Abstract
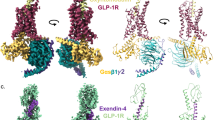
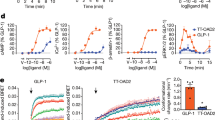
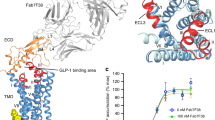
Recent advances in G-protein-coupled receptor (GPCR) structural elucidation have strengthened previous hypotheses that multidimensional signal propagation mediated by these receptors depends, in part, on their conformational mobility; however, the relationship between receptor function and static structures is inherently uncertain. Here, we examine the contribution of peptide agonist conformational plasticity to activation of the glucagon-like peptide 1 receptor (GLP-1R), an important clinical target. We use variants of the peptides GLP-1 and exendin-4 (Ex4) to explore the interplay between helical propensity near the agonist N terminus and the ability to bind to and activate the receptor. Cryo-EM analysis of a complex involving an Ex4 analog, the GLP-1R and Gs heterotrimer revealed two receptor conformers with distinct modes of peptide–receptor engagement. Our functional and structural data, along with molecular dynamics (MD) simulations, suggest that receptor conformational dynamics associated with flexibility of the peptide N-terminal activation domain may be a key determinant of agonist efficacy.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequencing data for NLuc-GLP-1R is available at Addgene (ID 124831). Atomic coordinates and cryo-EM density maps for Ex4-d-Ala-bound GLP-1R–Gs in conformer 1 and conformer 2 have been deposited in the PDB under accession numbers 7S1M and 7S3I and Electron Microscopy Data Bank entries EMD-24805 and EMD-24825, respectively. MD trajectories are available at https://doi.org/10.5281/zenodo.5578864. Source data are provided with this paper.
Code availability
No new code was used in this study. A list of software used is available in the Methods section and Reporting Summary.
References
Hilger, D., Masureel, M. & Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 25, 4–12 (2018).
Latorraca, N. R., Venkatakrishnan, A. J. & Dror, R. O. GPCR dynamics: structures in motion. Chem. Rev. 117, 139–155 (2017).
Graaf, C. D. et al. Glucagon-like peptide-1 and its class B G protein-coupled receptors: a long march to therapeutic successes. Pharmacol. Rev. 68, 954–1013 (2016).
Weis, W. I. & Kobilka, B. K. The molecular basis of G protein-coupled receptor activation. Annu. Rev. Biochem. 87, 897–919 (2018).
Runge, S., Thøgersen, H., Madsen, K., Lau, J. & Rudolph, R. Crystal structure of the ligand-bound glucagon-like peptide-1 receptor extracellular domain. J. Biol. Chem. 283, 11340–11347 (2008).
Zhang, X. et al. Differential GLP-1R binding and activation by peptide and non-peptide agonists. Mol. Cell 80, 485–500 (2020).
Liang, Y.-L. et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor–Gs complex. Nature 555, 121–125 (2018).
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017).
Dong, M. et al. Structure and dynamics of the active Gs-coupled human secretin receptor. Nat. Commun. 11, 4137 (2020).
Liang, Y.-L. et al. Toward a structural understanding of class B GPCR peptide binding and activation. Mol. Cell 77, 656–668 (2020).
Qiao, A. et al. Structural basis of Gs and Gi recognition by the human glucagon receptor. Science 367, 1346–1352 (2020).
Liang, Y.-L. et al. Phase-plate cryo-EM structure of a class B GPCR–G-protein complex. Nature 546, 118–123 (2017).
Hoang, H. N. et al. Short hydrophobic peptides with cyclic constraints are potent glucagon-like peptide-1 receptor (GLP-1R) agonists. J. Med. Chem. 58, 4080–4085 (2015).
Neidigh, J. W., Fesinmeyer, R. M., Prickett, K. S. & Andersen, N. H. Exendin-4 and glucagon-like-peptide-1: NMR structural comparisons in the solution and micelle-associated states. Biochemistry 40, 13188–13200 (2001).
Neumann, J.-M. et al. Class-B GPCR activation: is ligand helix-capping the key? Trends Biochem. Sci. 33, 314–319 (2008).
Oddo, A. et al. α-Helix or β-turn? An investigation into N-terminally constrained analogues of glucagon-like peptide 1 (GLP-1) and exendin-4. Biochemistry 57, 4148–4154 (2018).
Watanabe, Y. et al. Structure–activity relationships of glucagon-like peptide-1 (7–36) amide: insulinotropic activities in perfused rat pancreases, and receptor binding and cyclic AMP production in RINm5F cells. J. Endocrinol. 140, 45–52 (1994).
Fisher, B. F., Hong, S. H. & Gellman, S. H. Helix propensities of amino acid residues via thioester exchange. J. Am. Chem. Soc. 139, 13292–13295 (2017).
Pace, C. N. & Scholtz, J. M. A helix propensity scale based on experimental studies of peptides and proteins. Biophys. J. 75, 422–427 (1998).
Adelhorst, K., Hedegaard, B. B., Knudsen, L. B. & Kirk, O. Structure-activity studies of glucagon-like peptide-1. J. Biol. Chem. 269, 6275–6278 (1994).
Göke, R. et al. Exendin-4 is a high potency agonist and truncated exendin-(9–39)-amide an antagonist at the glucagon-like peptide 1-(7–36)-amide receptor of insulin-secreting β-cells. J. Biol. Chem. 268, 19650–19655 (1993).
Anil, B., Song, B., Tang, Y. & Raleigh, D. P. Exploiting the right side of the Ramachandran plot: substitution of glycines by d -alanine can significantly increase protein stability. J. Am. Chem. Soc. 126, 13194–13195 (2004).
Fremaux, J. et al. Ureidopeptide GLP-1 analogues with prolonged activity in vivo via signal bias and altered receptor trafficking. Chem. Sci. 10, 9872–9879 (2019).
Hager, M. V., Johnson, L. M., Wootten, D., Sexton, P. M. & Gellman, S. H. β-Arrestin-biased agonists of the GLP-1 receptor from β-amino acid residue incorporation into GLP-1 analogues. J. Am. Chem. Soc. 138, 14970–14979 (2016).
Sang, P. et al. The activity of sulfono-γ-AApeptide helical foldamers that mimic GLP-1. Sci. Adv. 6, eaaz4988 (2020).
Johnson, L. M. & Gellman, S. H. α-Helix mimicry with α/β-peptides. Methods Enzymol 523, 407–429 (2013).
Fisher, B. F., Hong, S. H. & Gellman, S. H. Thermodynamic scale of β-amino acid residue propensities for an α-helix-like conformation. J. Am. Chem. Soc. 140, 9396–9399 (2018).
Underwood, C. R. et al. Crystal structure of glucagon-like peptide-1 in complex with the extracellular domain of the glucagon-like peptide-1 receptor. J. Biol. Chem. 285, 723–730 (2010).
Johnson, L. M. et al. A potent α/β-peptide analogue of GLP-1 with prolonged action in vivo. J. Am. Chem. Soc. 136, 12848–12851 (2014).
Cary, B. P., Hager, M. V. & Gellman, S. H. Impact of substitution registry on receptor-activation profiles of backbone-modified glucagon-like peptide-1 analogues. ChemBioChem 20, 2834–2840 (2019).
Mortenson, D. E. et al. Evaluation of β-amino acid replacements in protein loops: effects on conformational stability and structure. ChemBioChem 19, 604–612 (2018).
Craig, C. M. et al. Efficacy and pharmacokinetics of subcutaneous exendin (9–39) in patients with post-bariatric hypoglycaemia. Diabetes Obes. Metab. 20, 352–361 (2018).
Furness, S. G. B. et al. Ligand-dependent modulation of G protein conformation alters drug efficacy. Cell 167, 739–749 (2016).
Yang, D. et al. Structural determinants of binding the seven-transmembrane domain of the glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 291, 12991–13004 (2016).
Zhang, H. et al. Autocrine selection of a GLP-1R G-protein biased agonist with potent antidiabetic effects. Nat. Commun. 6, 8918 (2015).
Zhao, P. et al. Activation of the GLP-1 receptor by a non-peptidic agonist. Nature 577, 432–436 (2020).
Dods, R. L. & Donnelly, D. The peptide agonist-binding site of the glucagon-like peptide-1 (GLP-1) receptor based on site-directed mutagenesis and knowledge-based modelling. Biosci. Rep. 36, e00285 (2016).
Wootten, D. et al. Key interactions by conserved polar amino acids located at the transmembrane helical boundaries in Class B GPCRs modulate activation, effector specificity and biased signalling in the glucagon-like peptide-1 receptor. Biochem. Pharmacol. 118, 68–87 (2016).
Fairman, R., Anthony-Cahill, S. J. & DeGrado, W. F. The helix-forming propensity of d-alanine in a right-handed α-helix. J. Am. Chem. Soc. 114, 5458–5459 (1992).
Fesinmeyer, R. M., Peterson, E. S., Dyer, R. B. & Andersen, N. H. Studies of helix fraying and solvation using 13C′ isotopomers. Protein Sci. 14, 2324–2332 (2005).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Deganutti, G., Moro, S. & Reynolds, C. A. A supervised molecular dynamics approach to unbiased ligand–protein unbinding. J. Chem. Inf. Model. 60, 1804–1817 (2020).
Wootten, D. et al. The extracellular surface of the GLP-1 receptor is a molecular trigger for biased agonism. Cell 165, 1632–1643 (2016).
O’Connor, C. et al. NMR structure and dynamics of the agonist dynorphin peptide bound to the human κ opioid receptor. Proc. Natl Acad. Sci. USA 112, 11852–11857 (2015).
Bumbak, F. et al. Conformational changes in tyrosine 11 of neurotensin are required to activate the neurotensin receptor 1. ACS Pharmacol. Transl. Sci 3, 690–705 (2020).
Deganutti, G. et al. Dynamics of GLP-1R peptide agonist engagement are correlated with kinetics of G protein activation. Preprint at bioRxiv https://doi.org/10.1101/2021.03.10.434902 (2021).
Chorev, M. et al. Modifications of position 12 in a parathyroid hormone and parathyroid hormone-related protein: toward the design of highly potent antagonists. Biochemistry 29, 1580–1586 (1990).
Binkowski, B. F. et al. A luminescent biosensor with increased dynamic range for intracellular cAMP. ACS Chem. Biol. 6, 1193–1197 (2011).
Liang, Y.-L. et al. Dominant negative G proteins enhance formation and purification of agonist–GPCR–G protein complexes for structure determination. ACS Pharmacol. Transl. Sci. 1, 12–20 (2018).
Danev, R. et al. Routine sub-2.5 Å cryo-EM structure determination of GPCRs. Nat. Commun. 12, 4333 (2021).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Struct. Biol. 74, 519–530 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Liang, Y.-L. et al. Structure and dynamics of adrenomedullin receptors AM1 and AM2 reveal key mechanisms in the control of receptor phenotype by receptor activity-modifying proteins. ACS Pharmacol. Transl. Sci. 3, 263–284 (2020).
Doerr, S., Harvey, M. J., Noé, F. & De Fabritiis, G. HTMD: high-throughput molecular dynamics for molecular discovery. J. Chem. Theory Comput. 12, 1845–1852 (2016).
Acknowledgements
This work was supported by the National Institutes of Health (R01 GM056414, to S.H.G.). B.P.C. was supported in part by a graduate fellowship from the NSF (DGE-1747503) and by a Biotechnology Training Grant from NIGMS (T32 GM008349). R.D. was supported by a Takeda Science Foundation 2019 Medical Research Grant and Japan Science and Technology Agency PRESTO (18069571). P.M.S. and D.W. were supported by an ARC Centre Grant (IC200100052). P.M.S. was supported by the National Health and Medical Research Council of Australia (NHMRC) Program Grant (1150083) and Senior Principal Research Fellowship (1154434). D.W. was supported by NHMRC Project Grants (1126857 and 1184726) and a NHMRC Senior Research Fellowship (1155302). This study made use of the National Magnetic Resonance Facility at Madison, which is supported by NIH grants P41GM136463 and P41RR002301; equipment was purchased with funds from the University of Wisconsin-Madison, the NIH (P41GM103399, S10RR02781, S10RR08438, S10RR023438, S10RR025062, S10RR029220) and the NSF (DMB-8415048, OIA-9977486, BIR-9214394). We thank Promega (Madison, WI) for sharing plasmid DNA encoding NLuc.
Author information
Authors and Affiliations
Contributions
B.P.C. designed the project, synthesized peptides, generated the NLuc-GLP-1R construct and expressed and purified the protein complex. G.D. performed and analyzed MD simulations. B.P.C., P.Z. and T.T.T. conducted in vitro assays. B.P.C., S.J.P. and M.J.B. processed the cryo-EM data, built the model and performed refinement. S.J.P. and M.J.B. performed multivariate analysis, assisted in data interpretation and assisted in figure preparation. X.L. performed NMR measurements and analyzed spectra. R.D. prepared the cryo-EM samples and collected EM data. P.M.S., D.W. and S.H.G. supervised the project. B.P.C., P.M.S., D.W. and S.H.G. interpreted data, generated figures and wrote the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.H.G. is a cofounder of Longevity Biotech, Inc., which is pursuing biomedical applications for α/β-peptides. P.M.S. receives research funding from Laboratoires Servier in the area of GPCR drug discovery. The current study is 100% independent from all academic or commercial collaborations with industry.
Additional information
Peer review information Nature Chemical Biology thanks Yan Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 G protein conformation assay time-courses.
The ligand induced BRET is baseline subtracted. Agonist was added at time 2 min and GTP (30 µM) was added after the 12.1 minute timepoint. The values indicated by the colored keys indicate concentrations (log[peptide (M)]). Data points represent the mean of at least three independent experiments. n = 3, 7, 7, 4, 4, and 3 independent replicates for GLP-1, Ex4, Ex4-D-Ala, Ex4-R,R-X, Ex4-L-Ala, and Ex4-S,S-X, respectively. Error bars represent standard error.
Extended Data Fig. 2 Purification and characterization of GLP-1R/Ex4-D-Ala/DNGas/Gβ1/Gγ2/Nb35 complex.
(a) Size-Exclusion chromatogram of crude anti-FLAG elution. (b) Fluorescence-detected size-exclusion chromatograph of purified complex. (c) Western blot. The channel corresponding to anti-His6 antibody is depicted in blue. The channel corresponding to anti-FLAG antibody channel is depicted in green. The channel corresponding to anti-Gas antibody is depicted in red. (d) Coomassie-stained, reducing SDS-PAGE gel. The lanes labeled 1, 2, and 3 correspond to LMNG/CHS solubilized fraction, anti-FLAG column flow-through, and purified complex, respectively for both (c) and (d). (e) Representative 2D-classes from negative-stain, single-particle electron microscopy of GLP-1R/Ex4-D-Ala/DNGas/Gβ1/Gγ2/Nb35 complex. Purification and characterization of GLP-1R/Ex4-S,S-X/DNGas/Gβ1/Gγ2/Nb35 complex. (f) Size-Exclusion chromatogram of crude anti-FLAG elution. (g) Fluorescence-detected size-exclusion chromatograph of purified complex. (h) Western blot. The channel corresponding to anti-His6 antibody is depicted in blue. The channel corresponding to anti-FLAG antibody channel is depicted in green. The channel corresponding to anti-Gas antibody is depicted in red. (i) Coomassie-stained, reducing SDS-PAGE gel. The lanes labeled 1, 2, 3, and 4 correspond to LMNG/CHS solubilized fraction, anti-FLAG column flow-through, anti-FLAG elution, and purified complex, respectively for both (c) and (d). (j) Representative 2D-classes from negative-stain, single-particle electron microscopy of GLP-1R/Ex4-S,S-X/DNGas/Gβ1/Gγ2/Nb35 complex. N = 2 biological replicates for c, d, (representative data) and N = 1 for h, and i.
Extended Data Fig. 3
An overview of the cryo-EM data processing pipeline for the Ex4-D-Ala/GLP-1R/Gs complex.
Extended Data Fig. 4 CryoEM map reconstructions.
(a, e, j) Gold-standard Fourier Shell Correlation (FSC) curves for the consensus map (a), conformer 1 (e), and conformer 2 (j) showing overall nominal resolutions of 2.3 Å, 2.4 Å, and 2.5 Å, respectively. Black, green, blue and red curves indicate corrected, unmasked, masked, and phase-randomized maps, respectively. (b, f, k) Euler angle distribution histograms of the particles used in reconstructions for the consensus map (B), conformer 1 (f), and conformer 2 (k). (c-d,g-i,l-n) Local resolution estimates shown as colored heatmaps. High threshold maps with resolution-estimate heatmaps for the consensus map (c), conformer 1 (g), and conformer 2 (l). Low threshold maps with resolution-estimate heatmaps for the consensus map (d), conformer 1 (h), and conformer 2 (m). Receptor-focused refinment maps for conformer 1 (i) and conformer 2 (n).
Extended Data Fig. 5 Map to model figures for selected features of Conformer 1 (a) and Conformer 2 (b).
Residues are show in paratheses. a indicates the map threshold.
Extended Data Fig. 6 Analysis of Ex4-D-Ala bound to GLP-1R.
(a) A Ligplot+ v2.2 diagram of the N-terminal interacting residues of Ex-4-D-Ala bound to GLP-1R as observed in the conformer 1 model. Residues of the agonist are indicated by the chain (P) denotation and GLP-1R residues are indicated by the (R) denotation. Hydrophobic interactions are shown with solid red lines, ionic interactions are show with dotted red lines, and hydrogen bonding is indicated with dotted green lines. Hydrogen bonds are shown with a maximum distance of 3.35 A and other non-bonded contacts are shown with 3.90 A. A comparison of GLP-1R structures with ligand removed (and ECD removed in top views) for clarity. (b) Side view and (c) top view of GLP-1R bound to TT-OAD2, PF-06882961, and Ex-4-D-Ala’s two conformers shown in purple, yellow, blue and orange, respectively. (d) Side view and (e) top view of GLP-1R bound to Ex-P5, Chu-128 and Ex-4-D-Ala’s two conformers shown in cyan, brown, blue, and orange, respectively.
Extended Data Fig. 7 Receptor-focused refinement of conformer 2.
The receptor-focused density map of Ex4-D-Alaconf. 2 with the model of exendin-9-39-bound to the GLP-1R ECD (PDB: 3C5T) docked to the density. b, A close view showing density for selected side chains of the docked model shown in a. c, Comparison of the docked model from a,b overlaid with the Ex4-D-Alaconf. 1 model. d, A model of Ex4-D-Ala fitted (using ISOLDE) to the receptor-focused density map of Ex4-D-Alaconf. 2 with a helical n-terminus. e, A model of Ex4-D-Ala fitted (using ISOLDE) to the receptor-focused density map of Ex4-D-Alaconf. 2 with an extended n-terminus (orange), and a snapshot of the molecular dynamics simulations of Ex4-D-Ala with the receptor docked to the receptor-focused density map of Ex4-D-Alaconf. 2 showing an extended N-terminus (red).
Extended Data Fig. 8 Conformation of GLP-1 and analogs in solution.
(a-b) Far-UV Circular Dichroism peptide of peptides reconstituted in ultrapure water at 25 °C. All measurements were performed at 25 µM except GLP-1-L-Ala which was measured at a 14 µM. Inset graphs show a close-up of the region from 190-220 nm. (c-d) Far-UV Circular Dichroism peptide of peptides reconstituted in 30% (%v/v) of 2,2,2-trifluoroethanol (TFE) in ultrapure water at 25 µM concentration and 25 °C. (g) CαH chemical shift patterns for the N-terminal region of peptides. The residue number corresponds to number of residues from the N-terminus. ΔδCαH = δCαH(obs) - δCαH(RC), with δCαH(RC) values obtained from Wishart, et al. No value is shown for residue 1 because the N-terminal histidine residue is free, and no value is shown for position 4 due to the presence of unnatural substitutions.
Extended Data Fig. 9 Secondary structure analysis of Exendin-4 and analogs simulated in water.
(a) Left: Per-residue time course analysis of secondary structure during three simulations. Right: Per-residue averaged secondary structure observed. (b) The local effects of position-4 substitution on the fraction of helical secondary structure during simulations. (c) The local effects of position-4 substitution on the fraction of bend secondary structure during simulations.
Extended Data Fig. 10 Molecular dynamics simulations including the GLP-1R.
a, b, c. An outline of the MD simulations performed on the GLP-1R bound to Ex4-L-Ala (a), Ex4 (b) and Ex4-D-Ala (c). d, e, f. MD snapshots taken at frame 4000 (200 ns) of Ex4-L-Ala (a), Ex4 (c) and Ex4-D-Ala (e) bound to the GLP-1R from the selected simulated rebinding replicates. The GLP-1R is colored grey. Ex4-L-Ala, Ex4, and Ex4-D-Ala are colored green, white, and red, respectively. The Gαs C-terminal, h5 helix (residues 370-394) is colored yellow. g, h, i. Per-residue, secondary structure fractions for Ex4-L-Ala (g), Ex4 (h) and Ex4-D-Ala (i) bound to the GLP-1R averaged across five MD simulation replicates.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Tables 1–6, video descriptions, methods and instrumentation.
Supplementary Video 1
Partial unbinding and binding simulation of Ex4-l-Ala. The MD trajectory is the merge of one metadynamics replica with one SuMD replica and 300 ns of classic MD. Ex4-l-Ala is represented as orange ribbon (backbone) and sticks (side chains, position 4 as thick stick), and GLP-1R is represented as white ribbon. GLP-1R side chains within 3 Å of Ex4-l-Ala are shown as cyan sticks. Red dotted lines indicate hydrogen bonds. The starting conformation of the backbone of Ex4-l-Ala (conformer 1) is shown in transparent orange.
Supplementary Video 2
Partial unbinding and binding simulation of Ex4. The MD trajectory is the merge of one metadynamics replica with one SuMD replica and 300 ns of classic MD. Ex4 is represented as orange ribbon (backbone) and sticks (side chains), and GLP-1R is represented as white ribbon. GLP-1R side chains within 3 Å of Ex4 are shown as cyan sticks. The starting conformation of the backbone of Ex4 is shown in transparent orange.
Supplementary Video 3
Partial unbinding and binding of Ex4-d-Ala. Hydrophobic interactions between Phe 9 and Y1521.47 and L1411.36 anchor the peptide to GLP-1R in both a conformer 1-like structure (beginning of the video) and a conformer 2-like structure (end of the video) during the simulation. The MD trajectory is the merge of one metadynamics replica with one SuMD replica and 300 ns of classic MD. Ex4-d-Ala is represented as orange ribbon (backbone) and sticks (side chains, position 4 as thick stick), and GLP-1R is represented as white ribbon. GLP-1R side chains within 3 Å of Ex4-d-Ala are shown as cyan sticks. Red dotted lines indicate hydrogen bonds. A model of the backbone of Ex4-d-Ala from the cryo-EM maps of GLP-1R conformer 2 is shown in transparent purple for reference.
Supplementary Video 4
A video summarizing cryoSPARC 3D variability analysis for the first three principal components of the consensus refinement.
Supplementary Video 5
Morphs between models generated by roughly fitting the Ex4-d-Ala–GLP-1R–Gαs complex to extreme frames from cryoSPARC 3D variability analysis. Supplementary Videos 2, 3 and 4 correspond to principle components 1, 2 and 3, respectively. Supplementary Video 2 does not include morphs for the peptide or extracellular domain because one extreme frame from component 1 showed poorly resolved density in these regions.
Supplementary Video 6
Same as Supplementary Video 5.
Supplementary Video 7
Same as Supplementary Video 5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Cary, B.P., Deganutti, G., Zhao, P. et al. Structural and functional diversity among agonist-bound states of the GLP-1 receptor. Nat Chem Biol 18, 256–263 (2022). https://doi.org/10.1038/s41589-021-00945-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00945-w
This article is cited by
-
Cryo-electron microscopy for GPCR research and drug discovery in endocrinology and metabolism
Nature Reviews Endocrinology (2024)
-
Understanding VPAC receptor family peptide binding and selectivity
Nature Communications (2022)