Abstract
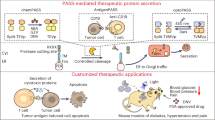
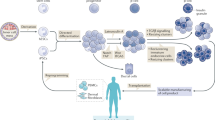
Inducer-triggered therapeutic protein expression from designer cells is a promising strategy for disease treatment. However, as most inducer systems harness transcriptional machineries, protein expression timeframes are unsuitable for many therapeutic applications. Here, we engineered a genetic code expansion-based therapeutic system, termed noncanonical amino acids (ncAAs)-triggered therapeutic switch (NATS), to achieve fast therapeutic protein expression in response to cognate ncAAs at the translational level. The NATS system showed response within 2 hours of triggering, whereas no signal was detected in a transcription-machinery-based system. Moreover, NATS system is compatible with transcriptional switches for multi-regulatory-layer control. Diabetic mice with microencapsulated cell implants harboring the NATS system could alleviate hyperglycemia within 90 min on oral delivery of ncAA. We also prepared ncAA-containing ‘cookies’ and achieved long-term glycemic control in diabetic mice implanted with NATS cells. Our proof-of-concept study demonstrates the use of NATS system for the design of next-generation cell-based therapies to achieve fast orally induced protein expression.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the main data supporting the study are provided within this article, its Supplementary Information and source data files. Source data are provided with this paper.
References
Saeedi, P. et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas. Diabetes Res. Clin. Pract. 157, 107843 (2019).
Ruder, W. C., Lu, T. & Collins, J. J. Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011).
Kitada, T., DiAndreth, B., Teague, B. & Weiss, R. Programming gene and engineered-cell therapies with synthetic biology. Science 359, eaad1067 (2018).
Scheller, L. & Fussenegger, M. From synthetic biology to human therapy: engineered mammalian cells. Curr. Opin. Biotechnol. 58, 108–116 (2019).
Chen, Z., Hu, Q. & Gu, Z. Leveraging engineering of cells for drug delivery. Acc. Chem. Res. 51, 668–677 (2018).
Kolar, K. & Weber, W. Synthetic biological approaches to optogenetically control cell signaling. Curr. Opin. Biotechnol. 47, 112–119 (2017).
P Teixeira, A. & Fussenegger, M. Engineering mammalian cells for disease diagnosis and treatment. Curr. Opin. Biotechnol. 55, 87–94 (2019).
Zaykov, A. N., Mayer, J. P. & DiMarchi, R. D. Pursuit of a perfect insulin. Nat. Rev. Drug Discov. 15, 425–439 (2016).
Krawczyk, K. et al. Electrogenetic cellular insulin release for real-time glycemic control in type 1 diabetic mice. Science 368, 993–1001 (2020).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Mandell, D. J. et al. Biocontainment of genetically modified organisms by synthetic protein design. Nature 518, 55–60 (2015).
Rovner, A. J. et al. Recoded organisms engineered to depend on synthetic amino acids. Nature 518, 89–93 (2015).
Ernst, R. J. et al. Genetic code expansion in the mouse brain. Nat. Chem. Biol. 12, 776–778 (2016).
Maywood, E. S. et al. Translational switching of Cry1 protein expression confers reversible control of circadian behavior in arrhythmic Cry-deficient mice. Proc. Natl Acad. Sci. USA 115, E12388–E12397 (2018).
Suzuki, T. et al. Switchable genome editing via genetic code expansion. Sci. Rep. 8, 10051 (2018).
Wang, F., Robbins, S., Guo, J., Shen, W. & Schultz, P. G. Genetic incorporation of unnatural amino acids into proteins in Mycobacterium tuberculosis. PLoS ONE 5, e9354 (2010).
Wang, N. et al. Construction of a live-attenuated HIV-1 vaccine through genetic code expansion. Angew. Chem. Int. Ed. 53, 4867–4871 (2014).
Si, L. et al. Generation of influenza A viruses as live but replication-incompetent virus vaccines. Science 354, 1170–1173 (2016).
Minaba, M. & Kato, Y. High-yield, zero-leakage expression system with a translational switch using site-specific unnatural amino acid incorporation. Appl. Environ. Microbiol. 80, 1718–1725 (2014).
Mukai, T., Lajoie, M. J., Englert, M. & Söll, D. Rewriting the genetic code. Annu. Rev. Microbiol. 71, 557–577 (2017).
Tang, H. et al. Proteomic identification of protein tyrosine phosphatase and substrate interactions in living mammalian cells by genetic encoding of irreversible enzyme inhibitors. J. Am. Chem. Soc. 140, 13253–13259 (2018).
Hu, L. et al. Thermophilic pyrrolysyl-tRNA synthetase mutants for enhanced mammalian genetic code expansion. ACS Synth. Biol. https://doi.org/10.1021/acssynbio.0c00257 (2020).
Qin, X. et al. An orthogonal tyrosyl-tRNA synthetase/tRNA pair from a thermophilic bacterium for an expanded eukaryotic genetic code. Biochemistry 59, 90–99 (2020).
Tannous, B. A. & Teng, J. Secreted blood reporters: insights and applications. Biotechnol. Adv. 29, 997–1003 (2011).
Gossen, M. et al. Transcriptional activation by tetracyclines in mammalian cells. Science 268, 1766–1769 (1995).
Volkwein, W., Maier, C., Krafczyk, R., Jung, K. & Lassak, J. A versatile toolbox for the control of protein levels using N ε -acetyl-l-lysine dependent amber suppression. ACS Synth. Biol. 6, 1892–1902 (2017).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Batty, K. T., Law, A. S. F., Stirling, V. & Moore, B. R. Pharmacodynamics of doxycycline in a murine malaria model. Antimicrob. Agents Chemother. 51, 4477–4479 (2007).
Ishiwata, K. et al. Preclinical and clinical evaluation of O-[11C]methyl-l-tyrosine for tumor imaging by positron emission tomography. Nucl. Med. Biol. 32, 253–262 (2005).
Farina, M., Alexander, J. F., Thekkedath, U., Ferrari, M. & Grattoni, A. Cell encapsulation: overcoming barriers in cell transplantation in diabetes and beyond. Adv. Drug Deliv. Rev. 139, 92–115 (2019).
Orive, G. et al. Cell encapsulation: technical and clinical advances. Trends Pharmacol. Sci. 36, 537–546 (2015).
Mathieu, C., Gillard, P. & Benhalima, K. Insulin analogues in type 1 diabetes mellitus: getting better all the time. Nat. Rev. Endocrinol. 13, 385–399 (2017).
Wu, K. K. & Huan, Y. Streptozotocin‐induced diabetic models in mice and rats. Curr. Protoc. Pharmacol. 40, 5.47.1–5.47.14 (2008).
Reaven, P. D. et al. Intensive glucose control in patients with type 2 diabetes - 15-year follow-up. N. Engl. J. Med. 380, 2215–2224 (2019).
Bai, P. et al. A fully human transgene switch to regulate therapeutic protein production by cooling sensation. Nat. Med. 25, 1266–1273 (2019).
Yin, J. et al. A green tea-triggered genetic control system for treating diabetes in mice and monkeys. Sci. Transl. Med. 11, eaav8826 (2019).
Wu, G. Important roles of dietary taurine, creatine, carnosine, anserine and 4-hydroxyproline in human nutrition and health. Amino Acids 52, 329–360 (2020).
Lean, M. E. J. Low-calorie diets in the management of type 2 diabetes mellitus. Nat. Rev. Endocrinol. 15, 251–252 (2019).
Bachmanov, A. A., Reed, D. R. & Beauchamp, G. K. Food intake, water intake, and drinking spout side preference of 28 mouse strains. Behav Genet. 32, 435–443 (2002).
Ellacott, K. L. J., Morton, G. J., Woods, S. C., Tso, P. & Schwartz, M. W. Assessment of feeding behavior in laboratory mice. Cell Metab. 12, 10–17 (2010).
Ashimova, A., Yegorov, S., Negmetzhanov, B. & Hortelano, G. Cell encapsulation within alginate microcapsules: immunological challenges and outlook. Front. Bioeng. Biotechnol. 7, 380 (2019).
Farah, S. et al. Long-term implant fibrosis prevention in rodents and non-human primates using crystallized drug formulations. Nat. Mater. 18, 892–904 (2019).
Arranz-Gibert, P., Patel, J. R. & Isaacs, F. J. The role of orthogonality in genetic code expansion. eLife 9, 58 (2019).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Wang, L. Genetically encoding new bioreactivity. New Biotechnol. 38, 16–25 (2017).
Kemmer, C. et al. Self-sufficient control of urate homeostasis in mice by a synthetic circuit. Nat. Biotechnol. 28, 355–360 (2010).
Saxena, P., Charpin-El Hamri, G., Folcher, M., Zulewski, H. & Fussenegger, M. Synthetic gene network restoring endogenous pituitary–thyroid feedback control in experimental Graves’ disease. Proc. Natl Acad. Sci. USA 113, 1244–1249 (2016).
Wang, H., Xie, M., Charpin-El Hamri, G., Ye, H. & Fussenegger, M. Treatment of chronic pain by designer cells controlled by spearmint aromatherapy. Nat. Biomed. Eng. 2, 114–123 (2018).
Grasso, K. T. et al. Structural robustness affects the engineerability of aminoacyl-tRNA synthetases for genetic code expansion. Biochemistry 60, 489–493 (2021).
Tharp, J. M. & Liu, W. R. in Noncanonical Amino Acids Vol. 1728 (ed. Lemke, E. A.) 147–154 (Springer, 2018).
Tom, R., Bisson, L. & Durocher, Y. Transfection of adherent HEK293-EBNA1 cells in a six-well plate with branched PEI for production of recombinant proteins. Cold Spring Harb. Protoc. 3, pdb.prot4978 (2008).
Schlatter, S., Rimann, M., Kelm, J. & Fussenegger, M. SAMY, a novel mammalian reporter gene derived from Bacillus stearothermophilus a-amylase. Gene 282, 19–31 (2002).
Acknowledgements
This work was financially supported by Beijing Natural Science Foundation (grant no. JQ20034), the National Natural Science Foundation of China (grant nos. 21922701, 91853111 and 21778005), the National Major Scientific and Technological Special Project for ‘Significant New Drugs Development’ (grant no. 2019ZX09739001) and Shenzhen Institute of Synthetic Biology Scientific Research Program (grant no. DWKF20190004) to T.L., the National Natural Science Foundation of China (grant no. 31971346 and 31861143016), the National Key R&D Program of China, Synthetic Biology Research (no. 2019YFA0904500) and the Science and Technology Commission of Shanghai Municipality (grant no. 18JC1411000) to H.Y. We also thank L. Zhong at the Medical and Health Analysis Center, Peking University, for help with mass spectrometry experiments and analysis.
Author information
Authors and Affiliations
Contributions
C.C., T.L., Y.H. and H.Y. designed the research. C.C., Y.H. and W.C. performed plasmid construction, cell culture and analytical assays. C.C., Y.H., W.C. and Y.L. performed hollow fiber-implanted mouse experiments. Y.S., Y.H. and W.C. performed pharmacokinetics experiments. C.C. and G.Y. performed cell line generation and microcapsule-implanted mouse experiments. C.C., G.Y. and Y.H. analyzed the data. C.C., Y.H., T.L. and H.Y. wrote the paper. H.Y. and Y.S. reviewed the paper. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Zhen Gu, Mingqi Xie and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 POI expression mediated by the NATS system composed of aaRS/tRNA pair.
a, The chemical structure of OmeY and BocK and used in this study. b, Exploring the impact of amber codon position on the expression of a POI (SEAP) in HEK293T cells. MbPylRS/tRNAPyl pair was used for BocK incorporation. SEAP levels in culture supernatants were measured 48 h after stimulation with BocK. The signal-to-noise ratio is indicated above the bars. Data are presented as the mean ± SD; n = 3 independent samples.
Extended Data Fig. 2 SEAP expression Ratio of the NATS or Tet-On systems.
a, Fold-change of SEAP expression at the indicated time points with respect to SEAP expression at t0 were profiled after stimulation with 1 mM OmeY or Dox. b, Fold-change of SEAP expression at the indicated time points with respect to SEAP expression at t0 were profiled after stimulation with 1 mM OmeY or Dox. Data are presented as the mean ± SD; n = 3 independent samples.
Extended Data Fig. 3 The transcription–translation combination switch-mediated protein expression.
a, Schematic showing the design of the transcription-translation combination switch. A ncAA could trigger translation of tetracycline-controlled transactivator (tTA) bearing an ectopic amber codon, then Dox could stop SEAP transcription by regulating the activity of the tTA. b, The translation-transcription combination switch consists of triple plasmids, encoding an OmeYRS/tRNA pair, tTA with an amber codon, and wild-type SEAP. c, Exploration of the impact on amber codon position on the expression of a POI (transactivator tTA) in HEK293T cells. OmeYRS/tRNA pair was used for OmeY incorporation. SEAP levels in culture supernatants were measured 48 h after OmeY or Dox treatment. Data are presented as the mean ± SD; n = 3 independent samples. The signal-to-noise ratio is indicated above the bars. Red rectangle represents the final construct used in subsequent studies. d, Constructs of translation-transcription combination switch using bacterial TyrRS/tRNA, and fluorescence micrographs of designer cells cultivated within or without OmeY. Each experiment was repeated three times independently with similar results. e, Constructs of translation-transcription combination switch using archaea PylRS/tRNA, and fluorescence micrographs of designer cells cultivated within or without BocK. Each experiment was repeated three times independently with similar results.
Extended Data Fig. 4 POI expression of NATS stable cell line.
a, Sleeping Beauty transposon-based ncAA system used to introduce an orthogonal aaRS/tRNA pair and a gene-of-interest based on an ectopic amber codon. ITR, Inverted repeats; P, Promoter; BleoR, Bleomycin resistance marker; PuroR, Puromycin resistance marker. b, Selected cell line clone No. 76 was incubated in the presence of or absence of 1mM OmeY. The signal-to-noise ratio is indicated above the bars. c, Long-term OmeY-dependent SEAP expression. SEAP levels in culture supernatants were measured every two days after stimulation. d, Selection of NATS stable cell lines using hMSC-TERT cell as carrier. The selected cell clones were profiled for their OmeY-inducible SEAP production. Red rectangle represents the final construct used in subsequent studies. e, Fluorescence micrographs of iPSC cells harboring NATS system cultivated within or without OmeY. Each experiment was repeated three times independently with similar results. f, OmeY-inducible SEAP expression in different cell types. The signal-to-noise ratio is indicated above the bars. Data are presented as the mean ± SD; n = 3 independent samples.
Extended Data Fig. 5 Biosafety of long-term intake of OmeY or BocK.
a, Body weights of mice feeding with ncAA cookies (equal to OmeY or BocK 200 mg/kg) or standard chow were recorded every 3 days during a month. Two-way ANOVA. b, Blood chemical indexes and complete blood count were determined in mice after one-month feeding with ncAA cookies, or standard chow. Data are presented as the mean ± SEM; n = 8 mice. ns, not significant. Exact P values are provided in Source Data files. One-way ANOVA. ALT, Alanine transaminase; AST, Aspartate transaminase; TP, Total protein; ALB, Albumin; CREA Creatinine; UA, Uric acid; GR, Granulocytes; HCT, Hematocrit; HGB, Hemoglobin; LY Lymphocytes; MCH, Mean cell hemoglobin; MCV, Mean cell volume; MO, Monocytes; MPV, Mean platelet volume; PCT, Plateletcrit; PDW, Platelet Distribution Width; PLT, Platelet count; RBC, red blood cells; WBC, white blood cells.
Extended Data Fig. 6 Implantation of microencapsulated NATS designer cells.
a, Schematic diagram of fast POI response to oral administration of ncAAs. Male C57BL/6 mice aged 8-10 weeks were implanted with microencapsulated engineered cells harboring the NATS system. Upon oral intake of OmeY, mice rapidly produce the POI based on translational activation. b, Representative micrographs of microencapsulated NATS designer cells. c, Oral OmeY-inducible SEAP expression in mice implanted with microencapsulated cells harboring the NATS system. After cell implantation, mice received OmeY (200mg/kg) or vesicle by o.g. administration 3 times per day. Data are presented as the mean ± SEM; n = 5 mice. Two-tailed Student’s t-test, ***P < 0.001. Exact P values are provided in Source Data files.
Extended Data Fig. 7 Implantation of hollow fibers harboring NATS designer cells.
a, Exploration of the effect on SEAP production by cell densities. Data are presented as the mean ± SD, n = 3 independent samples. b, Dose-dependent OmeY-inducible SEAP production in mice implanted with hollow fibers harboring NATS designer cells. After cell implantation, mice received OmeY or vesicle by o.g. administration 3 times per day. Serum SEAP level were measured 48 h after implantation. Data are presented as the mean ± SEM; n = 5 mice. Two-tailed Student’s t test, *P < 0.05, **P < 0.01, ***P < 0.001. Exact P values are provided in Source Data files.
Extended Data Fig. 8 Standard Chow did not affect OmeY absorption.
a, Schematic diagram of the NATS-mediated rapid insulin response to feeding on ncAA ‘cookies;’. Mice were implanted with microencapsulated engineered cells harboring the NATS system. Through intake of ncAA ‘cookies’, mice absorb ncAAs, which trigger a fast insulin response based on translational activation. b, Photos of homemade ncAA ‘cookies’ for feeding the T1DM mice. c, Pharmacokinetic analysis of serum OmeY concentrations in mice after oral gavage (o.g.) administration. Each point represents the mean ± SEM from 5 mice. Two-way ANOVA. d, Treatment schedules for long-term therapies in T1DM by OmeY oral gavage or ncAA ‘cookies’.
Supplementary information
Supplementary Information
Supplementary Table 1 and Notes.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Chen, C., Yu, G., Huang, Y. et al. Genetic-code-expanded cell-based therapy for treating diabetes in mice. Nat Chem Biol 18, 47–55 (2022). https://doi.org/10.1038/s41589-021-00899-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00899-z
This article is cited by
-
Tracking endogenous proteins based on RNA editing-mediated genetic code expansion
Nature Chemical Biology (2024)
-
How scientists are hacking the genetic code to give proteins new powers
Nature (2023)
-
Applications of synthetic biology in medical and pharmaceutical fields
Signal Transduction and Targeted Therapy (2023)
-
Unleashing the potential of noncanonical amino acid biosynthesis to create cells with precision tyrosine sulfation
Nature Communications (2022)
-
Treating diabetes with ‘cookies’
Nature Chemical Biology (2022)