Abstract
Cell death can be executed by regulated apoptotic and nonapoptotic pathways, including the iron-dependent process of ferroptosis. Small molecules are essential tools for studying the regulation of cell death. Using time-lapse imaging and a library of 1,833 bioactive compounds, we assembled a large compendium of kinetic cell death modulatory profiles for inducers of apoptosis and ferroptosis. From this dataset we identify dozens of ferroptosis suppressors, including numerous compounds that appear to act via cryptic off-target antioxidant or iron chelating activities. We show that the FDA-approved drug bazedoxifene acts as a potent radical trapping antioxidant inhibitor of ferroptosis both in vitro and in vivo. ATP-competitive mechanistic target of rapamycin (mTOR) inhibitors, by contrast, are on-target ferroptosis inhibitors. Further investigation revealed both mTOR-dependent and mTOR-independent mechanisms that link amino acid metabolism to ferroptosis sensitivity. These results highlight kinetic modulatory profiling as a useful tool to investigate cell death regulation.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Uncropped western blots are available in the Supplementary Information. Source data are provided with this paper.
Code availability
The manuscript does not report any custom code.
References
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Stockwell, B. R., Jiang, X. & Gu, W. Emerging mechanisms and disease relevance of ferroptosis. Trends Cell Biol. 30, 478–490 (2020).
Conrad, M. & Pratt, D. A. The chemical basis of ferroptosis. Nat. Chem. Biol. 15, 1137–1147 (2019).
Li, Y., Qian, L. & Yuan, J. Small molecule probes for cellular death machines. Curr. Opin. Chem. Biol. 39, 74–82 (2017).
Smith, C. E. et al. Non-steroidal anti-inflammatory drugs are caspase inhibitors. Cell Chem. Biol. 24, 281–292 (2017).
Corsello, S. M. et al. The Drug Repurposing Hub: a next-generation drug library and information resource. Nat. Med. 23, 405–408 (2017).
Mishima, E. et al. Drugs repurposed as antiferroptosis agents suppress organ damage, including AKI, by functioning as lipid peroxyl radical scavengers. J. Am. Soc. Nephrol. 31, 280–296 (2020).
Shah, R., Shchepinov, M. S. & Pratt, D. A. Resolving the role of lipoxygenases in the initiation and execution of ferroptosis. ACS Cent. Sci. 4, 387–396 (2018).
Wolpaw, A. J. et al. Modulatory profiling identifies mechanisms of small molecule-induced cell death. Proc. Natl Acad. Sci. USA 108, E771–E780 (2011).
Shimada, K. et al. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat. Chem. Biol. 12, 497–503 (2016).
Forcina, G. C., Conlon, M., Wells, A., Cao, J. Y. & Dixon, S. J. Systematic quantification of population cell death kinetics in mammalian cells. Cell Syst. 4, 600–610.e6 (2017).
Fitzgerald, J. B., Schoeberl, B., Nielsen, U. B. & Sorger, P. K. Systems biology and combination therapy in the quest for clinical efficacy. Nat. Chem. Biol. 2, 458–466 (2006).
Soula, M. et al. Metabolic determinants of cancer cell sensitivity to canonical ferroptosis inducers. Nat. Chem. Biol. 16, 1351–1360 (2020).
Jost, M. et al. Combined CRISPRi/a-based chemical genetic screens reveal that rigosertib is a microtubule-destabilizing agent. Mol. Cell 68, 210–223.e6 (2017).
Bohnacker, T. et al. Deconvolution of Buparlisib’s mechanism of action defines specific PI3K and tubulin inhibitors for therapeutic intervention. Nat. Commun. 8, 14683 (2017).
Smolinski, M. P. et al. Discovery of novel dual mechanism of action SRC signaling and tubulin polymerization inhibitors (KX2-391 and KX2-361). J. Med. Chem. 61, 4704–4719 (2018).
Viswanathan, V. S. et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 547, 453–457 (2017).
Song, X. et al. JTC801 induces pH-dependent death specifically in cancer cells and slows growth of tumors in mice. Gastroenterology 154, 1480–1493 (2018).
Tomlin, F. M. et al. Inhibition of NGLY1 inactivates the transcription factor Nrf1 and potentiates proteasome inhibitor cytotoxicity. ACS Cent. Sci. 3, 1143–1155 (2017).
Richards, R. et al. Drug antagonism and single-agent dominance result from differences in death kinetics. Nat. Chem. Biol. 16, 791–800 (2020).
Zilka, O. et al. On the mechanism of cytoprotection by ferrostatin-1 and liproxstatin-1 and the role of lipid peroxidation in ferroptotic cell death. ACS Cent. Sci. 3, 232–243 (2017).
Shah, R., Margison, K. & Pratt, D. A. The potency of diarylamine radical-trapping antioxidants as inhibitors of ferroptosis underscores the role of autoxidation in the mechanism of cell death. ACS Chem. Biol. 12, 2538–2545 (2017).
Gao, M., Monian, P., Quadri, N., Ramasamy, R. & Jiang, X. Glutaminolysis and transferrin regulate ferroptosis. Mol. Cell 59, 298–308 (2015).
Skouta, R. et al. Ferrostatins inhibit oxidative lipid damage and cell death in diverse disease models. J. Am. Chem. Soc. 136, 4551–4556 (2014).
Lu, T. et al. Up-regulation of hypoxia-inducible factor antisense as a novel approach to treat ovarian cancer. Theranostics 10, 6959–6976 (2020).
Jeitner, T. M. Optimized ferrozine-based assay for dissolved iron. Anal. Biochem. 454, 36–37 (2014).
Shah, R., Farmer, L. A., Zilka, O., Van Kessel, A. T. M. & Pratt, D. A. Beyond DPPH: use of fluorescence-enabled inhibited autoxidation to predict oxidative cell death rescue. Cell Chem. Biol. 26, 1594–1607.e7 (2019).
Silverman, S. L. et al. Efficacy of bazedoxifene in reducing new vertebral fracture risk in postmenopausal women with osteoporosis: results from a 3-year, randomized, placebo-, and active-controlled clinical trial. J. Bone Miner. Res. 23, 1923–1934 (2008).
Komm, B. S. et al. Bazedoxifene acetate: a selective estrogen receptor modulator with improved selectivity. Endocrinology 146, 3999–4008 (2005).
Le Naour, A. et al. EO771, the first luminal B mammary cancer cell line from C57BL/6 mice. Cancer Cell Int 20, 328 (2020).
Perez, M. A., Magtanong, L., Dixon, S. J. & Watts, J. L. Dietary lipids induce ferroptosis in caenorhabditiselegans and human cancer cells. Dev. Cell 54, 447–454.e4 (2020).
Valvezan, A. J. & Manning, B. D. Molecular logic of mTORC1 signalling as a metabolic rheostat. Nat. Metab. 1, 321–333 (2019).
Yang, W. S. & Stockwell, B. R. Synthetic lethal screening identifies compounds activating iron-dependent, nonapoptotic cell death in oncogenic-RAS-harboring cancer cells. Chem. Biol. 15, 234–245 (2008).
Kang, S. A. et al. mTORC1 phosphorylation sites encode their sensitivity to starvation and rapamycin. Science 341, 1236566 (2013).
Rees, M. G. et al. Correlating chemical sensitivity and basal gene expression reveals mechanism of action. Nat. Chem. Biol. 12, 109–116 (2016).
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012).
Wolfson, R. L. & Sabatini, D. M. The dawn of the age of amino acid sensors for the mTORC1 pathway. Cell Metab. 26, 301–309 (2017).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164 (2016).
Harding, H. P. et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol. Cell 11, 619–633 (2003).
Dai, C. et al. Transcription factors in ferroptotic cell death. Cancer Gene Ther. 27, 645–656 (2020).
Nakamura, A. et al. Inhibition of GCN2 sensitizes ASNS-low cancer cells to asparaginase by disrupting the amino acid response. Proc. Natl Acad. Sci. USA 115, E7776–E7785 (2018).
Ye, J. et al. The GCN2-ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 29, 2082–2096 (2010).
Jover-Mengual, T. et al. Molecular mechanisms mediating the neuroprotective role of the selective estrogen receptor modulator, bazedoxifene, in acute ischemic stroke: a comparative study with 17beta-estradiol. J. Steroid Biochem. Mol. Biol. 171, 296–304 (2017).
Lan, Y. L. et al. Bazedoxifene protects cerebral autoregulation after traumatic brain injury and attenuates impairments in blood-brain barrier damage: involvement of anti-inflammatory pathways by blocking MAPK signaling. Inflamm. Res 68, 311–323 (2019).
Yu, X. & Long, Y. C. Crosstalk between cystine and glutathione is critical for the regulation of amino acid signaling pathways and ferroptosis. Sci. Rep. 6, 30033 (2016).
Ratan, R. R., Murphy, T. H. & Baraban, J. M. Macromolecular synthesis inhibitors prevent oxidative stress-induced apoptosis in embryonic cortical neurons by shunting cysteine from protein synthesis to glutathione. J. Neurosci. 14, 4385–4392 (1994).
Nofal, M., Zhang, K., Han, S. & Rabinowitz, J. D. mTOR inhibition restores amino acid balance in cells dependent on catabolism of extracellular protein. Mol. Cell 67, 936–946.e5 (2017).
Baba, Y. et al. Protective effects of the mechanistic target of rapamycin against excess iron and ferroptosis in cardiomyocytes. Am. J. Physiol. Heart Circ. Physiol. 314, H659–H668 (2018).
Cramer, S. L. et al. Systemic depletion of l-cyst(e)ine with cyst(e)inase increases reactive oxygen species and suppresses tumor growth. Nat. Med. 23, 120–127 (2017).
Dixon, S. J. et al. Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. eLife 3, e02523 (2014).
Tarangelo, A. et al. p53 suppresses metabolic stress-induced ferroptosis in cancer cells. Cell Rep. 22, 569–575 (2018).
Haidasz, E. A., Van Kessel, A. T. & Pratt, D. A. A continuous visible light spectrophotometric approach to accurately determine the reactivity of radical-trapping antioxidants. J. Org. Chem. 81, 737–744 (2016).
Li, B. et al. Besting vitamin E: sidechain substitution is key to the reactivity of naphthyridinol antioxidants in lipid bilayers. J. Am. Chem. Soc. 135, 1394–1405 (2013).
Watts, J. L. & Browse, J. Genetic dissection of polyunsaturated fatty acid synthesis in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 99, 5854–5859 (2002).
Cao, J. Y. et al. A genome-wide haploid genetic screen identifies regulators of glutathione abundance and ferroptosis sensitivity. Cell Rep. 26, 1544–1556.e8 (2019).
Acknowledgements
We thank J. Cao, A. Tarangelo, Z. Inde and E. Kolebrander Ho for experimental assistance, and B. Stockwell, L. Li, M. Bassik, J. Ye and M. Cyert for reagents. Certain constructs were obtained from Addgene. This work was supported by awards from the NSERC (grant no. RGPIN-06741-2016) to D.A.P., and the National Institutes of Health (grant no. 5T32GM007276) to D.A.A. (grant no. 1R01GM133883) to J.L.W. and (grant nos. 4R00CA166517-03 and 1R01GM122923) to S.J.D.
Author information
Authors and Affiliations
Contributions
M.C., G.C.F. and A.W. performed kinetic modulatory profiling and follow-up experiments. A.K. and L.M. performed S. cerevisiae experiments. M.A.P. performed C. elegans experiments. M.M. performed liposomal experiments. C.D.P. and D.A.A. performed mTOR and amino acid deprivation experiments. J.L.W., D.A.P. and S.J.D. supervised experiments. All authors analyzed the data. M.C., C.D.P. and S.J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.J.D. is a member of the scientific advisory board for Ferro Therapeutics, has consulted for Toray Industries and AbbVie Inc., and is an inventor on patents related to ferroptosis.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Examination of compound interactions on cell death.
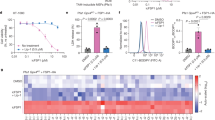
a, Chemical structures. b, Cell death determined using STACK. c, NRF1 (NFE2L1) protein levels. Blot is representative of three independent experiments. d, Cell death determined using STACK. e, Quantification of the timing of population cell death onset (DO) from the lethal fraction curves in d. f, Cell death in two different cell lines determined using STACK. Results in b, d and e are mean ± SD from three independent experiments, while in f results are from two or three independent experiments.
Extended Data Fig. 2 Investigating ferroptosis inhibitors.
a, Cell death data extracted from the compendium for erastin treatment. Each dot represents a single modulator compound, tested once at 5 µM, organized together by major target class. Lower nAUC values indicate greater death suppression. MEKi: mitogen activated protein kinase kinase 1/2 inhibitors (n = 14); AdRm: adrenergic receptor modulators (n = 55); ER/PRm: estrogen/progesterone receptor modulators (n = 29); VEGFRi: vascular endothelial growth factor receptor inhibitors (n = 20); K+ Cm: potassium channel modulators (n = 19). The vertical dotted line indicates the mean lethality of the control erastin + DMSO conditions. b, Cell death determined by counting SYTOX Green positive (SG+) objects. The experiment was performed twice on different days and data represents mean ± SD.
Extended Data Fig. 3 Cell free RTA and iron chelator profiling.
a, Overview of cell free compound profiling for radical trapping and Fe2+-binding activity. b, Cell death at 48 h in HT-1080N cells treated with erastin2 (1 µM) and candidate radical trapping compounds (50 µM, n = 100) plotted against predicted hydrophilicity (LogS), predicted lipophilicity (LogP) and the %DPPH inhibition values from the cell-free assay. Dotted lines indicate a lethal fraction of 0.2. Spearman correlation values are reported with the 95% confidence interval (C.I.). Exact P values (two-tailed) are reported where computable.
Extended Data Fig. 4 Bazedoxifene suppresses ferroptosis in mammalian cells.
a, Cell death quantified as the number of SYTOX Green positive (SG+) objects (that is dead cells) over time. Data are from two independent experiments. b,c, Cell death quantified by SG+ object counting. Data are from three or four independent experiments. d, Outline of the STY-BODIPY kinetic competition assay. Egg-phosphatidylcholine (1 mM) and STY-BODIPY (10 µM) are incubated with 0.2 mM di-tert-undecyl hyponitrite (DTUN), in addition to a radical trapping antioxidant (RTA-H).
Extended Data Fig. 5 Bazedoxifene prevents ferroptosis in C. elegans.
a, Representative images of DAPI-stained adult C. elegans under the different treatment conditions indicated below each image. The gonads of fertile worms are indicated (arrows). DGLA: dihomo-γ-linolenic acid (125 µM); Baz: bazedoxifene (150 µM). Scale bar = 100 µm. Imaging was repeated twice and representative animals from one experiment are shown. b, Polyunsaturated fatty acid levels as a function of total lipids determined in worms using gas chromatography/mass spectrometry. Results are from two independent experiments on separate populations of worms.
Extended Data Fig. 6 mTOR regulates ferroptosis sensitivity.
a, Cell death over time determined using STACK. Cells were infected with control (scrambled) shRNA or shRNAs targeting RPTOR or RICTOR for 72 h prior to compound treatment. INK128 was used as a positive control. Results are mean ± SD from three independent experiments. b, Expression and phosphorylation of proteins in the mTOR pathway following infection of HT-1080 cells as in a. Blot is representative of three independent experiments.
Extended Data Fig. 7 mTOR inhibitors suppress ferroptosis.
a, Cell death determined using STACK in three different cell lines. Results are mean ± SD from three independent experiments. b, 4E-BP1 protein phosphorylation and total levels. Blot is representative of three independent experiments. c, Total glutathione (GSH + GSSG) levels measured using Ellman’s reagent (DNTB). d, System xc- activity inferred from glutamate release over 2 h from HT-1080 and U-2 OS cells treated as indicated. e, Cell death determined using STACK. Results in c-e are from three independent experiments.
Extended Data Fig. 8 mTOR and protein synthesis regulate ferroptosis.
a, Phosphorylation and levels of mTOR pathway effectors in U-2 OS cells. Blot is representative of three independent experiments. b, Analysis of Cancer Therapeutics Response Portal (CTRP) dataset for ferroptosis-inducing compounds. c, Cell death determined using STACK. Erastin2 was used at 2 µM in all cell lines except Caki-1N (1 µM), ML162 was used at 4 µM in all cell lines except Caki-1N (2 µM). CHX: cycloheximide, BSO: buthionine sulfoximine. Results are mean ± SD from three independent experiments.
Extended Data Fig. 9
a, Phosphorylation and levels of mTOR pathway effectors. Blot is representative of two independent experiments. b, Fold-change in amino acids levels in HT-1080 cells determined using liquid chromatography coupled to mass spectrometry.
Extended Data Fig. 10 Arginine uptake regulates ferroptosis.
a, Dose-dependent effect of arginine (Arg) resupplementation on erastin2-induced cell death determined using STACK. Erastin2 was used at 2 µM (U-2 OSN) or 4 µM (A549N). b, Cell death as determined using STACK in cells grown in complete medium (CM), switched to -Arg medium at the time of compound addition, or 24 h before compound addition. c, Detection of puromycylated peptides. Results are representative of three independent experiments. d, Quantification of results from three puromycylation experiments, as in c. e, SYTOX Green positive (SG+) dead cell counts normalized to initial cell confluence. f, Confirmation of ATF4 knockdown at the protein level. Blot is representative of two independent experiments. Results in a,b and e represent mean ± SD from three independent experiments.
Supplementary information
Supplementary Information
Supplementary Table 1 and Figs. 1–5.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 4
Uncropped western blots for Fig. 4.
Source Data Fig. 6
Uncropped western blots for Fig. 6.
Source Data Extended Data Fig. 1
Uncropped western blots.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3a.
Source Data Extended Data Fig. 6
Uncropped western blots.
Source Data Extended Data Fig. 7
Uncropped western blots.
Source Data Extended Data Fig. 8
Uncropped western blots.
Source Data Extended Data Fig. 9
Uncropped western blots.
Source Data Extended Data Fig. 10
Uncropped western blots.
Rights and permissions
About this article
Cite this article
Conlon, M., Poltorack, C.D., Forcina, G.C. et al. A compendium of kinetic modulatory profiles identifies ferroptosis regulators. Nat Chem Biol 17, 665–674 (2021). https://doi.org/10.1038/s41589-021-00751-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00751-4
This article is cited by
-
Synergism of non-thermal plasma and low concentration RSL3 triggers ferroptosis via promoting xCT lysosomal degradation through ROS/AMPK/mTOR axis in lung cancer cells
Cell Communication and Signaling (2024)
-
Polyamine-mediated ferroptosis amplification acts as a targetable vulnerability in cancer
Nature Communications (2024)
-
The cell biology of ferroptosis
Nature Reviews Molecular Cell Biology (2024)
-
CGI1746 targets σ1R to modulate ferroptosis through mitochondria-associated membranes
Nature Chemical Biology (2024)
-
Nanomedicine targeting ferroptosis to overcome anticancer therapeutic resistance
Science China Life Sciences (2024)