Abstract
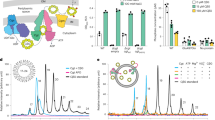
In ovoid-shaped, Gram-positive bacteria, MapZ guides FtsZ-ring positioning at cell equators. The cell wall of the ovococcus Streptococcus mutans contains peptidoglycan decorated with serotype c carbohydrates (SCCs). In the present study, we identify the major cell separation autolysin AtlA as an SCC-binding protein. AtlA binding to SCC is attenuated by the glycerol phosphate (GroP) modification. Using fluorescently labeled AtlA constructs, we mapped SCC distribution on the streptococcal surface, revealing enrichment of GroP-deficient immature SCCs at the cell poles and equators. The immature SCCs co-localize with MapZ at the equatorial rings throughout the cell cycle. In GroP-deficient mutants, AtlA is mislocalized, resulting in dysregulated cellular autolysis. These mutants display morphological abnormalities associated with MapZ mislocalization, leading to FtsZ-ring misplacement. Altogether, our data support a model in which maturation of a cell wall polysaccharide provides the molecular cues for the recruitment of cell division machinery, ensuring proper daughter cell separation and FtsZ-ring positioning.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are provided with this paper. All other data generated during this study are included in the article and supplementary information files.
References
Brown, S., Santa Maria, J. P. Jr. & Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 67, 313–336 (2013).
Higgins, M. L. & Shockman, G. D. Model for cell wall growth of Streptococcus faecalis. J. Bacteriol. 101, 643–648 (1970).
Land, A. D. & Winkler, M. E. The requirement for pneumococcal MreC and MreD is relieved by inactivation of the gene encoding PBP1a. J. Bacteriol. 193, 4166–4179 (2011).
Sham, L. T., Tsui, H. C., Land, A. D., Barendt, S. M. & Winkler, M. E. Recent advances in pneumococcal peptidoglycan biosynthesis suggest new vaccine and antimicrobial targets. Curr. Opin. Microbiol. 15, 194–203 (2012).
Wheeler, R., Mesnage, S., Boneca, I. G., Hobbs, J. K. & Foster, S. J. Super-resolution microscopy reveals cell wall dynamics and peptidoglycan architecture in ovococcal bacteria. Mol. Microbiol. 82, 1096–1109 (2011).
Massidda, O., Novakova, L. & Vollmer, W. From models to pathogens: how much have we learned about Streptococcus pneumoniae cell division? Environ. Microbiol. 15, 3133–3157 (2013).
van Raaphorst, R., Kjos, M. & Veening, J. W. Chromosome segregation drives division site selection in Streptococcus pneumoniae. Proc. Natl Acad. Sci. USA 114, E5959–E5968 (2017).
Fleurie, A. et al. MapZ marks the division sites and positions FtsZ rings in Streptococcus pneumoniae. Nature 516, 259–262 (2014).
Holeckova, N. et al. LocZ is a new cell division protein involved in proper septum placement in Streptococcus pneumoniae. mBio 6, e01700–e01714 (2014).
Perez, A. J. et al. Movement dynamics of divisome proteins and PBP2x:FtsW in cells of Streptococcus pneumoniae. Proc. Natl Acad. Sci. USA 116, 3211–3220 (2019).
Manuse, S. et al. Structure–function analysis of the extracellular domain of the pneumococcal cell division site positioning protein MapZ. Nat. Commun. 7, 12071 (2016).
Yamamoto, H., Miyake, Y., Hisaoka, M., Kurosawa, S. & Sekiguchi, J. The major and minor wall teichoic acids prevent the sidewall localization of vegetative dl-endopeptidase LytF in Bacillus subtilis. Mol. Microbiol. 70, 297–310 (2008).
Schirner, K., Marles-Wright, J., Lewis, R. J. & Errington, J. Distinct and essential morphogenic functions for wall- and lipo-teichoic acids in Bacillus subtilis. EMBO J. 28, 830–842 (2009).
Schlag, M. et al. Role of staphylococcal wall teichoic acid in targeting the major autolysin Atl. Mol. Microbiol. 75, 864–873 (2010).
Lunderberg, J. M., Liszewski Zilla, M., Missiakas, D. & Schneewind, O. Bacillus anthracis tagO Is required for vegetative growth and secondary cell wall polysaccharide synthesis. J. Bacteriol. 197, 3511–3520 (2015).
Bonnet, J. et al. Nascent teichoic acids insertion into the cell wall directs the localization and activity of the major pneumococcal autolysin LytA. Cell Surf. 2, 24–37 (2018).
Mistou, M. Y., Sutcliffe, I. C. & van Sorge, N. M. Bacterial glycobiology: rhamnose-containing cell wall polysaccharides in Gram-positive bacteria. FEMS Microbiol. Rev. 40, 464–479 (2016).
St Michael, F. et al. Investigating the candidacy of the serotype specific rhamnan polysaccharide based glycoconjugates to prevent disease caused by the dental pathogen Streptococcus mutans. Glycoconj. J. 35, 53–64 (2018).
Coligan, J. E., Kindt, T. J. & Krause, R. M. Structure of the streptococcal groups A, A-variant and C carbohydrates. Immunochemistry 15, 755–760 (1978).
McCarty, M. Variation in the group-specific carbohydrate of group A streptococci. II. Studies on the chemical basis for serological specificity of the carbohydrates. J. Exp. Med. 104, 629–643 (1956).
Edgar, R. J. et al. Discovery of glycerol phosphate modification on streptococcal rhamnose polysaccharides. Nat. Chem. Biol. 15, 463–471 (2019).
Rush, J. S. et al. The molecular mechanism of N-acetylglucosamine side-chain attachment to the Lancefield group A carbohydrate in Streptococcus pyogenes. J. Biol. Chem. 292, 19441–19457 (2017).
Brown, T. A. Jr. et al. A hypothetical protein of Streptococcus mutans is critical for biofilm formation. Infect. Immun. 73, 3147–3151 (2005).
Shibata, Y., Kawada, M., Nakano, Y., Toyoshima, K. & Yamashita, Y. Identification and characterization of an autolysin-encoding gene of Streptococcus mutans. Infect. Immun. 73, 3512–3520 (2005).
Ahn, S. J. & Burne, R. A. The atlA operon of Streptococcus mutans: role in autolysin maturation and cell surface biogenesis. J. Bacteriol. 188, 6877–6888 (2006).
Yoshimura, G. et al. Identification and molecular characterization of an N-acetylmuraminidase, Aml, involved in Streptococcus mutans cell separation. Microbiol Immunol. 50, 729–742 (2006).
Yamashita, Y. et al. A novel gene required for rhamnose-glucose polysaccharide synthesis in Streptococcus mutans. J. Bacteriol. 181, 6556–6559 (1999).
Shibata, Y., Yamashita, Y., Ozaki, K., Nakano, Y. & Koga, T. Expression and characterization of streptococcal rgp genes required for rhamnan synthesis in Escherichia coli. Infect. Immun. 70, 2891–2898 (2002).
Zorzoli, A. et al. Group A, B, C, and G Streptococcus Lancefield antigen biosynthesis is initiated by a conserved alpha-d-GlcNAc-beta-1,4-l-rhamnosyltransferase. J. Biol. Chem. 294, 15237–15256 (2019).
Li, Y. et al. MapZ forms a stable ring structure that acts as a nanotrack for FtsZ treadmilling in Streptococcus mutans. ACS Nano. 12, 6137–6146 (2018).
Neuhaus, F. C. & Baddiley, J. A continuum of anionic charge: structures and functions of d-alanyl-teichoic acids in Gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 686–723 (2003).
Yu, Y. et al. Systematic hydrogen-bond manipulations to establish polysaccharide structure–property correlations. Angew. Chem. Int. Ed. Engl. 58, 13127–13132 (2019).
Yu, Y. & Delbianco, M. Conformational studies of oligosaccharides. Chemistry 26, 9814–9825 (2020).
Merzlyak, E. M. et al. Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nat. Methods 4, 555–557 (2007).
Pedelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Catalao, M. J., Figueiredo, J., Henriques, M. X., Gomes, J. P. & Filipe, S. R. Optimization of fluorescent tools for cell biology studies in Gram-positive bacteria. PLoS ONE 9, e113796 (2014).
Rosenow, M. A., Huffman, H. A., Phail, M. E. & Wachter, R. M. The crystal structure of the Y66L variant of green fluorescent protein supports a cyclization–oxidation–dehydration mechanism for chromophore maturation. Biochemistry 43, 4464–4472 (2004).
Dufour, D. & Levesque, C. M. Cell death of Streptococcus mutans induced by a quorum-sensing peptide occurs via a conserved streptococcal autolysin. J. Bacteriol. 195, 105–114 (2013).
Templeton, D. W., Quinn, M., Van Wychen, S., Hyman, D. & Laurens, L. M. Separation and quantification of microalgal carbohydrates. J. Chromatogr. A 1270, 225–234 (2012).
Kojima, N., Araki, Y. & Ito, E. Structural studies on the linkage unit of ribitol teichoic acid of Lactobacillus plantarum. Eur. J. Biochem. 148, 29–34 (1985).
Lehrman, M. A. & Gao, N. Alternative and sources of reagents and supplies of fluorophore-assisted carbohydrate electrophoresis (FACE). Glycobiology 13, 1G–3G (2003).
Chaturvedi, S. K., Ma, J., Brown, P. H., Zhao, H. & Schuck, P. Measuring macromolecular size distributions and interactions at high concentrations by sedimentation velocity. Nat. Commun. 9, 4415 (2018).
Schuck, P. Size–distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Brautigam, C. A. Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol. 562, 109–133 (2015).
Stafford, W. F. & Braswell, E. H. Sedimentation velocity, multi-speed method for analyzing polydisperse solutions. Biophys. Chem. 108, 273–279 (2004).
Acknowledgements
We thank S.-J. Ahn (University of Florida) for the kind gift of anti-AtlA antibodies, J. Abranches (University of Florida) for providing S. mutans serotype e, f and k strains; J. F. Timoney (University of Kentucky) and J. Huebner (von Hauner Children’s Hospital, LMU) for providing S. equi and E. faecalis, respectively; J. M. Bosken and E. D. Hall (University of Kentucky) for the use of the Thermo Fisher Scientific GC–MS instrument and C. Velez-Ortega (University of Kentucky) for access to a Leica SP8 confocal microscope. This work was supported by National Institutes of Health (NIH) grants (nos. R01 DE028916 from the National Institute of Dental and Craniofacial Research (NIDCR) and R01 AI143690 from the National Institute of Allergy and Infectious Diseases to N.K., R01 GM094363 from the National Institute of General Medical Sciences to A.B.H. and R01 DC014658 from the NIDCD to G.I.F.), Tenovus Scotland large research grant (no. T17/17) and University of Dundee Wellcome Fund (grant no. 105606/Z/14/Z) to S.A.C. and H.C.D., and the Wellcome and Royal Society grant (no. 109357/Z/15/Z) to H.C.D.. SEM was performed at the Electron Microscopy Center, which belongs to the National Science Foundation NNCI Kentucky Multiscale Manufacturing and Nano Integration Node, supported by ECCS-1542174. Carbohydrate composition analysis at the Complex Carbohydrate Research Center was supported by the Chemical Sciences, Geosciences and Biosciences Division, Office of Basic Energy Sciences, US Department of Energy grant (no. DE-FG02-93ER20097). The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
S.Z., C.T.C., J.S.R., S.A.C., A.E.Y., A.B.H., N.M.v.S., H.C.D., G.I.F., K.V.K. and N.K. designed the experiments. S.Z., C.T.C., J.S.R., C.W.K., S.A.C., A.E.Y., H.C.D., K.V.K. and N.K. performed functional and biochemical experiments. S.Z. and G.I.F. performed microscopy analysis. N.K., K.V.K. and N.M.v.S. constructed plasmids and isolated mutants. S.Z., C.T.C., J.S.R., S.A.C., A.E.Y., A.B.H., H.C.D., K.V.K. and N.K. analyzed the data. N.K. wrote the manuscript with contributions from all authors. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Rut Carballido-Lopez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Modifications of SCC control S. mutans cell aggregation and morphology.
a, Sedimentation phenotype of S. mutans strains after overnight growth in THY broth. b, Sedimentation phenotype of sacculi purified from S. mutans strains. Sacculi were resuspended in PBS (OD600 of 8), and allowed to sediment for 72 hours at 4 °C. c, DIC images of S. mutans strains taken from mid-log growth phase. Blue arrows denote small round cells. Scale bar is 1 µm. Representative images from at least three independent experiments are shown in a, b and c.
Extended Data Fig. 2 Fluorescent microscopy of S. mutans intact cells and sacculi labeled by AtlABSP-GFP.
a, Binding of AtlABSP-GFP to the intact WT, ΔsccH, ΔsccH:psccH, ΔsccN and ΔsccN:psccN bacterial cells. b, Binding of AtlABSP-GFP to the sacculi purified from WT, ΔsccH and ΔsccH:psccH. In a and b, yellow and blue arrows show polar and equatorial sites labeled by AtlABSP-GFP, respectively. Scale bar is 1 µm in a and b. The experiments depicted in a and b were performed independently three times and yielded the same results.
Extended Data Fig. 3 Gating Strategy for Flow Cytometry (Fig. 4b).
In this sample gating, E. coli expressing polyrhamnose on the cell surface and its parental strain (E. coli without polyrhamnose) were gated based on the presence of fluorescent signal (AtlABSP-GFP). Bacterial gating occurred at the GFP/SCC density plot omitting signals derived from bacteria stained with GFP alone (GFP). For flow cytometric analysis, at least 10,000 events were collected. Experiments were performed independently three times and yielded the same results. Dot plots of representative results are shown.
Extended Data Fig. 4 S. mutans WT produces SCCs with different degrees of GroP modification.
a, Ion exchange chromatography of SCCs purified from ΔsccH. b, Ion exchange chromatography of SCCs purified from WT S. mutans. SCC material was loaded onto Toyopearl DEAE-650M and eluted with a NaCl gradient (0-0.5 M). Fractions were analyzed for Rha and Glc contents by anthrone assay and total phosphate (Pi) content by malachite green assay following digestion with perchloric acid. c, Composition analysis of minor and major fractions from b. Fractions were pooled, concentrated, and desalted by spin column and analyzed by GC-MS to determine the Rha and Glc concentrations. The concentration of Glc is presented relative to the Rha concentration. Data are means ± S.D., n = 3 biologically independent experiments. P values were calculated by a two-tailed t-test. d, SDS-PAGE analysis of ANDS-labeled SCCs extracted from S. mutans mutants. In a, b and d, the experiments were performed at least three times and yielded the same results. Data from one representative experiment are shown.
Supplementary information
Supplementary Information
Supplementary Figs. 1–18, Supplementary Tables 1–6, Supplementary References.
Source data
Source Data Fig. 3b
Uncropped SDS–PAGE gel corresponding to Fig 3b.
Source Data Extended Data Fig. 4d
Uncropped SDS–PAGE gel corresponding to Extended Data Fig 4d.
Rights and permissions
About this article
Cite this article
Zamakhaeva, S., Chaton, C.T., Rush, J.S. et al. Modification of cell wall polysaccharide guides cell division in Streptococcus mutans. Nat Chem Biol 17, 878–887 (2021). https://doi.org/10.1038/s41589-021-00803-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00803-9
This article is cited by
-
PplD is a de-N-acetylase of the cell wall linkage unit of streptococcal rhamnopolysaccharides
Nature Communications (2022)
-
Undermodification cues division
Nature Chemical Biology (2021)