Abstract
The natural antivitamin 2′-methoxy-thiamine (MTh) is implicated in the suppression of microbial growth. However, its mode of action and enzyme-selective inhibition mechanism have remained elusive. Intriguingly, MTh inhibits some thiamine diphosphate (ThDP) enzymes, while being coenzymatically active in others. Here we report the strong inhibition of Escherichia coli transketolase activity by MTh and unravel its mode of action and the structural basis thereof. The unique 2′-methoxy group of MTh diphosphate (MThDP) clashes with a canonical glutamate required for cofactor activation in ThDP-dependent enzymes. This glutamate is forced into a stable, anticatalytic low-barrier hydrogen bond with a neighboring glutamate, disrupting cofactor activation. Molecular dynamics simulations of transketolases and other ThDP enzymes identify active-site flexibility and the topology of the cofactor-binding locale as key determinants for enzyme-selective inhibition. Human enzymes either retain enzymatic activity with MThDP or preferentially bind authentic ThDP over MThDP, while core bacterial metabolic enzymes are inhibited, demonstrating therapeutic potential.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The refined structural protein models and corresponding structure-factor amplitudes are deposited in the PDB under accession codes 6TJ8 (EcTK in complex with cofactor analog MThDP) and 6TJ9 (EcTK in complex with cofactor analog MThDP and substrate X5P). The structures cited in this publication (1QGD, 2R8O, 3MOS, 2IEA and 3EXE) are available under their respective PDB accession codes. Input files for the MD simulations are available as part of the Supplementary Information. All other data are available on request.
References
Alanis, A. J. Resistance to antibiotics: are we in the post-antibiotic era? Arch. Med. Res. 36, 697–705 (2005).
Kohansky, M. A., Dwyer, D. J. & Collins, J. J. How antibiotics kill bacteria: from targets to networks. Nat. Rev. Microbiol. 8, 423–435 (2010).
Otani, S., Takatsu, M., Nakano, M., Kasai, S. & Miura, R. Letter: roseoflavin, a new antimicrobial pigment from Streptomyces. J. Antibiot. 27, 86–87 (1974).
Wada, K. & Haga, M. Ginkgo Biloba—A Global Treasure (eds. Hori, T. et al.) 309–321 (Springer Japan, 1997).
Drautz, H., Messerer, W., Zähner, H., Breiding-Mack, S. & Zeeck, A. Metabolic products of microorganisms. 239. Bacimethrin isolated from Streptomyces albus identification, derivatives, synthesis and biological properties. J. Antibiot. 40, 1431–1439 (1987).
Reddick, J. J. et al. The mechanism of action of bacimethrin, a naturally occurring thiamin antimetabolite. Bioorg. Med. Chem. Lett. 11, 2245–2248 (2001).
Pedrolli, D. B. et al. The antibiotics roseoflavin and 8-demethyl-8-amino-riboflavin from Streptomyces davawensis are metabolized by human flavokinase and human FAD synthetase. Biochem. Pharmacol. 82, 1853–1859 (2011).
Leistner, E. & Drewke, C. Ginkgo biloba and ginkgotoxin. J. Nat. Prod. 73, 86–92 (2010).
Lee, E. R., Blount, K. F. & Breaker, R. R. Roseoflavin is a natural antibacterial compound that binds to FMN riboswitches and regulates gene expression. RNA Biol. 6, 187–194 (2009).
Langer, S., Hashimoto, M., Hobl, B., Mathes, T. & Mack, M. Flavoproteins are potential targets for the antibiotic roseoflavin in Escherichia coli. J. Bacteriol. 195, 4037–4045 (2013).
Nemeria, N. S. et al. Competence of thiamin diphosphate-dependent enzymes with 2′-methoxythiamin diphosphate derived from bacimethrin, a naturally occurring thiamin antivitamin. Biochemistry 55, 1135–1148 (2016).
Schneider, G. & Lindqvist, Y. Crystallography and mutagenesis of transketolase: mechanistic implications for enzymatic thiamin catalysis. Biochim. Biophys. Acta 1385, 387–398 (1998).
Tittmann, K. Sweet siblings with different faces: the mechanisms of FBP and F6P aldolase, transaldolase, transketolase and phosphoketolase revisited in light of recent structural data. Bioorg. Chem. 57, 263–280 (2014).
Dai, S. et al. Low-barrier hydrogen bonds in enzyme cooperativity. Nature 573, 609–613 (2019).
Frank, R. A., Titman, C. M., Pratap, J. V., Luisi, B. F. & Perham, R. N. A molecular switch and proton wire synchronize the active sites in thiamine enzymes. Science 306, 872–876 (2004).
Kern, D. et al. How thiamine diphosphate is activated in enzymes. Science 275, 67–70 (1997).
Asztalos, P. et al. Strain and near attack conformers in enzymic thiamin catalysis: X-ray crystallographic snapshots of bacterial transketolase in covalent complex with donor ketoses xylulose 5-phosphate and fructose 6-phosphate, and in noncovalent complex with acceptor aldose ribose 5-phosphate. Biochemistry 46, 12037–12052 (2007).
Meyer, D., Neumann, P., Ficner, R. & Tittmann, K. Observation of a stable carbene at the active site of a thiamin enzyme. Nat. Chem. Biol. 9, 488–490 (2013).
Nemeria, N. S., Chakraborty, S., Balakrishnan, A. & Jordan, F. Reaction mechanisms of thiamin diphosphate enzymes: defining states of ionization and tautomerization of the cofactor at individual steps. FEBS J. 276, 2432–2446 (2009).
Paulikat, M., Wechsler, C., Tittmann, K. & Mata, R. A. Theoretical studies of the electronic absorption spectra of thiamin diphosphate in pyruvate decarboxylase. Biochemistry 56, 1854–1864 (2017).
Tittmann, K. et al. NMR analysis of covalent intermediates in thiamin diphosphate enzymes. Biochemistry 42, 7885–7891 (2003).
Kluger, R. & Tittmann, K. Thiamin diphosphate catalysis: enzymic and nonenzymic covalent intermediates. Chem. Rev. 108, 1797–1833 (2008).
Muller, Y. A. et al. A thiamin diphosphate binding fold revealed by comparison of the crystal structures of transketolase, pyruvate oxidase and pyruvate decarboxylase. Structure 1, 95–103 (1993).
Kaplun, A. et al. Glyoxylate carboligase lacks the canonical active site glutamate of thiamine-dependent enzymes. Nat. Chem. Biol. 4, 113–118 (2008).
Burgi, H. B., Dunitz, J. D., Lehn, J. M. & Wipff, G. Stereochemistry of reaction paths at carbonyl centers. Tetrahedron 30, 1563–1572 (1974).
Lüdtke, S. et al. Sub-ångström-resolution crystallography reveals physical distortions that enhance reactivity of a covalent enzymatic intermediate. Nat. Chem. 5, 762–767 (2013).
Neumann, P. & Tittmann, K. Marvels of enzyme catalysis at true atomic resolution: distortions, bond elongations, hidden flips, protonation states and atom identities. Curr. Opin. Struct. Biol. 29, 122–133 (2014).
Booth, C. K. & Nixon, P. F. Reconstitution of holotransketolase is by a thiamin‐diphosphate‐magnesium complex. Eur. J. Biochem. 218, 261–265 (1993).
Mitschke, L. et al. The crystal structure of human transketolase and new insights into its mode of action. J. Biol. Chem. 285, 31559–31570 (2010).
Ciszak, E. M., Korotchkina, L. G., Dominiak, P. M., Sidhu, S. & Patel, M. S. Structural basis for flip-flop action of thiamin pyrophosphate-dependent enzymes revealed by human pyruvate dehydrogenase. J. Biol. Chem. 278, 21240–21246 (2003).
Jarzynski, C. Equilibrium free-energy differences from nonequilibrium measurements: a master-equation approach. Phys. Rev. E 56, 5018–5035 (1997).
Warshel, A., Papazyan, A. & Kollman, P. A. On low-barrier hydrogen bonds and enzyme catalysis. Science 269, 102–106 (1995).
Warshel, A. & Papazyan, A. Energy considerations show that low-barrier hydrogen bonds do not offer a catalytic advantage over ordinary hydrogen bonds. Proc. Natl Acad. Sci. USA 93, 13665–13670 (1996).
Lehwess-Litzmann, A. et al. Twisted Schiff base intermediates and substrate locale revise transaldolase mechanism. Nat. Chem. Biol. 7, 678–684 (2011).
Light, S. H., Minasov, G., Duban, M. E. & Anderson, W. F. Adherence to Burgi–Dunitz stereochemical principles requires significant structural rearrangements in Schiff-base formation: insights from transaldolase complexes. Acta Crystallogr. D Biol. Crystallogr. 70, 544–552 (2014).
Hur, S. & Bruice, T. C. The near attack conformation approach to the study of the chorismate to prephenate reaction. Proc. Natl Acad. Sci. USA 100, 12015–12020 (2003).
Kluger, R. Catalyzing decarboxylation by taming carbon dioxide. Pure Appl. Chem. 87, 353–360 (2015).
Bailey, S. S. et al. Enzymatic control of cycloadduct conformation ensures reversible 1,3-dipolar cycloaddition in a prFMN-dependent decarboxylase. Nat. Chem. 11, 1049–1057 (2019).
Fersht, A. Structure and Mechanism in Protein Science (W.H. Freeman and Company, 1999).
Fujihashi, M. et al. Substrate distortion contributes to the catalysis of orotidine 5'-monophosphate decarboxylase. J. Am. Chem. Soc. 135, 17432–17443 (2013).
Jafari, R. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 9, 2100–2122 (2014).
Begley, T. The mechanistic enzymology of thiamin biosynthesis. FASEB J. 29, (2015).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Bailey, S. The CCP4 suite—programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Marks, C. et al. Sphinx: merging knowledge-based and ab initio approaches to improve protein loop prediction. Bioinformatics 33, 1346–1353 (2017).
Doerr, S., Harvey, M. J., Noe, F. & De Fabritiis, G. HTMD: high-throughput molecular dynamics for molecular discovery. J. Chem. Theory Comput. 12, 1845–1852 (2016).
Olsson, M. H. M., Sondergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Dolinsky, T. J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Hornak, V. et al. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Wang, J. M., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
da Silva, A. & Vranken, W. ACPYPE—AnteChamber PYthon Parser interfacE. BMC Res. Notes 5, 367 (2012).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2, 19–25 (2015).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., Dinola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals—a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Gapsys, V., Michielssens, S., Seeliger, D. & de Groot, B. L. Pmx: automated protein structure and topology generation for alchemical perturbations. J. Comput. Chem. 36, 348–354 (2015).
Shirts, M. R., Bair, E., Hooker, G. & Pande, V. S. Equilibrium free energies from nonequilibrium measurements using maximum-likelihood methods. Phys. Rev. Lett. 91, 140601 (2003).
Aldeghi, M., Gapsys, V. & de Groot, B. L. Accurate estimation of ligand binding affinity changes upon protein mutation. ACS Cent. Sci. 4, 1708–1718 (2018).
Acknowledgements
This study was supported by the Deutsche Forschungsgemeinschaft (FOR1296/TP3 to K.T.). We acknowledge access to beamline P14 at DESY/EMBL and thank G. Bourenkov and T. Schneider for local support. We thank R. Mata, M. McLeish and R. Kluger for discussion.
Author information
Authors and Affiliations
Contributions
K.T. designed and coordinated the project. F.R.v.P. expressed and purified E. coli and human TK, crystallized proteins and determined the X-ray crystallographic protein structures supervised by K.T. F.R.v.P. and K.T. carried out functional analyses and interpreted functional and structural data. M.A. designed, performed, analyzed and interpreted the computer simulations. B.L.d.G. designed and supervised the execution of the computer simulations and interpreted their results. B.S. synthesized MThDP under the supervision of T.B. All authors discussed the project data. F.R.v.P., M.A. and K.T. wrote the paper with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
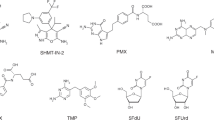
Extended Data Fig. 1 Structure and presumed mode of action of naturally occurring vitamin B antivitamins.
a, Chemical structures of vitamins B2 (riboflavin), B1 (thiamin) and B6 (pyridoxin/pyridoxal), and of corresponding antivitamins highlighting the site of modification (colored in magenta). b, Suggested mode of inhibition for the antivitamins shown in (a).
Extended Data Fig. 2 Transketolase reactions and mechanism of cofactor activation in ThDP enzymes.
a, Physiological substrates and reactions of transketolase showing the donor ketose D-xylulose-5-phosphate in blue (scissile C2-C3 bond indicated in black) and alternative aldose acceptors D-erythrose-4-phosphate and D-ribose-5-phosphate in red. b, Proposed mechanism of cofactor activation highlighting different cofactor protonation states and critical proton transfers. Protonation of the cofactor aminopyrimidine (AP) at atom N1′ by canonical residue E411 is thought to generate the aminopyrimidinium cation (APH+) form of the cofactor. Liberation of a proton from the exocyclic 4′-amino group yields the 1′,4′-iminotautomer (IP), which, owing to its high basicity, catalyses the deprotonation of the thiazolium at atom C2 (directly or via a water). The resultant carbanion/carbene nucleophilically attacks the donor substrate affording the covalent substrate-ThDP conjugate that undergoes further processing.
Extended Data Fig. 3 Spectroscopic analysis of cofactor binding to E. coli transketolase.
a, Quantitative analysis of cofactor binding to E. coli transketolase using genuine ThDP (black) and MThDP (orange) and monitoring fluorescence quenching of the protein. Experimental conditions are detailed in the Methods section. In case of ThDP, data were fitted with a quadratic function and yielded an apparent equilibrium binding constant of KDapp = 0.23 ± 0.01 µM. For MThDP, an equation with two quadratic terms was used for fitting the data as we observed a high-affinity binding regime (KDapp 1 = 0.09 ± 0.02 µM; 20% amplitude) and a medium-affinity binding regime (KDapp 2 = 13.39 ± 1.99 µM; 80% amplitude). All measurements were carried out in triplicate and are shown as mean ± s.d. b, Absorbance and near-UV circular dichroism (CD) spectra of E. coli transketolase reconstituted with either genuine ThDP at saturating concentration (red spectra) or MThDP at increasing concentrations (0–150 µM, colored spectra). Experimental conditions are detailed in the Methods section. Note the prominent CD signal with a negative signature at ~325 nm for EcTK in complex with ThDP that is absent for the enzyme complex with MThDP implying a different binding mode of MThDP. All experiments were independently repeated twice with similar results.
Extended Data Fig. 4 SA omit electron density maps for substrate X5P and structural superposition of E. coli transketolase in non-covalent versus covalent complex with X5P.
a, Simulated-annealing mFo-DFc omit electron density maps of substrate X5P noncovalently bound to E. coli transketolase reconstituted with 2′-methoxy-ThDP (MThDP). Omit maps are shown at contour levels of 7σ (in blue) and 4σ (in grey). The two chains of the homodimer (chain A and B) are colored individually. Simulated annealing omit maps were generated after omitting substrate X5P from the structural model. Five cycles of PHENIX.REFINE45 were run applying cartesian simulated annealing in cycles 2 and 4, with a start temperature of 5000 K and an end temperature of 300 K. b, Superposition of the active site of E. coli transketolase in noncovalent (in yellow, this study) and covalent complex with substrate X5P (in grey, pdb code 2R8O) in stereo view showing the MThDP cofactor, substrate X5P, the covalent X5P-ThDP conjugate and selected protein groups. The active sites are shown from two different perspectives (top panel: side view; bottom panel: viewed down the substrate channel).
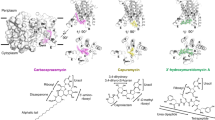
Extended Data Fig. 5 Structures of cooperativity proton wires linking the two remote active sites in transketolases and pyruvate dehydrogenases.
a, b, Structure of the cooperativity proton wire in E. coli (top) and human (bottom) transketolase showing selected amino acid residues, the substrate-ThDP intermediates and water molecules (pdb codes 2R8O & 4KXW). Hydrogen-bonding interactions are highlighted with dashed lines. Note that the wire architecture is highly conserved in both enzymes and consists of 6 glutamate residues and bridging water molecules (cyan spheres) providing a direct proton transfer pathway. c, d, Structure of the cooperativity communication channel in E. coli (top) and human (bottom) PDH E1 showing selected amino acid residues, the substrate-ThDP intermediates and water molecules (pdb codes 2IEA & 3EXE). Hydrogen-bonding interactions are highlighted with dashed lines. Note that the wire architecture in E. coli PDH is similar to the ones in E. coli and human transketolase, and consists of in total 6 glutamate residues and bridging water molecules providing a direct proton transfer pathway. In contrast, the putative communciation channel in human PDH features glutamines (Q172A, Q172C) that replace a critical wire glutamate (E235 in E. coli PDH) thus argueing against a direct proton transfer pathway.
Extended Data Fig. 6 Molecular dynamics analysis of cofactor binding in ThDP enzymes.
a–d, Scatter plots of the binding free energy calculations. The Amber99sb*-ILDN/GAFF(v2.1) force field is referred to as “Amber”, and the Charmm36/CGenFF(v3.0.1) force files is referred to as “Charmm”. The mean and 95% confidence intervals of the calculated ΔΔG values are shown based on n = 10 independent calculations. ThDP: thiamine diphosphate; MThDP: methoxythiamin diphosphate; TP1: 4′-desamino ThDP; TP2: N3′-pyridyl ThDP; WT: wild-type; RMSE: root-mean-square error. e,f, Computed isotropic B-factors from molecular dynamics simulations. e, Binding pocket plasticity. The ThDP ligand is shown as green ball and sticks, and the residues within 5 Å distance are shown as sticks and color-coded according to their atomic B-factors. The Mg2+ ion is shown as a larger green sphere. B-factors were derived from the protein heavy-atom RMSFs, which were calculated using the final snapshots of the 500 short non-equilibrium trajectories performed with the Amber force field, as described in the main text (PDB-ID 1QGD for E. coli TK; PDB-ID 3MOS for human TK; PDB-ID 2IEA for E. coli PDH; PDB-ID 3EXE for human PDH). f, Distribution of the average binding pocket B-factors for the four complexes studied. The average RMSF for each of n = 10 independent simulations is shown as a swarmplot. The box plots are derived from these n = 10 average RMSF values. The centres of the boxes indicate the median, the bounds of the boxes indicate the first and third quartiles of the distributions, and the whiskers extend to samples up to 1.5 of the interquartile range. Samples outside the marked extrema are classified as potential outliers. Overall, the data shown in e and f suggest a higher plasticity for the binding pockets of HsTK, EcPDH, and HsPDH than for that of EcTK.
Extended Data Figure 7 Distributions of proton donor-acceptor distances and angles for enzyme-bound ThDP and MThDP.
a, Distributions of proton donor-acceptor distances for enzyme-bound ThDP and MThDP in EcTK, HsTK, EcPDH, and HsPDH. The donor is the oxygen atom on the canonical, cofactor activating glutamic acid, and the acceptor is the N1’ atom on the aminopyrimidine ring of ThDP and MThDP. The area of the plots, where the distance is in the favourable regime <3 Å is highlighted in grey. The fraction of simulation frames in which the distance was <3 Å is reported for both ThDP and MThDP; the fraction difference between MThDP and ThDP is also reported (number highlighted in red). A negative difference means that MThDP has a smaller fraction of frames with distances <3 Å. b, Distributions of proton donor-acceptor angles for enzyme-bound ThDP and MThDP in EcTK, HsTK, EcPDH, and HsPDH. The angle is defined between the acceptor N1’ atom in ThDP and MThDP, respectively, the donor oxygen atom on the canonial, cofactor activating glutamic acid, and the neighboring δ-carbon on the same glutamic acid. The area of the plots, where the angle is in the favourable regime between 90 and 150 degrees is highlighted in grey. The fraction of simulation frames within this area is reported for both ThDP and MThDP. The fraction difference between MThDP and ThDP is also reported. A positive difference means that MThDP has a larger fraction of frames where this angle is between 90 and 150 degrees. Note that the fractions in the favourable regime are larger for the human enzymes in complex with MThDP compared to the E. coli orthologs.
Supplementary information
Supplementary Information
Supplementary Tables 1–5.
Supplementary Data
Input files for MD calculations.
Rights and permissions
About this article
Cite this article
Rabe von Pappenheim, F., Aldeghi, M., Shome, B. et al. Structural basis for antibiotic action of the B1 antivitamin 2′-methoxy-thiamine. Nat Chem Biol 16, 1237–1245 (2020). https://doi.org/10.1038/s41589-020-0628-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0628-4