomics and chemical biology, such as RNA labeling and imaging. Understanding the structural basis for these RNA-modifying reactions will enable the development of better tools for studying RNA. Nevertheless, high-resolution structures and underlying catalytic mechanisms for members of this ribozyme class remain elusive. Here, we focus on a self-alkylating ribozyme that catalyzes nitrogen–carbon bond formation between a specific guanine and a 2,3-disubstituted epoxide substrate and report the crystal structures of a self-alkylating ribozyme, including both alkylated and apo forms, at 1.71-Å and 2.49-Å resolution, respectively. The ribozyme assumes an elongated hairpin-like architecture preorganized to accommodate the epoxide substrate in a hook-shaped conformation. Observed reactivity of substrate analogs together with an inverse, log-linear pH dependence of the reaction rate suggests a requirement for epoxide protonation, possibly assisted by the ether oxygens within the substrate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates and structure factors for the reported crystal structures have been deposited at the PDB under accession codes 6XJZ, 6XJY, 6XJW and 6XJQ for the SAR structures corresponding to the non-alkylated (apo), short incubation with biotinylated substrate (2), alkylated with non-biotinylated substrate (1) and alkylated with biotinylated substrate (2) forms, respectively. The raw data, additional information and materials, including purified Fab HAVx and plasmid for its expression, will be provided by the authors upon request. The requests should be addressed to J.A.P. Source data are provided with this paper.
References
Leung, E. K. Y., Suslov, N., Tuttle, N., Sengupta, R. & Piccirilli, J. A. The mechanism of peptidyl transfer catalysis by the ribosome. Annu. Rev. Biochem. 80, 527–555 (2011).
Lassila, J. K., Zalatan, J. G. & Herschlag, D. Biological phosphoryl-transfer reactions: understanding mechanism and catalysis. Annu. Rev. Biochem. 80, 669–702 (2011).
Ekland, E. H., Szostak, J. W. & Bartel, D. P. Structurally complex and highly active RNA ligases derived from random RNA sequences. Science 269, 364–370 (1995).
Bartel, D. P. & Szostak, J. W. Isolation of new ribozymes from a large pool of random sequences [see comment]. Science 261, 1411–1418 (1993).
Piccirilli, J. A., McConnell, T. S., Zaug, A. J., Noller, H. F. & Cech, T. R. Aminoacyl esterase activity of the Tetrahymena ribozyme. Science 256, 1420–1424 (1992).
Ma, L. & Liu, J. Catalytic nucleic acids: biochemistry, chemical biology, biosensors, and nanotechnology. iScience 23, 100815 (2020).
Wilson, C. & Szostak, J. W. In vitro evolution of a self-alkylatlng ribozyme. Nature 374, 777–782 (1995).
Wecker, M., Smith, D. & Gold, L. In vitro selection of a novel catalytic RNA: characterization of a sulfur alkylation reaction and interaction with a small peptide. RNA 2, 982–994 (1996).
Ameta, S. & Jäschke, A. An RNA catalyst that reacts with a mechanistic inhibitor of serine proteases. Chem. Sci. 4, 957–964 (2013).
Sharma, A. K. et al. Fluorescent RNA labeling using self-alkylating ribozymes. ACS Chem. Biol. 9, 1680–1684 (2014).
Scheitl, C. P. M., Ghaem Maghami, M., Lenz, A.-K. & Höbartner, C. Site-specific RNA methylation by a methyltransferase ribozyme. Nature 587, 663–667 (2020).
McDonald, R. I. et al. Electrophilic activity-based RNA probes reveal a self-alkylating RNA for RNA labeling. Nat. Chem. Biol. 10, 1049–1054 (2014).
Bevilacqua, P. C. et al. An ontology for facilitating discussion of catalytic strategies of RNA-cleaving enzymes. ACS Chem. Biol. 14, 1068–1076 (2019).
Gaines, C. S., Piccirilli, J. A. & York, D. M. The L-platform/L-scaffold framework: a blueprint for RNA-cleaving nucleic acid enzyme design. RNA 26, 111–125 (2020).
Huang, H. et al. A G-quadruplex-containing RNA activates fluorescence in a GFP-like fluorophore. Nat. Chem. Biol. 10, 686–691 (2014).
Koirala, D. et al. Affinity maturation of a portable Fab–RNA module for chaperone-assisted RNA crystallography. Nucleic Acids Res. 46, 2624–2635 (2018).
Shelke, S. A. et al. Structural basis for activation of fluorogenic dyes by an RNA aptamer lacking a G-quadruplex motif. Nat. Commun. 9, 4542 (2018).
Koldobskaya, Y. et al. A portable RNA sequence whose recognition by a synthetic antibody facilitates structural determination. Nat. Struct. Mol. Biol. 18, 100–106 (2011).
Ferré-D’Amaré, A. R. & Doudna, J. A. Crystallization and structure determination of a hepatitis delta virus ribozyme: use of the RNA-binding protein U1A as a crystallization module. J. Mol. Biol. 295, 541–556 (2000).
Koirala, D. et al. A conserved RNA structural motif for organizing topology within picornaviral internal ribosome entry sites. Nat. Commun. 10, 3629 (2019).
Nozière, B. et al. The hydrolysis of epoxides catalyzed by inorganic ammonium salts in water: kinetic evidence for hydrogen bond catalysis. Phys. Chem. Chem. Phys. 20, 1583–1590 (2018).
Yan, Y., Tao, H., He, J. & Huang, S.-Y. The HDOCK server for integrated protein–protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Perrotta, A. T., Shih, I & Been, M. D. Imidazole rescue of a cytosine mutation in a self-cleaving ribozyme. Science 286, 123–126 (1999).
Piletic, I. R., Edney, E. O. & Bartolotti, L. J. A computational study of acid catalyzed aerosol reactions of atmospherically relevant epoxides. Phys. Chem. Chem. Phys. 15, 18065–18076 (2013).
Bevilacqua, P. C., Brown, T. S., Nakano, S. I. & Yajima, R. Catalytic roles for proton transfer and protonation in ribozymes. Biopolymers 73, 90–109 (2004).
Das, S. R. & Piccirilli, J. A. General acid catalysis by the hepatitis delta virus ribozyme. Nat. Chem. Biol. 1, 45–52 (2005).
Zhang, J., Lau, M. W. & Ferré-D’Amaré, A. R. Ribozymes and riboswitches: modulation of RNA function by small molecules. Biochemistry 49, 9123–9131 (2010).
Cochrane, J. C. & Strobel, S. A. Catalytic strategies of self-cleaving ribozymes. Acc. Chem. Res. 41, 1027–1035 (2008).
Viladoms, J. & Fedor, M. J. The glmS ribozyme cofactor is a general acid–base catalyst. J. Am. Chem. Soc. 134, 19043–19049 (2012).
Wilson, T. J., Liu, Y. & Lilley, D. M. J. Ribozymes and the mechanisms that underlie RNA catalysis. Front. Chem. Sci. Eng. 10, 178–185 (2016).
Ye, J.-D. et al. Synthetic antibodies for specific recognition and crystallization of structured RNA. Proc. Natl Acad. Sci. USA 105, 82–87 (2008).
Shao, Y. et al. Specific recognition of a single-stranded RNA sequence by a synthetic antibody fragment. J. Mol. Biol. 428, 4100–4114 (2016).
Kao, C., Rüdisser, S. & Zheng, M. A simple and efficient method to transcribe RNAs with reduced 3′ heterogeneity. Methods 23, 201–205 (2001).
Rio, D. C. Expression and purification of active recombinant T7 RNA polymerase from E. coli. Cold Spring Harb. Protoc. 2013, pdb.prot078527 (2013).
Hall, B. et al. In vitro selection of RNA aptamers to a protein target by filter immobilization. Curr. Protoc. Nucleic Acid Chem. 40, 9.3.1–9.3.27 (2010).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Bailey, T. L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Goujon, M. et al. A new bioinformatics analysis tools framework at EMBL–EBI. Nucleic Acids Res. 38, W695–W699 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Acknowledgements
This work was supported by grants from the National Institutes of Health (R01AI081987, R01GM102489) and the Chicago Biomedical Consortium with support from the Searle Funds at The Chicago Community Trust to J.A.P. and start-up funds from the University of Maryland Baltimore County to D. Koirala. The crystallographic work is based on research conducted at the Advanced Photon Source on the Northeastern Collaborative Access Team beamlines, 24-ID-C and 24-ID-E, which is supported by a grant from the National Institute of General Medical Sciences (P41 GM103403) from the National Institutes of Health. This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract number DE-AC02-06CH11357. We would like to thank the staff of the Advanced Photon Source at Argonne National Laboratory for providing technical advice during data collection. We are also thankful to B. Weissman, a former Piccirilli laboratory member, for helping with pH data analysis and all current Piccirilli laboratory members for critical review of the manuscript.
Author information
Authors and Affiliations
Contributions
D. Krochmal, D. Koirala and J.A.P. conceived and designed the experiments. D. Krochmal and D. Koirala prepared the samples and conducted most of the crystallographic and biochemical experiments. Y.S. phased and solved the crystal structures with D. Koirala. N.-S.L. synthesized and purified the substrates. S.D. and S.A.S. made contributions to design and set up crystallization trials with initial constructs. D. Krochmal and D. Koirala analyzed most of the biochemical and crystallization data and interpreted the results with J.A.P. D. Krochmal, D. Koirala and J.A.P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Solen Ekesan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the self-alkylating ribozyme (SAR) crystallization construct.
(a) Biochemically derived secondary structure of the self-alkylating ribozyme, showing the site of alkylation and critical nucleotides required for the reactivity according to McDonald et al2. (b) In the crystallization construct, the variable loop of the RNA is replaced by the Fab HAVx binding loop sequence. (c) Representative plots of fraction RNA bound as a function of varying concentration of Fab HAVx for the parent HAV IRES dV (red circles), 4H SM hairpin (blue diamonds, see supplementary Fig. 2d for the hairpin sequence), and the SAR crystallization construct (green triangles) as accessed from filter binding assay in 25 mM HEPES-Na pH 7.4, 150 mM NaCl, 10 mM MgCl2 buffer at 23 °C. The reported Kds represent the values (average ± standard deviation) obtained from 3 independent measurements. (d) The crystallization construct is catalytically active in the presence or absence of Fab HAVx. These results are representatives of 3 independent experiments. See methods for details of the assay.
Extended Data Fig. 2 Overall crystal structures of the apo SAR-HAVx complexes.
(a) The structure of the apo form of the SAR in complex with Fab HAVx solved at 2.49 Å resolution. (b) The same complex structure was solved at 2.16 Å resolution but pre-incubated with the biotinylated epoxide (substrate 2) for 30 minutes before setting up the crystallization trials. Both crystals contained two complexes per asymmetric unit. The RNA molecules are colored blue and green for clarity. In the crystal lattice, including the epitope-paratope binding interface, Fab HAVx interactions account for 95% and 94% of the buried surface area, respectively. The RNA-RNA interactions (5% and 6% of the buried surface area, respectively) involve end-to-end stacking of terminal helices from symmetry-related RNA molecules.
Extended Data Fig. 3 Overall crystal structures of the alkylated SAR-HAVx complexes.
(a) The structure of the alkylated SAR complex with Fab HAVx solved at 1.71 Å resolution. The ribozyme was pre-reacted with the biotinylated epoxide (substrate 2) for 24 hours before setting up the crystallization trials. (b) Structure of the alkylated SAR complex with Fab HAVx solved at 1.91 Å resolution. The ribozyme was pre-reacted with the non-biotinylated epoxide (substrate 1) for 24 hours before setting up the crystallization trials. Both crystals contained two complexes per asymmetric unit. The RNA molecules are colored blue and green for clarity. In both cases, including the epitope-paratope binding interface, Fab HAVx interactions account for all crystal contacts.
Extended Data Fig. 4 Comparison of the structures of SAR catalytic core pre-alkylated with biotinylated and non-biotinylated epoxide substrates.
(a) Superposition of the catalytic cores of the SAR pre-alkylated with biotinylated (red) and the non-biotinylated (yellow) substrates (RMSD: 0.119 Å). The substrate structure after alkylation, which is linked covalently with the N7 of ribozyme’s G16 nucleobase, for (b) biotinylated and (c) non-biotinylated substrates. (d) Superposition of the biotinylated and non-biotinylated substrates within the catalytic cores (as shown in a), including G16 nucleotides. Both substrates assume similar hook-shaped conformations with minor deviations in some areas, which could be due to the differences in the resolution of the crystallographic data and the substrate modeling with relatively poor electron density in the region, especially for the non-biotinylated substrate. The green (b and d) and blue (c and d) meshes represent the 2|Fo| − |Fc| electron density map at 1σ contour level and carve radius 1.8 Å. All figures are colored analogously for facile comparison.
Extended Data Fig. 5 Comparison of apo and alkylated RNA structures.
(a) Superposition of the crystal structures of the self-alkylating ribozyme in apo (blue), alkylated with non-biotinylated (yellow), and with biotinylated (red) forms. The RMSDs for superpositions of apo and alkylated with the non-biotinylated substrate, apo and alkylated with the biotinylated substrate, and alkylated with non-biotinylated substrate and with the biotinylated substrate are 2.507 Å, 2.574 Å, 0.141 Å, respectively. These structures are almost identical, including the catalytic core structure (b), suggesting that the ribozyme is pre-organized for substrate binding and subsequent catalysis. Minor deviations in some areas could be due to the differences in crystal packing or the resolution of the crystallographic data (the alkylated form with biotinylated substrate: 1.71 Å, alkylated form with non-biotinylated substrate: 1.91 Å and apo form: 2.49 Å).
Extended Data Fig. 6 Comparison of two SAR-HAVx complexes within the crystallographic asymmetric unit.
(a) The superposition of two Fab-RNA complexes within the crystallographic asymmetric unit (colored blue and green) for the apo SAR – HAVx complex (RMSD: 0.477 Å), (b) the SAR – HAVx complex that was pre-incubated with the substrate 2 for 30 minutes before crystallization trials (RMSD: 0.573 Å), (c) the SAR – HAVx complex alkylated with substrate 1 (RMSD: 0.502 Å), and (d) the SAR – HAVx complex alkylated with substrate 2 (RMSD: 0.559 Å). Overall, two complexes within the asymmetric unit are similar except for some deviations in RNA helices peripheral to the catalytic core for the alkylated forms (c, d). The superposition of the catalytic cores is also shown for the alkylated ribozymes. The substrate conformation appears slightly different, perhaps due to modeling errors, as the electron density map for the substrates is the weakest within the catalytic core.
Extended Data Fig. 7 Structural features of the Fab-RNA binding interface.
(a) Molecular surface of the Fab HAVx and a cartoon of the RNA epitope showing the Fab CDRs (L1, L2, L3, H1, H2, and H3) and the interacting nucleotides of the RNA epitope. Three consecutive Cs within the RNA epitope (C27-C29) that do not directly interact with the Fab residues are not shown for clarity. (b) Schematic summary of interactions between the RNA epitope and the Fab residues. Orange spheres represent water molecules that mediate hydrogen bonds. Hydrogen bonding and stacking interactions are indicated by — and |—|, respectively. Epitope nucleotides and Fab CDR residues are colored analogously in both panels for facile comparisons. The residues in gray represent scaffold residues from the constant part of the light chain’s variable domain. Analysis of the engaged interfaces in all four Fab-RNA structures using PDBePISA (http://www.ebi.ac.uk/pdbe/pisa/) reveals that within the epitope-paratope interface involving Fab CDRs, the Fab-RNA interactions bury a total of 1091 ± 22 Å2 (average ± standard deviation for the four structures). The heavy- and light-chain CDRs contribute equally to the buried surface area (50 ± 1%). This same Fab HAVx, when bound to a bulge within the HAV IRES domain V, buries 1315 Å2 of surface area, with the heavy- and light-chain CDRs contributing 819 Å2 (62%) and 496 Å2 (38%), respectively20. For comparison, the interfacial area observed here is higher than that observed Fab BL3-6 – hairpin RNA epitope complex (821 Å2)15 but lower than that observed for Fab2 – P4P6 RNA complex (1316 Å2)31. The Fab binding RNA motif adopts a well-structured fold and contains two loops, L3a and L3b, which harbor three (U24-G26) and six (G30-C35) nucleotides, respectively, separated by three coaxially stacked Cs (C27-C29). Tertiary interactions between L3a and L3b loops involving two base triples, U25-U34•G30 and G26-C35•C29, stabilize the overall architecture of the motif. Within the interface with Fab HAVx, the L3a and L3b loops interact with the residues of two separate pockets formed by Fab CDRs L1, L2, L3, and H3, and H1, H2, H3, and L3, respectively. The highly conserved C27-C29 nucleotides do not interact with the Fab but serve as a spacer that allows correct positioning of the L3a and L3b loops within these Fab binding pockets.
Extended Data Fig. 8 Specific interactions between the RNA epitope nucleotides and the Fab light-chain residues.
(a) Beginning from the P3 helix, the CDR-L1 serine (S29) and some scaffold residues, including glutamine (Q28) and arginine (R67) from the light chain’s variable domain, make hydrogen bonding (direct and water-mediated) and electrostatic interactions with the RNA phosphate backbone. (b, c) Continuing upward from the P3 helix, two CDR-L1 tyrosines (Y31 and Y32) and three CDR-L2 residues (S51, S53, and Y54) recognize the L3a loop through stacking and hydrogen bonding interactions and a CDR-L1 serine (S33) makes a hydrogen bond with the L3b U34. (d) The residues from CDR-H3, including tyrosines (Y109 and Y110), and an asparagine (N108), interact with the nucleotides U25, G30 and U34. The yellow, cyan, and orange represent the residues from CDR L1, L2, and H3, respectively. Magenta and blue represent nucleotides from the SAR P3 helix and RNA epitope loop, respectively. The dashed lines reflect heteroatoms within hydrogen-bonding distance. Orange spheres represent water molecules. Blue mesh represents the 2|Fo| − |Fc| electron density map at 1σ contour level and carve radius 1.8 Å. Gray residues represent the scaffold residues from the constant part of the light chain’s variable domain.
Extended Data Fig. 9 Specific interactions between the RNA epitope nucleotides and the Fab heavy-chain residues.
(a) The residues from CDR-H3, including a tyrosine (Y104), a histidine (H105), an asparagine (N108), and a tryptophan (W111), interact with the nucleotides U32, A33, and U34. (b, c) At the top of the loop, three CDR-H1 (S33, S35, and S36) and three CDR-H2 (S55, S58, and S60) serines and a CDR-H2 tyrosine (Y57) contact U32 and A33 through an extensive network of direct and water-mediated hydrogen bonding and stacking interactions. (d) Among three consecutive CDR-L3 arginines, R95 and R94 contact L3b U34 and C20 via the phosphate backbone, respectively, through hydrogen bonding and electrostatic interactions. The green, red, orange, and purple represent the residues from CDR H1, H2, H3, and L3, respectively. Blue and magenta (C20) represent nucleotides from the SAR P3 helix and RNA epitope loop. The dashed lines reflect heteroatoms within hydrogen-bonding distance. Orange spheres represent water molecules. Blue mesh represents the 2|Fo| − |Fc| electron density map at 1σ contour level and carve radius 1.8 Å.
Extended Data Fig. 10 Comparison of Fab HAVx binding hairpin loop and asymmetric bulge epitopes.
(a) Molecular surface of the Fab HAVx and a cartoon of the RNA epitope showing the Fab CDRs (L1, L2, L3, H1, H2, and H3) and the interacting nucleotides of the RNA epitope in SAR-HAVx complex (hairpin-loop epitope) and (b) in HAV dV-HAVx complex20 (bulge epitope). Molecular surface of the Fab HAVx and a cartoon of the common RNA nucleotides (UUAU), showing Fab-RNA interactions in the context of (c) hairpin loop epitope (L3b) and (d) bulge epitope (right-side of the bulge). (e) Superposition of common nucleotide sequence (UAU) from these epitopes on the molecular surface of the Fab HAVx from the SAR-HAVx complex. A cartoon of the other set of common nucleotides (UUG), showing Fab-RNA interactions in the context of (f) hairpin loop epitope (loop L3a) and (g) bulge epitope (left-side of the bulge). Most of the interactions within the epitope-paratope interfaces are analogous. Compared to the only four of the six CDRs, L1, L3, H2, and H3 of the Fab HAVx being used in the bulge epitope and HAVx interface, all six CDRs engage in the in vitro selected epitope and HAVx interactions. In contrast to a few hydrogen-bonding and electrostatic interactions observed in HAVx-loop interface for three consecutive CDR-L3 residues R93, R94, and R95, these residues in the HAVx-bulge structure interact heavily with two uridine nucleobases. Within the bulge, CDR-H3 Y109 stacks with a uridine nucleobase, but in the HAVx-loop interface, both Y110 and Y109 stack with a uridine (U34) and the guanine (G30) nucleobases, respectively. Although most of the interactions are well-defined and comparable, direct comparison of many other interactions between HAVx-bulge and HAVx-loop might be ambiguous due to a large difference in resolution of the crystal structures (HAVx-loop: 1.71 Å vs. HAVx-bulge: 2.84 Å). In both the HAVx-bulge and HAVx-loop interfaces, the CDR-L1 residues, a serine (S29) and a tyrosine (Y31 for HAVx-bulge but Y32 for HAVx-loop interface), and a scaffold residue, R67, interact with the phosphate backbone of a paired helical stem through direct and water-mediated hydrogen bonding and electrostatic interactions. Similarly, the heavy chain CDRs H2 and H3 make direct or water-mediated hydrogen bonds and stacking interactions with the RNA with CDR-H3 utilizing the highest number of residues among all CDRs. The CDR-H2 Y57 and CDR-H3 Y104 sandwich an uracil nucleobase from top and bottom, respectively. The S55 and S58 interact with phosphate backbone and 2′-OH within the consecutive A and U nucleotides in a similar fashion. Within the asymmetric bulge, unpaired sequence UUAU analogous to L3b sequence spans through a pocket formed at the interface of CDRs H1, H2, H3, and L3, interacting with the residues Y104, H105, N108, Y109, Y110, and W111, analogous to those observed in the HAVx-loop interface. Nevertheless, N108 and Y109 do not interact directly with the UUAU sequence in the HAVx-loop interface. We also note that loop L3a consists of a UUG sequence identical to that located opposite the UUAU sequence within the asymmetric bulge epitope. However, the interactions of this sequence with the Fab HAVx in the context of loop L3a and asymmetric bulge are different. The interactions of L3a involve several residues from the interface of L1, L2, L3, and H3 CDRs, whereas within the bulge, we observed only a few interactions involving CDRs L1 and H3 (see Extended Data Figures 7–9 for details).
Supplementary information
Supplementary Information
Supplementary Figs. 1–5, Table 1, Notes 1–3 and additional Supplementary Figs. 1 and 2.
Source data
Source Data Extended Data Fig. 1
Unmodified gel image for Extended Data Fig. 1d.
Rights and permissions
About this article
Cite this article
Krochmal, D., Shao, Y., Li, NS. et al. Structural basis for substrate binding and catalysis by a self-alkylating ribozyme. Nat Chem Biol 18, 376–384 (2022). https://doi.org/10.1038/s41589-021-00950-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00950-z
This article is cited by
-
Crystal structure of a highly conserved enteroviral 5′ cloverleaf RNA replication element
Nature Communications (2023)
-
A SAM analogue-utilizing ribozyme for site-specific RNA alkylation in living cells
Nature Chemistry (2023)
-
A new RNA performs old chemistry
Nature Chemical Biology (2022)