Abstract
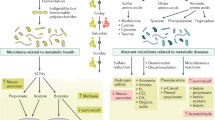
Human-associated microorganisms play a vital role in human health, and microbial imbalance has been linked to a wide range of disease states. In this Review, we explore recent efforts to progress from correlative studies that identify microorganisms associated with human disease to experiments that establish causal relationships between microbial products and host phenotypes. We propose that successful efforts to uncover phenotypes often follow a chain of evidence that proceeds from (1) association studies; to (2) observations in germ-free animals and antibiotic-treated animals and humans; to (3) fecal microbiota transplants (FMTs); to (4) identification of strains; and then (5) molecules that elicit a phenotype. Using this experimental ‘funnel’ as our guide, we explore how the microbiota contributes to metabolic disorders and hypertension, infections, and neurological conditions. We discuss the potential to use FMTs and microbiota-inspired therapies to treat human disease as well as the limitations of these approaches.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Nicolas, G. R. & Chang, P. V. Deciphering the chemical lexicon of host–gut microbiota interactions. Trends Pharmacol. Sci. 40, 430–445 (2019).
Luczynski, P. et al. Growing up in a bubble: using germ-free animals to assess the influence of the gut microbiota on brain and behavior. Int. J. Neuropsychopharmacol. 19, pyw020 (2016).
Kennedy, E. A., King, K. Y. & Baldridge, M. T. Mouse microbiota models: comparing germ-free mice and antibiotics treatment as tools for modifying gut bacteria. Front. Physiol. 9, 1534 (2018).
Bramante, C. T., Lee, C. J. & Gudzune, K. A. Treatment of obesity in patients with diabetes. Diabetes Spectr. 30, 237–243 (2017).
Schnurr, T. M. et al. Obesity, unfavourable lifestyle and genetic risk of type 2 diabetes: a case-cohort study. Diabetologia 63, 1324–1332 (2020).
Jiang, S. Z., Lu, W., Zong, X. F., Ruan, H. Y. & Liu, Y. Obesity and hypertension. Exp. Ther. Med. 12, 2395–2399 (2016).
Grigorescu, I. & Dumitrascu, D. L. Implication of gut microbiota in diabetes mellitus and obesity. Acta Endocrinol. 12, 206–214 (2016).
Castaner, O. et al. The gut microbiome profile in obesity: a systematic review. Int. J. Endocrinol. 2018, 4095789 (2018).
Dao, M. C. et al. Akkermansia muciniphila abundance is lower in severe obesity, but its increased level after bariatric surgery is not associated with metabolic health improvement. Am. J. Physiol. Endocrinol. Metab. 317, E446–E459 (2019).
Dao, M. C. et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: relationship with gut microbiome richness and ecology. Gut 65, 426–436 (2016).
Yan, Q. et al. Alterations of the gut microbiome in hypertension. Front. Cell Infect. Microbiol. 7, 381 (2017).
Li, J. et al. Gut microbiota dysbiosis contributes to the development of hypertension. Microbiome 5, 14 (2017).
Liu, J. et al. Correlation analysis of intestinal flora with hypertension. Exp. Ther. Med. 16, 2325–2330 (2018).
Scott, F. I. et al. Administration of antibiotics to children before age 2 years increases risk for childhood obesity. Gastroenterology 151, 120–129 (2016).
Hwang, I. et al. Alteration of gut microbiota by vancomycin and bacitracin improves insulin resistance via glucagon-like peptide 1 in diet-induced obesity. FASEB J. 29, 2397–2411 (2015).
Miao, Z. et al. Antibiotics can cause weight loss by impairing gut microbiota in mice and the potent benefits of lactobacilli. Biosci. Biotechnol. Biochem. 84, 411–420 (2020).
Hooper, L. V. Bacterial contributions to mammalian gut development. Trends Microbiol. 12, 129–134 (2004).
Davis, C. D. The gut microbiome and its role in obesity. Nutr. Today 51, 167–174 (2016).
Honour, J. W., Borriello, S. P., Ganten, U. & Honour, P. Antibiotics attenuate experimental hypertension in rats. J. Endocrinol. 105, 347–350 (1985).
Sanada, T. J. et al. Gut microbiota modification suppresses the development of pulmonary arterial hypertension in an SU5416/hypoxia rat model. Pulm. Circ. 10, 2045894020929147 (2020).
Galla, S. et al. Disparate effects of antibiotics on hypertension. Physiol. Genomics 50, 837–845 (2018).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214 (2013).
Wang, S. et al. Gut microbiota mediates the anti-obesity effect of calorie restriction in mice. Sci. Rep. 8, 13037 (2018).
Lai, Z. L. et al. Fecal microbiota transplantation confers beneficial metabolic effects of diet and exercise on diet-induced obese mice. Sci. Rep. 8, 15625 (2018).
de Groot, P. et al. Donor metabolic characteristics drive effects of faecal microbiota transplantation on recipient insulin sensitivity, energy expenditure and intestinal transit time. Gut 69, 502–512 (2020).
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23, 850–858 (2017).
Wang, H. et al. Promising treatment for type 2 diabetes: fecal microbiota transplantation reverses insulin resistance and impaired islets. Front. Cell Infect. Microbiol. 9, 455 (2019).
Kootte, R. S. et al. Improvement of insulin sensitivity after lean donor feces in metabolic syndrome is driven by baseline intestinal microbiota composition. Cell Metab. 26, 611–619 (2017).
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–6 (2012).
Zhang, Z. et al. Impact of fecal microbiota transplantation on obesity and metabolic syndrome—a systematic review. Nutrients 11, 2291 (2019).
Durgan, D. J. et al. Role of the gut microbiome in obstructive sleep apnea-induced hypertension. Hypertension 67, 469–474 (2016).
Adnan, S. et al. Alterations in the gut microbiota can elicit hypertension in rats. Physiol. Genomics 49, 96–104 (2017).
Ridlon, J. M., Kang, D.-J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 47, 241–259 (2006).
Broeders, E. P. et al. The bile acid chenodeoxycholic acid increases human brown adipose tissue activity. Cell Metab. 22, 418–426 (2015).
Kars, M. et al. Tauroursodeoxycholic acid may improve liver and muscle but not adipose tissue insulin sensitivity in obese men and women. Diabetes 59, 1899–1905 (2010).
Zhang, H. M. et al. Beneficial effect of farnesoid X receptor activation on metabolism in a diabetic rat model. Mol. Med. Rep. 13, 2135–2142 (2016).
Sun, L. et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat. Med. 24, 1919–1929 (2018).
Schittenhelm, B. et al. Role of FXR in beta-cells of lean and obese mice. Endocrinology 156, 1263–1271 (2015).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Liu, Y. et al. Gut microbiome fermentation determines the efficacy of exercise for diabetes prevention. Cell Metab. 31, 77–91 (2020).
La Rosa, S. L. et al. The human gut Firmicute Roseburia intestinalis is a primary degrader of dietary β-mannans. Nat. Commun. 10, 905 (2019).
Chambers, E. S. et al. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 64, 1744–1754 (2015).
van der Hee, B. & Wells, J. M. Microbial regulation of host physiology by short-chain fatty acids. Trends Microbiol. https://doi.org/10.1016/j.tim.2021.02.001 (2021).
Kim, K. N., Yao, Y. & Ju, S. Y. Short chain fatty acids and fecal microbiota abundance in humans with obesity: a systematic review and meta-analysis. Nutrients 11, 2512 (2019).
Muller, M. et al. Circulating but not faecal short-chain fatty acids are related to insulin sensitivity, lipolysis and GLP-1 concentrations in humans. Sci. Rep. 9, 12515 (2019).
Canfora, E. E., Jocken, J. W. & Blaak, E. E. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat. Rev. Endocrinol. 11, 577–591 (2015).
den Besten, G. et al. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 54, 2325–2340 (2013).
Pluznick, J. L. Microbial short-chain fatty acids and blood pressure regulation. Curr. Hypertension Rep. 19, 25 (2017).
Oyama, J.-I. & Node, K. Gut microbiota and hypertension. Hypertension Res. 42, 741–743 (2019).
Latif, S. A., Pardo, H. A., Hardy, M. P. & Morris, D. J. Endogenous selective inhibitors of 11β-hydroxysteroid dehydrogenase isoforms 1 and 2 of adrenal origin. Mol. Cell. Endocrinol. 243, 43–50 (2005).
Feighner, S. D. & Hylemon, P. B. Characterization of a corticosteroid 21-dehydroxylase from the intestinal anaerobic bacterium, Eubacterium lentum. J. Lipid Res. 21, 585–593 (1980).
Kumar, A., Ellermann, M. & Sperandio, V. Taming the beast: interplay between gut small molecules and enteric pathogens. Infect. Immun. 87, 277 (2019).
Cameron, E. A. & Sperandio, V. Frenemies: signaling and nutritional integration in pathogen–microbiota–host interactions. Cell Host Microbe 18, 275–284 (2015).
Manfredo Vieira, S. et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 359, 1156–1161 (2018).
Aykut, B. et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature 574, 264–267 (2019).
Wortelboer, K., Nieuwdorp, M. & Herrema, H. Fecal microbiota transplantation beyond Clostridioides difficile infections. EBioMedicine 44, 716–729 (2019).
Willing, B. P., Vacharaksa, A., Croxen, M., Thanachayanont, T. & Finlay, B. B. Altering host resistance to infections through microbial transplantation. PLoS ONE 6, e26988 (2011).
Buffie, C. G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015).
Vuong, H. E., Yano, J. M., Fung, T. C. & Hsiao, E. Y. The microbiome and host behavior. Annu. Rev. Neurosci. 40, 21–49 (2017).
Scheperjans, F. et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov. Disord. 30, 350–358 (2015).
Keshavarzian, A. et al. Colonic bacterial composition in Parkinson’s disease. Mov. Disord. 30, 1351–1360 (2015).
Peng, A. et al. Altered composition of the gut microbiome in patients with drug-resistant epilepsy. Epilepsy Res. 147, 102–107 (2018).
Xie, G. et al. Ketogenic diet poses a significant effect on imbalanced gut microbiota in infants with refractory epilepsy. World J. Gastroenterol. 23, 6164–6171 (2017).
Naseribafrouei, A. et al. Correlation between the human fecal microbiota and depression. Neurogastroenterol. Motil. 26, 1155–1162 (2014).
Jiang, H. et al. Altered fecal microbiota composition in patients with major depressive disorder. Brain Behav. Immun. 48, 186–194 (2015).
Cekanaviciute, E. et al. Gut bacteria from multiple sclerosis patients modulate human T cells and exacerbate symptoms in mouse models. Proc. Natl Acad. Sci. USA 114, 10713–10718 (2017).
Berer, K. et al. Gut microbiota from multiple sclerosis patients enables spontaneous autoimmune encephalomyelitis in mice. Proc. Natl Acad. Sci. USA 114, 10719–10724 (2017).
De Angelis, M. et al. Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS ONE 8, e76993 (2013).
Kang, D.-W. et al. Reduced incidence of Prevotella and other fermenters in intestinal microflora of autistic children. PLoS ONE 8, e68322 (2013).
Sharon, G. et al. Human gut microbiota from autism spectrum disorder promote behavioral symptoms in mice. Cell 177, 1600–1618 (2019).
Sampson, T. R. et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson’s disease. Cell 167, 1469–1480 (2016).
Berer, K. et al. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature 479, 538–541 (2011).
Kim, S. et al. Maternal gut bacteria promote neurodevelopmental abnormalities in mouse offspring. Nature 549, 528–532 (2017).
Blacher, E. et al. Potential roles of gut microbiome and metabolites in modulating ALS in mice. Nature 572, 474–480 (2019).
Olson, C. A. et al. The gut microbiota mediates the anti-seizure effects of the ketogenic diet. Cell 173, 1728–1741 (2018).
Chu, C. et al. The microbiota regulate neuronal function and fear extinction learning. Nature 574, 543–548 (2019).
Makkawi, S., Camara-Lemarroy, C. & Metz, L. Fecal microbiota transplantation associated with 10 years of stability in a patient with SPMS. Neurol. Neuroimmunol. Neuroinflamm. 5, e459 (2018).
Kang, D.-W. et al. Microbiota transfer therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: an open-label study. Microbiome 5, 10 (2017).
He, Z. et al. Fecal microbiota transplantation cured epilepsy in a case with Crohn’s disease: the first report. World J. Gastroenterol. 23, 3565–3568 (2017).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019).
Strandwitz, P. et al. GABA-modulating bacteria of the human gut microbiota. Nat. Microbiol. 4, 396–403 (2019).
Luscher, B., Shen, Q. & Sahir, N. The GABAergic deficit hypothesis of major depressive disorder. Mol. Psychiatry 16, 383–406 (2011).
Devlin, A. S. et al. Modulation of a circulating uremic solute via rational genetic manipulation of the gut microbiota. Cell Host Microbe 20, 709–715 (2016).
Yu, E. W. et al. Fecal microbiota transplantation for the improvement of metabolism in obesity: the FMT-TRIM double-blind placebo-controlled pilot trial. PLoS Med. 17, e1003051 (2020).
Young, M. T., Phelan, M. J. & Nguyen, N. T. A decade analysis of trends and outcomes of male vs female patients who underwent bariatric surgery. J. Am. Coll. Surg. 222, 226–231 (2016).
DeFilipp, Z. et al. Drug-resistant E. coli bacteremia transmitted by fecal microbiota transplant. N. Engl. J. Med. 381, 2043–2050 (2019).
Depommier, C. et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: a proof-of-concept exploratory study. Nat. Med. 25, 1096–1103 (2019).
Chiang, J. Y. L. & Ferrell, J. M. Bile acids as metabolic regulators and nutrient sensors. Annu. Rev. Nutr. 39, 175–200 (2019).
Bedford, A. & Gong, J. Implications of butyrate and its derivatives for gut health and animal production. Anim. Nutr. 4, 151–159 (2018).
Sato, F. T. et al. Tributyrin attenuates metabolic and inflammatory changes associated with obesity through a GPR109A-dependent mechanism. Cells 9, 2007 (2020).
Nguyen, T. D., Prykhodko, O., Hållenius, F. F. & Nyman, M. Monobutyrin reduces liver cholesterol and improves intestinal barrier function in rats fed high-fat diets. Nutrients 11, 308 (2019).
Yao, L. et al. A selective gut bacterial bile salt hydrolase alters host metabolism. eLife 7, e37182 (2018).
Bai, L. et al. Engineered butyrate-producing bacteria prevents high fat diet-induced obesity in mice. Microbe Cell Fact. 19, 94–13 (2020).
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 5, e9085 (2010).
Kan, H., Zhao, F., Zhang, X. X., Ren, H. & Gao, S. Correlations of gut microbial community shift with hepatic damage and growth inhibition of Carassius auratus induced by pentachlorophenol exposure. Environ. Sci. Technol. 49, 11894–11902 (2015).
Haro, C. et al. Intestinal microbiota is influenced by gender and body mass index. PLoS ONE 11, e0154090 (2016).
Patil, D. P. et al. Molecular analysis of gut microbiota in obesity among Indian individuals. J. Biosci. 37, 647–657 (2012).
Mullish, B. H. & Williams, H. R. Clostridium difficile infection and antibiotic-associated diarrhoea. Clin. Med. 18, 237–241 (2018).
Zheng, P. et al. Gut microbiome remodeling induces depressive-like behaviors through a pathway mediated by the host’s metabolism. Mol. Psychiatry 21, 786 (2016).
Acknowledgements
This work was supported by National Institutes of Health grants R35 GM128618 and R01 DK126855 (A.S.D.). S.N.C. acknowledges an American Heart Association Postdoctoral Fellowship. M.D.M. acknowledges an NSF Graduate Research Fellowship (DGE1745303). Figures created with BioRender.com.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
A.S.D. is an ad hoc consultant for Takeda Pharmaceuticals and Axial Therapeutics. The other authors have declared no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Andrew Gewirtz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chaudhari, S.N., McCurry, M.D. & Devlin, A.S. Chains of evidence from correlations to causal molecules in microbiome-linked diseases. Nat Chem Biol 17, 1046–1056 (2021). https://doi.org/10.1038/s41589-021-00861-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00861-z