Abstract
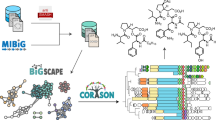
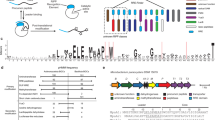
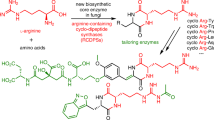
Multidomain enzymes orchestrate two or more catalytic activities to carry out metabolic transformations with increased control and speed. Here, we report the design and development of a genome-mining approach for targeted discovery of biochemical transformations through the analysis of co-occurring enzyme domains (CO-ED) in a single protein. CO-ED was designed to identify unannotated multifunctional enzymes for functional characterization and discovery based on the premise that linked enzyme domains have evolved to function collaboratively. Guided by CO-ED, we targeted an unannotated predicted ThiF–nitroreductase di-domain enzyme found in more than 50 proteobacteria. Through heterologous expression and biochemical reconstitution, we discovered a series of natural products containing the rare oxazolone heterocycle and characterized their biosynthesis. Notably, we identified the di-domain enzyme as an oxazolone synthetase, validating CO-ED-guided genome mining as a methodology with potential broad utility for both the discovery of unusual enzymatic transformations and the functional annotation of multidomain enzymes.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated during this study are included in this article and its Supplementary Information. CO-ED networks used to generate Fig. 2 and several Supplementary Figs. showing CO-ED networks are available in Supplementary Data 1. The curated set of enzyme domains for CO-ED can be found as the node table of the ‘all_of_uniprot’ network in Supplementary Data 1, as well as at https://github.com/tderond/CO-ED/blob/master/pfamID_to_name_desc_longdesc.tsv. This same set of domains is also the default setting for the web tool. NMR spectra of newly reported structures are available in the Supplementary Note. Tandem MS spectra of newly reported structures are available in the Supplementary Note and were also deposited to the GNPS spectral library at the URLs shown in the Supplementary Note. The following bioinformatic databases were employed in this study: BRENDA (https://www.brenda-enzymes.org/), MIBiG (https://mibig.secondarymetabolites.org/), UniProt (https://www.uniprot.org/) and Pfam-A (http://pfam.xfam.org). Source data are provided with this paper.
Code availability
Jupyter notebooks containing Python code for CO-ED analysis and for generating the statistics shown in Supplementary Figs. 1 and 18 are publicly available at https://github.com/tderond/CO-ED.
References
Gu, C., Kim, G. B., Kim, W. J., Kim, H. U. & Lee, S. Y. Current status and applications of genome-scale metabolic models. Genome Biol. 20, 121 (2019).
Bornscheuer, U. T. The fourth wave of biocatalysis is approaching. Philos. Trans. A Math. Phys. Eng. Sci. 376, 20170063 (2018).
Gerlt, J. A. et al. The enzyme function initiative. Biochemistry 50, 9950–9962 (2011).
Hanson, A. D., Pribat, A., Waller, J. C. & de Crécy-Lagard, V. Unknown proteins and orphan enzymes: the missing half of the engineering parts list—and how to find it. Biochem. J. 425, 1–11 (2009).
Scott, T. A. & Piel, J. The hidden enzymology of bacterial natural product biosynthesis. Nat. Rev. Chem. 3, 404–425 (2019).
Ellens, K. W. et al. Confronting the catalytic dark matter encoded by sequenced genomes. Nucleic Acids Res. 45, 11495–11514 (2017).
Medema, M. H., de Rond, T. & Moore, B. S. Mining genomes to illuminate the specialized chemistry of life. Nat. Rev. Genet. https://doi.org/10.1038/s41576-021-00363-7 (2021).
Michael, A. J. Evolution of biosynthetic diversity. Biochem. J. 474, 2277–2299 (2017).
Hagel, J. M. & Facchini, P. J. Tying the knot: occurrence and possible significance of gene fusions in plant metabolism and beyond. J. Exp. Bot. 68, 4029–4043 (2017).
Bashton, M. & Chothia, C. The generation of new protein functions by the combination of domains. Structure 15, 85–99 (2007).
Weissman, K. J. The structural biology of biosynthetic megaenzymes. Nat. Chem. Biol. 11, 660–670 (2015).
Winzer, T. et al. Plant science. Morphinan biosynthesis in opium poppy requires a P450–oxidoreductase fusion protein. Science 349, 309–312 (2015).
Ng, T. L., Rohac, R., Mitchell, A. J., Boal, A. K. & Balskus, E. P. An N-nitrosating metalloenzyme constructs the pharmacophore of streptozotocin. Nature 566, 94–99 (2019).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Jeske, L., Placzek, S., Schomburg, I., Chang, A. & Schomburg, D. BRENDA in 2019: a European ELIXIR core data resource. Nucleic Acids Res. 47, D542–D549 (2019).
Kautsar, S. A. et al. MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 48, D454–D458 (2020).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Ishino, F., Mitsui, K., Tamaki, S. & Matsuhashi, M. Dual enzyme activities of cell wall peptidoglycan synthesis, peptidoglycan transglycosylase and penicillin-sensitive transpeptidase, in purified preparations of Escherichia coli penicillin-binding protein 1A. Biochem. Biophys. Res. Commun. 97, 287–293 (1980).
Goodlove, P. E., Cunningham, P. R., Parker, J. & Clark, D. P. Cloning and sequence analysis of the fermentative alcohol-dehydrogenase-encoding gene of Escherichia coli. Gene 85, 209–214 (1989).
Williams, G. J., Breazeale, S. D., Raetz, C. R. H. & Naismith, J. H. Structure and function of both domains of ArnA, a dual function decarboxylase and a formyltransferase, involved in 4-amino-4-deoxy-l-arabinose biosynthesis. J. Biol. Chem. 280, 23000–23008 (2005).
Rusnak, F., Sakaitani, M., Drueckhammer, D., Reichert, J. & Walsh, C. T. Biosynthesis of the Escherichia coli siderophore enterobactin: sequence of the entF gene, expression and purification of EntF, and analysis of covalent phosphopantetheine. Biochemistry 30, 2916–2927 (1991).
De Rond, T. et al. Oxidative cyclization of prodigiosin by an alkylglycerol monooxygenase-like enzyme. Nat. Chem. Biol. 13, 1155–1157 (2017).
Agarwal, V. et al. Biosynthesis of polybrominated aromatic organic compounds by marine bacteria. Nat. Chem. Biol. 10, 640–647 (2014).
Ross, A. C., Gulland, L. E. S., Dorrestein, P. C. & Moore, B. S. Targeted capture and heterologous expression of the Pseudoalteromonas alterochromide gene cluster in Escherichia coli represents a promising natural product exploratory platform. ACS Synth. Biol. 4, 414–420 (2015).
Herz, S., Eberhardt, S. & Bacher, A. Biosynthesis of riboflavin in plants. The ribA gene of Arabidopsis thaliana specifies a bifunctional GTP cyclohydrolase II/3,4-dihydroxy-2-butanone 4-phosphate synthase. Phytochemistry 53, 723–731 (2000).
Schulman, B. A. & Harper, J. W. Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nat. Rev. Mol. Cell Biol. 10, 319–331 (2009).
Miyauchi, K., Kimura, S. & Suzuki, T. A cyclic form of N6-threonylcarbamoyladenosine as a widely distributed tRNA hypermodification. Nat. Chem. Biol. 9, 105–111 (2013).
Regni, C. A. et al. How the MccB bacterial ancestor of ubiquitin E1 initiates biosynthesis of the microcin C7 antibiotic. EMBO J. 28, 1953–1964 (2009).
Xi, J., Ge, Y., Kinsland, C., McLafferty, F. W. & Begley, T. P. Biosynthesis of the thiazole moiety of thiamin in Escherichia coli: identification of an acyldisulfide-linked protein–protein conjugate that is functionally analogous to the ubiquitin/E1 complex. Proc. Natl Acad. Sci. USA 98, 8513–8518 (2001).
Godert, A. M., Jin, M., McLafferty, F. W. & Begley, T. P. Biosynthesis of the thioquinolobactin siderophore: an interesting variation on sulfur transfer. J. Bacteriol. 189, 2941–2944 (2007).
Akiva, E., Copp, J. N., Tokuriki, N. & Babbitt, P. C. Evolutionary and molecular foundations of multiple contemporary functions of the nitroreductase superfamily. Proc. Natl Acad. Sci. USA 114, E9549–E9558 (2017).
Mondal, S., Raja, K., Schweizer, U. & Mugesh, G. Chemistry and biology in the biosynthesis and action of thyroid hormones. Angew. Chem. Int. Ed. Engl. 55, 7606–7630 (2016).
Schneider, T. L., Shen, B. & Walsh, C. T. Oxidase domains in epothilone and bleomycin biosynthesis: thiazoline to thiazole oxidation during chain elongation. Biochemistry 42, 9722–9730 (2003).
Taga, M. E., Larsen, N. A., Howard-Jones, A. R., Walsh, C. T. & Walker, G. C. BluB cannibalizes flavin to form the lower ligand of vitamin B12. Nature 446, 449–453 (2007).
Bashiri, G. et al. A revised biosynthetic pathway for the cofactor F420 in prokaryotes. Nat. Commun. 10, 1558 (2019).
Gondry, M. et al. Cyclic dipeptide oxidase from Streptomyces noursei. Isolation, purification and partial characterization of a novel, amino acyl α,β-dehydrogenase. Eur. J. Biochem. 268, 1712–1721 (2001).
Zenno, S., Saigo, K., Kanoh, H. & Inouye, S. Identification of the gene encoding the major NAD(P)H-flavin oxidoreductase of the bioluminescent bacterium Vibrio fischeri ATCC 7744. J. Bacteriol. 176, 3536–3543 (1994).
Guella, G., N’Diaye, I., Fofana, M. & Mancini, I. Isolation, synthesis and photochemical properties of almazolone, a new indole alkaloid from a red alga of Senegal. Tetrahedron 62, 1165–1170 (2006).
Seyedsayamdost, M. R. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc. Natl Acad. Sci. USA 111, 7266–7271 (2014).
Walsh, C. & Wencewicz, T. Antibiotics: Challenges, Mechanisms, Opportunities https://doi.org/10.1128/9781555819316 (American Society for Microbiology, 2016).
Hon, J. et al. EnzymeMiner: automated mining of soluble enzymes with diverse structures, catalytic properties and stabilities. Nucleic Acids Res. 48, W104–W109 (2020).
Gerlt, J. A. Genomic enzymology: web tools for leveraging protein family sequence–function space and genome context to discover novel functions. Biochemistry 56, 4293–4308 (2017).
Wuchty, S. & Almaas, E. Evolutionary cores of domain co-occurrence networks. BMC Evol. Biol. 5, 24 (2005).
Barrera, A., Alastruey-Izquierdo, A., Martín, M. J., Cuesta, I. & Vizcaíno, J. A. Analysis of the protein domain and domain architecture content in fungi and its application in the search of new antifungal targets. PLoS Comput. Biol. 10, e1003733 (2014).
Suhre, K. Inference of gene function based on gene fusion events: the Rosetta-stone method. In Methods in Molecular Biology 396, 31–41 (2007).
Promponas, V. J., Ouzounis, C. A. & Iliopoulos, I. Experimental evidence validating the computational inference of functional associations from gene fusion events: a critical survey. Brief. Bioinform. 15, 443–454 (2014).
Alborzi, S. Z., Devignes, M.-D. & Ritchie, D. W. ECDomainMiner: discovering hidden associations between enzyme commission numbers and Pfam domains. BMC Bioinformatics 18, 107 (2017).
De Castro, P. P., Carpanez, A. G. & Amarante, G. W. Azlactone reaction developments. Chem. Eur. J. 22, 10294–10318 (2016).
Kenney, G. E. et al. The biosynthesis of methanobactin. Science 359, 1411–1416 (2018).
Haft, D. H., Paulsen, I. T., Ward, N. & Selengut, J. D. Exopolysaccharide-associated protein sorting in environmental organisms: the PEP-CTERM/EpsH system. Application of a novel phylogenetic profiling heuristic. BMC Biol. 4, 29 (2006).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Xie, B.-B. et al. Genome sequence of the cycloprodigiosin-producing bacterial strain Pseudoalteromonas rubra ATCC 29570(T). J. Bacteriol. 194, 1637–1638 (2012).
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002).
Udwary, D. W. et al. Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica. Proc. Natl Acad. Sci. USA 104, 10376–10381 (2007).
Paulsen, I. T. et al. Complete genome sequence of the plant commensal Pseudomonas fluorescens Pf-5. Nat. Biotechnol. 23, 873–878 (2005).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Acknowledgements
We thank our UCSD colleagues J. Li, V. Shende, T. Fallon and B. Duggan for helpful discussions. This work was supported by National Institutes of Health awards F32GM129960 to T.d.R. and R01GM085770 to B.S.M., as well as an American Society for Pharmacognosy Undergraduate Research Award and a UC San Diego ‘Eureka’ Undergraduate Research Scholarship to J.E.A.
Author information
Authors and Affiliations
Contributions
T.d.R. and B.S.M. designed research; T.d.R. and J.E.A. performed research; T.d.R. analyzed data; T.d.R. and B.S.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks A. James Link, Maude Pupin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Survey of transformations known to be catalyzed by the ThiF and nitroreductase domains, and properties of organisms harboring a ThiF-nitroreductase di-domain enzyme identified by CO-ED.
a, OxzB exhibits homology to both the ThiF and nitroreductase enzyme families. Shown is a selection of characterized members of these enzyme families along with the transformations they catalyze, along with midpoint-rooted gene trees showing the relationships between each other and OxzB. ThiF family enzymes are known to catalyze ATP-dependent carboxylate activating reactions, while nitroreductase family enzymes catalyze a variety of redox reactions. Inferred phylogenies were generated from protein sequences using neighbor joining on the MAFFT web server51 and midpoint-rooted. Scale bars designate 1 substitution per site. b,c, Phylogenetic distribution (b) and habitat (c) of the host organisms harboring genes encoding OxzB proteins represented in the Uniprot database. Organisms with unknown habitats are not included. d, Genomic context of oxzB genes as determined using the Enzyme Function Initiative Genome Neighborhood Tool42. Most oxzB homologs are accompanied by oxzA, which codes for an N-acyltransferase. The arrow labeled ‘?’ represents genes unrelated to oxzA or oxzB. Organisms with unknown oxzB genomic context (for example, at the edge of a contig) are not included.
Extended Data Fig. 2 Induction of oxazolone production in P. rubra and C. chukchiensis by various antibiotics.
Bacteria were grown as a lawn on Marine Agar 2216 with a drop of antibiotic. A consistent amount of biomass adjacent to the zone of inhibition was harvested, extracted and analyzed by HPLC. Dots indicate summed peak areas between wavelengths of 300 nm and 400 nm. Three biological replicates were analyzed for each condition.
Supplementary information
Supplementary Information
Supplementary Figs. 1–19, Tables 1 and 2 and Note
Source data
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1b–d.
Source Data Extended Data Fig. 2
Statistical source data.
Rights and permissions
About this article
Cite this article
de Rond, T., Asay, J.E. & Moore, B.S. Co-occurrence of enzyme domains guides the discovery of an oxazolone synthetase. Nat Chem Biol 17, 794–799 (2021). https://doi.org/10.1038/s41589-021-00808-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00808-4
This article is cited by
-
Non-enzymatic reactions in biogenesis of fungal natural products
Journal of Natural Medicines (2024)
-
Strategies to access biosynthetic novelty in bacterial genomes for drug discovery
Nature Reviews Drug Discovery (2022)
-
Mining genomes to illuminate the specialized chemistry of life
Nature Reviews Genetics (2021)