Abstract
Inefficient homology-directed repair (HDR) constrains CRISPR–Cas9 genome editing in organisms that preferentially employ nonhomologous end joining (NHEJ) to fix DNA double-strand breaks (DSBs). Current strategies used to alleviate NHEJ proficiency involve NHEJ disruption. To confer precision editing without NHEJ disruption, we identified the shortcomings of the conventional CRISPR platforms and developed a CRISPR platform—lowered indel nuclease system enabling accurate repair (LINEAR)—which enhanced HDR rates (to 67–100%) compared to those in previous reports using conventional platforms in four NHEJ-proficient yeasts. With NHEJ preserved, we demonstrate its ability to survey genomic landscapes, identifying loci whose spatiotemporal genomic architectures yield favorable expression dynamics for heterologous pathways. We present a case study that deploys LINEAR precision editing and NHEJ-mediated random integration to rapidly engineer and optimize a microbial factory to produce (S)-norcoclaurine. Taken together, this work demonstrates how to leverage an antagonizing pair of DNA DSB repair pathways to expand the current collection of microbial factories.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are provided for the results presented in Main and Extended Data figures. Vectors containing LINEAR constructs for Y. lipolytica, K. marxianus, H. polymorpha and S. stipitis are available from Addgene (plasmid nos. 174837–174840). Raw reads and assembled genome sequences for the final engineered norcoclaurine-producing strains have been deposited in NCBI under BioProject ID PRJNA753835. Additional data are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
All code used for processing and assembly of genome sequencing reads (along with an accompanying method for how the code was implemented) is available on Zenodo: https://doi.org/10.5281/zenodo.5544230.
References
Hittinger, C. T. et al. Genomics and the making of yeast biodiversity. Curr. Opin. Genet. Dev. 35, 100–109 (2015).
Thorwall, S., Schwartz, C., Chartron, J. W. & Wheeldon, I. Stress-tolerant non-conventional microbes enable next-generation chemical biosynthesis. Nat. Chem. Biol. 16, 113–121 (2020).
Goncalves, F. A., Colen, G. & Takahashi, J. A. Yarrowia lipolytica and its multiple applications in the biotechnological industry. ScientificWorldJournal 2014, 476207 (2014).
Fonseca, G. G., Heinzle, E., Wittmann, C. & Gombert, A. K. The yeast Kluyveromyces marxianus and its biotechnological potential. Appl. Microbiol. Biotechnol. 79, 339–354 (2008).
Manfrao-Netto, J. H. C., Gomes, A. M. V. & Parachin, N. S. Advances in using Hansenula polymorpha as chassis for recombinant protein production. Front. Bioeng. Biotechnol. 7, 94 (2019).
Gao, M. et al. Innovating a nonconventional yeast platform for producing shikimate as the building block of high-value aromatics. ACS Synth. Biol. 6, 29–38 (2017).
Lobs, A. K., Schwartz, C. & Wheeldon, I. Genome and metabolic engineering in non-conventional yeasts: current advances and applications. Synth. Syst. Biotechnol. 2, 198–207 (2017).
Kuck, U. & Hoff, B. New tools for the genetic manipulation of filamentous fungi. Appl. Microbiol. Biotechnol. 86, 51–62 (2010).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair (Amst.) 7, 1765–1771 (2008).
Cao, M., Gao, M., Ploessl, D., Song, C. & Shao, Z. CRISPR-mediated genome editing and gene repression in Scheffersomyces stipitis. Biotechnol. J. 13, e1700598 (2018).
Sishc, B. J. & Davis, A. J. The role of the core non-homologous end joining factors in carcinogenesis and cancer. Cancers (Basel) 9, 81 (2017).
Ferguson, D. O. et al. The nonhomologous end-joining pathway of DNA repair is required for genomic stability and the suppression of translocations. Proc. Natl Acad. Sci. USA 97, 6630–6633 (2000).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Janssen, J. M., Chen, X., Liu, J. & Goncalves, M. The chromatin structure of CRISPR-Cas9 target DNA controls the balance between mutagenic and homology-directed gene-editing events. Mol. Ther. Nucleic Acids 16, 141–154 (2019).
Lieber, M. R. & Karanjawala, Z. E. Ageing, repetitive genomes and DNA damage. Nat. Rev. Mol. Cell Biol. 5, 69–75 (2004).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells. Cell Cycle 7, 2902–2906 (2008).
Tang, X. F. et al. Two Geminin homologs regulate DNA replication in silkworm, Bombyx mori. Cell Cycle 16, 830–840 (2017).
Kerns, S. L., Torke, S. J., Benjamin, J. M. & McGarry, T. J. Geminin prevents rereplication during xenopus development. J. Biol. Chem. 282, 5514–5521 (2007).
Quinn, L. M., Herr, A., McGarry, T. J. & Richardson, H. The Drosophila Geminin homolog: roles for Geminin in limiting DNA replication, in anaphase and in neurogenesis. Genes Dev. 15, 2741–2754 (2001).
McGarry, T. J. & Kirschner, M. W. Geminin, an inhibitor of DNA replication, is degraded during mitosis. Cell 93, 1043–1053 (1998).
Gutschner, T., Haemmerle, M., Genovese, G., Draetta, G. F. & Chin, L. Post-translational regulation of Cas9 during G1 enhances homology-directed repair. Cell Rep. 14, 1555–1566 (2016).
Slater, M. L., Sharrow, S. O. & Gart, J. J. Cell cycle of Saccharomyces cerevisiae in populations growing at different rates. Proc. Natl Acad. Sci. USA 74, 3850–3854 (1977).
Donaldson, A. D. et al. CLB5-dependent activation of late replication origins in S. cerevisiae. Mol. Cell 2, 173–182 (1998).
Collado, M., Blasco, M. A. & Serrano, M. Cellular senescence in cancer and aging. Cell 130, 223–233 (2007).
YMotoko Sasaki, Y. N. in Autophagy: Cancer, Other Pathologies, Inflammation, Immunity, Infection, and Aging (ed. Hayat, M. A.) pp. 293–303 (Academic Press, 2014).
Akhmetov, A. et al. Single-step precision genome editing in yeast using CRISPR-Cas9. Bio. Protoc. 8, e2765 (2018).
Lobs, A. K., Engel, R., Schwartz, C., Flores, A. & Wheeldon, I. CRISPR-Cas9-enabled genetic disruptions for understanding ethanol and ethyl acetate biosynthesis in Kluyveromyces marxianus. Biotechnol. Biofuels 10, 164 (2017).
Numamoto, M., Maekawa, H. & Kaneko, Y. Efficient genome editing by CRISPR/Cas9 with a tRNA-sgRNA fusion in the methylotrophic yeast Ogataea polymorpha. J. Biosci. Bioeng. 124, 487–492 (2017).
Juergens, H. et al. Contribution of Complex I NADH dehydrogenase to respiratory energy coupling in glucose-grown cultures of Ogataea parapolymorpha. Appl. Environ. Microbiol. 86, e00678–20 (2020).
Gao, S. et al. Multiplex gene editing of the Yarrowia lipolytica genome using the CRISPR-Cas9 system. J. Ind. Microbiol. Biotechnol. 43, 1085–1093 (2016).
Schwartz, C. M., Hussain, M. S., Blenner, M. & Wheeldon, I. Synthetic RNA polymerase III promoters facilitate high-efficiency CRISPR-Cas9-mediated genome editing in Yarrowia lipolytica. ACS Synth. Biol. 5, 356–359 (2016).
Vogel, C. & Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
Park, S. H. & Hahn, J. S. Development of an efficient cytosolic isobutanol production pathway in Saccharomyces cerevisiae by optimizing copy numbers and expression of the pathway genes based on the toxic effect of alpha-acetolactate. Sci. Rep. 9, 3996 (2019).
Li, N., Zeng, W., Xu, S. & Zhou, J. Toward fine-tuned metabolic networks in industrial microorganisms. Synth. Syst. Biotechnol. 5, 81–91 (2020).
Sato, H., Das, S., Singer, R. H. & Vera, M. Imaging of DNA and RNA in living eukaryotic cells to reveal spatiotemporal dynamics of gene expression. Annu. Rev. Biochem. 89, 159–187 (2020).
Chen, R., Yang, S., Zhang, L. & Zhou, Y. J. Advanced strategies for production of natural products in yeast. iScience 23, 100879 (2020).
Hagel, J. M. & Facchini, P. J. Benzylisoquinoline alkaloid metabolism: a century of discovery and a brave new world. Plant Cell Physiol. 54, 647–672 (2013).
DeLoache, W. C. et al. An enzyme-coupled biosensor enables (S)-reticuline production in yeast from glucose. Nat. Chem. Biol. 11, 465–471 (2015).
Liu, Q. et al. Rewiring carbon metabolism in yeast for high level production of aromatic chemicals. Nat. Commun. 10, 4976 (2019).
Zhao, Y. et al. Leveraging the Hermes transposon to accelerate the development of nonconventional yeast-based microbial cell factories. ACS Synth. Biol. 9, 1736–1752 (2020).
Jung, S. T., Lauchli, R. & Arnold, F. H. Cytochrome P450: taming a wild type enzyme. Curr. Opin. Biotechnol. 22, 809–817 (2011).
Veith, A. & Moorthy, B. Role of cytochrome P450s in the generation and metabolism of reactive oxygen species. Curr. Opin. Toxicol. 7, 44–51 (2018).
Hrycay, E. G. & Bandiera, S. M. Involvement of cytochrome P450 in reactive oxygen species formation and cancer. Adv. Pharmacol. 74, 35–84 (2015).
Bendich, A. J. The end of the circle for yeast mitochondrial DNA. Mol. Cell 39, 831–832 (2010).
Cao, M. et al. Centromeric DNA facilitates nonconventional yeast genetic engineering. ACS Synth. Biol. 6, 1545–1553 (2017).
Cao, M., Seetharam, A. S., Severin, A. J. & Shao, Z. Rapid isolation of centromeres from Scheffersomyces stipitis. ACS Synth. Biol. 6, 2028–2034 (2017).
Stanford, W. L., Cohn, J. B. & Cordes, S. P. Gene-trap mutagenesis: past, present and beyond. Nat. Rev. Genet. 2, 756–768 (2001).
Nguyen, V. Q., Co, C. & Li, J. J. Cyclin-dependent kinases prevent DNA re-replication through multiple mechanisms. Nature 411, 1068–1073 (2001).
Sopko, R. et al. Mapping pathways and phenotypes by systematic gene overexpression. Mol. Cell 21, 319–330 (2006).
Boeke, J. D., Trueheart, J., Natsoulis, G. & Fink, G. R. 5-Fluoroorotic acid as a selective agent in yeast molecular genetics. Methods Enzymol. 154, 164–175 (1987).
Shao, Z., Zhao, H. & Zhao, H. DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res. 37, e16 (2009).
Herricks, T. et al. One-cell doubling evaluation by living arrays of yeast, ODELAY! G3 (Bethesda) 7, 279–288 (2017).
Calvert, M. E., Lannigan, J. A. & Pemberton, L. F. Optimization of yeast cell cycle analysis and morphological characterization by multispectral imaging flow cytometry. Cytometry A 73, 825–833 (2008).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Laplaza, B. R. T. JoseM., Jin, Yong-Su & Jeffries, ThomasW. Sh ble and Cre adapted for functional genomics and metabolic engineering of Pichia stipitis. Enzyme Microb. Technol. 38, 741–747 (2006).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M. & Van Broeckhoven, C. NanoPack: visualizing and processing long-read sequencing data. Bioinformatics 34, 2666–2669 (2018).
Kolmogorov, M. et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat. Methods 17, 1103–1110 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Acknowledgements
This work was supported by the National Science Foundation Grant (no. 1716837 to D.P., Y.Z., M.C., S.G., C.L., M.S., S.C., A.S., L.H. and Z.S.), the NSF Graduate Research Fellowship Program (to D.P.) and Griswold Internship (to M.G.). We thank T. W. Jeffries (professor emeritus, University of Wisconsin-Madison and the Founder of Xylome Corporation) for sharing S. stipitis FLP-UC7 (ura3-3, NRRL Y21448) and the plasmid pJML545. We thank I. Wheeldon, University of California at Riverside, for sharing K. marxianus YS402 (CBC6556 Δura3); J. Dueber, University of California at Berkeley for sharing the S. cerevisiae plasmid containing TyrH* and DOD; and S. Yang, Chinese Academy of Science, for sharing the Y. lipolytica plasmids pCas1-yl and pCAS2-yl. We also thank S. M. Rigby for flow cytometry and L. Showman and K. Narayanaswamy at the W. M. Keck Metabolomics Research Laboratory of ISU for the detection of (S)-norcoclaurine. Finally, we thank T. Murtha at the ISU DNA facility for preparing gDNA samples for Nanopore sequencing.
Author information
Authors and Affiliations
Contributions
D.P. and Z.S. conceptualized the initial idea, designed the experiments, performed troubleshooting and wrote the manuscript. Y.Z. contributed to aberrant recombination characterization and hGem tag characterization in S. stipitis. M.C. contributed to the initial design of the hGem tag. S.G. contributed to Y. lipolytica PEX10 deletion. C.L. contributed to the construction of H. polymorpha DL1 Δleu2. M.S., S.C., and A.S. contributed to genome sequence analysis. M.G. and L.H. aided in cloning efforts and LINEAR CRISPR editing efficiency determination.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Zihe Liu, Eric Young and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
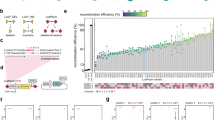
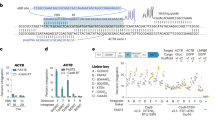
Extended Data Fig. 1 Literature survey to determine the most implemented CRISPR platform.
A compilation of 400 instances where studies deployed CRISPR for genome editing revealed that most researchers implement the conventional CRISPR plasmid and donor DNA Platform I. Studies were identified on PubMed using ‘Yeast CRISPR’ and ‘Mammal CRISPR’ in the search query.
Extended Data Fig. 2 The shortcomings associated with the conventional CRISPR platforms in S. stipitis.
a, The workflow for the conventional CRISPR plasmid and donor DNA platform (Platform I in Extended Data Fig. 1) deployed in S. stipitis, depicting various shortcomings our group has encountered that impeded obtaining correctly edited cells. b, The vector map of the consolidated ‘self-digesting’ plasmid, pCashGem-SD-Xyl2, highlighting the key design components (Platform IV in Extended Data Fig. 1). c, Transformation of UC7 with the pCashGem-SD-Xyl2 plasmid failed to address the shortcomings, with almost all the colonies displaying an episomal GFP expression profile.
Extended Data Fig. 3 Aberrant recombination characterization in S. stipitis.
a, Visual depiction of visibly intense GFP expression and subsequent loss of expression b, Vector map of the pGAOU plasmid, harboring the GAOU fragment flanked by SacI and XhoI restriction sites. The digested GAOU fragment was transformed into UC7. c, Ten strongly intense green colonies observed under the DR46B transilluminator. d, The GFP expression intensity of the ten variants was evaluated by flow cytometry after 36 h of cultivation. Numbers correspond to the mean GFP expression intensity. Among these ten variants, only #5, #7, and #9 exhibited genomic expression of GFP (sharp peaks). The gating methodology was summarized in Supplementary Information.
Extended Data Fig. 4 Verification of ‘pseudo’ plasmids formed by aberrant recombination events in S. stipitis.
a, Transformation of E. coli BW25141 with the five plasmids isolated from S. stipitis variants. b, Enzyme digestion verification of pGAOU-X vectors isolated from E. coli DH5α. Lane 1: M, GeneRuler 1 kb DNA Ladder. Lanes 2 and 3: pGAOU-1 isolates. Lanes 4 and 5: pGAOU-2 isolates. Lanes 6 and 7: pGAOU-3 isolates. Individual restriction digestions for each biological sample were performed once. c, Vector maps of the isolated pGAOU-X plasmids. d, Transformation of UC7 with pGAOU-2 and pGAOU-3. Plates were observed under a DR46B transilluminator.
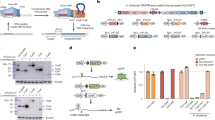
Extended Data Fig. 5 Design protocol for the LINEAR platform.
a, Cloning of individual LINEAR components. b, Assembly of LINEAR components into a cloning vector. Digestion and gel electrophoresis of the assembled vector yields the LINEAR fragment used in genome editing.
Extended Data Fig. 6 Comparison of the conventional CRISPR platforms versus LINEAR highlights growth deficiency associated with constitutive Cas9 expression.
All platforms utilized hGem tagged-Cas9. Plate pictures were taken at various timepoints post-transformation.
Extended Data Fig. 7 Assessing the contribution of the hGem tag on Xyl2:URA3 LINEAR editing efficiency with 500-bp homology arms.
a, Equimolar quantities of each LINEAR fragment were transformed to S. stipitis UC7. Plate pictures were taken at 48 h post-transformation. Ten colonies were selected at random (circled) for downstream analysis. b, The expected edit outcome on Xyl2 locus. c, PCR products obtained from the primer pair in b using the genomic DNA isolated from the colonies circled in a as templates. Individual PCR reactions for each biological sample were performed once. d, Sequencing of the amplicon associated with Cas9 #4 revealed an erroneous insertion of the donor template at the 5’ end of Xyl2.
Extended Data Fig. 8 Assessing the contribution of the hGem tag on Xyl2:URA3 LINEAR editing efficiency with 50-bp homology arms.
a, Equimolar quantities of each LINEAR fragment were transformed to S. stipitis UC7. Plate pictures were taken at 48 h post-transformation. Ten colonies selected at random (circled) for downstream analysis. Note that GFP cassette was placed in between sgRNA cassette and the donor DNA as a negative reporter to assess aberrant recombination. The colonies with strong GFP expression levels were false positives. b, The expected edit outcome on Xyl2 locus and the intact locus. c, PCR products obtained from the primer pair in b using the genomic DNA isolated from the colonies circled in a as templates. Individual PCR reactions for each biological sample were performed once. d, Sequencing of the amplicons associated with Cas9-hGem #10 and Cas9 #6 confirmed the presence of the intact wild type Xyl2. e, Sequencing of the amplicon associated with Cas9 #2 revealed an erroneous insertion of the donor template along with a portion of the upstream region of the LINEAR fragment.
Extended Data Fig. 9 Assessing the contribution of the hGem tag on Xyl2:GSAU LINEAR editing efficiency.
a, Equimolar quantities of each LINEAR fragment were transformed to S. stipitis UC7. Plate pictures were taken at 48 h post-transformation. Ten colonies that displayed genomic GFP expression as visualized on transilluminator selected (circled) for downstream analysis. Slightly different from the design used to achieve Xyl2:URA3 LINEAR editing with 50-bp homology arms, GFP cassette was included in the donor DNA as a positive reporter. b, PCR analysis of the 5’ end of the Xyl2 locus. Due to the large size of the inserted pathway, two reactions were designed to assess the nature of integration at each 500-bp homology arm. PCR products were obtained from the primer pair using the genomic DNA isolated from the colonies circled in a as templates. Individual PCR reactions for each biological sample were performed once. c, PCR analysis of the 3’ end of the Xyl2 locus. Individual PCR reactions for each biological sample were performed once. d, Sequencing of the amplicon associated with Cas9 #9 revealed an erroneous insertion of the downstream portion of the donor template at the 3’ end of Xyl2. Wild type stands for a ~100-bp sequence originating from the wild type Xyl2 locus between the upstream and the downstream homology arms.
Extended Data Fig. 10 Pathway map depicting the aromatic amino acid pathway and the de novo synthesis of (S)-norcoclaurine.
a, Glycolysis and the pentose phosphate pathway (PPP) channel carbon fluxes into the aromatic amino acid (AAA) pathway. Shikimate can serve as a reporter for assessing AAA pathway flux. Tkt1, Aro4*, Aro1, and Aro7* comprise the four-enzyme, upstream L-tyrosine precursor module. The red lines depict L-tyrosine feedback inhibition on the native Aro4 and Aro7 enzymes. b, (S)-betalamic acid, a product of the biosensing module of TyrH* and DOD, spontaneously condenses with endogenous amino acids to produce yellow pigmented betaxanthins. c, TyrH*, DODC, and NCS comprise the three-enzyme, downstream (S)-norcoclaurine module. Metabolite abbreviations: G6P, glucose-6-phosphate; PEP, phosphoenolpyruvate; E4P, erythrose 4-phosphate; DAHP, 3-deoxy-D-arabino-heptulosonate 7-phosphate; DHQ, 3-dehydroquinic acid; DHS, 3-dehydroshikimate; S3P, shikmate-3-phosphate; EPSP, 5-enolpyruvylshikimate-3-phosphate; L-DOPA, L-3,4-dihydroxyphenylalanine; 4-HPAA, 4-hydroxyphenylacetaldehyde. Enzyme abbreviations: Tkt1, transketolase; Aro4*, DAHP synthase feedback inhibition-resistant mutant (Aro4K220L); Aro1, pentafunctional Aro1 polypeptide; Aro1*, Aro1 mutant (Aro1D900A) with the shikimate kinase domain inactivated, enabling shikimate accumulation; Aro7*, chorismate mutase feedback inhibition-resistant mutant (Aro7G139S); TyrH*, tyrosine hydroxylase mutant (TyrHW13L, F309L) from Berberis vulgaris with low L-DOPA oxidase activity; DOD, L-DOPA dioxygenase from Mirabilis jalapa; DODC, L-DOPA decarboxylase from Pseudomonas putida; NCS, norcoclaurine synthase from Papaver somniferum.
Supplementary information
Supplementary Information
Supplementary Discussions 1–4, Figs. 1–22 and Tables 1–5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Compilation of references.
Source Data Extended Data Fig. 7
Unprocessed gel.
Source Data Extended Data Fig. 8
Unprocessed gel.
Source Data Extended Data Fig. 9
Unprocessed gel.
Rights and permissions
About this article
Cite this article
Ploessl, D., Zhao, Y., Cao, M. et al. A repackaged CRISPR platform increases homology-directed repair for yeast engineering. Nat Chem Biol 18, 38–46 (2022). https://doi.org/10.1038/s41589-021-00893-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00893-5
This article is cited by
-
Fusing an exonuclease with Cas9 enhances homologous recombination in Pichia pastoris
Microbial Cell Factories (2022)