Abstract
The p53 homolog TAp63α is the transcriptional key regulator of genome integrity in oocytes. After DNA damage, TAp63α is activated by multistep phosphorylation involving multiple phosphorylation events by the kinase CK1, which triggers the transition from a dimeric and inactive conformation to an open and active tetramer that initiates apoptosis. By measuring activation kinetics in ovaries and single-site phosphorylation kinetics in vitro with peptides and full-length protein, we show that TAp63α phosphorylation follows a biphasic behavior. Although the first two CK1 phosphorylation events are fast, the third one, which constitutes the decisive step to form the active conformation, is slow. Structure determination of CK1 in complex with differently phosphorylated peptides reveals the structural mechanism for the difference in the kinetic behavior based on an unusual CK1/TAp63α substrate interaction in which the product of one phosphorylation step acts as an inhibitor for the following one.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Johnston, R. J. & Wallace, W. H. Normal ovarian function and assessment of ovarian reserve in the survivor of childhood cancer. Pediatr. Blood Cancer 53, 296–302 (2009).
Maltaris, T., Beckmann, M. W. & Dittrich, R. Review. Fertility preservation for young female cancer patients. Vivo 23, 123–130 (2009).
Suh, E. K. et al. p63 protects the female germ line during meiotic arrest. Nature 444, 624–628 (2006).
Wallace, W. H., Thomson, A. B. & Kelsey, T. W. The radiosensitivity of the human oocyte. Hum. Reprod. 18, 117–121 (2003).
Quast, U. Whole body radiotherapy: a TBI-guideline. J. Med. Phys. 31, 5–12 (2006).
Woodard, T. L. & Bolcun-Filas, E. Prolonging reproductive life after cancer: the need for fertoprotective therapies. Trends Cancer 2, 222–233 (2016).
Livera, G. et al. p63 null mutation protects mouse oocytes from radio-induced apoptosis. Reproduction 135, 3–12 (2008).
Deutsch, G. B. et al. DNA damage in oocytes induces a switch of the quality control factor TAp63α from dimer to tetramer. Cell 144, 566–576 (2011).
Kerr, J. B. et al. DNA damage-induced primordial follicle oocyte apoptosis and loss of fertility require TAp63-mediated induction of Puma and Noxa. Mol. Cell 48, 343–352 (2012).
Kim, S. Y. et al. Transient inhibition of p53 homologs protects ovarian function from two distinct apoptotic pathways triggered by anticancer therapies. Cell Death Differ. 26, 502–515 (2019).
Bolcun-Filas, E., Rinaldi, V. D., White, M. E. & Schimenti, J. C. Reversal of female infertility by Chk2 ablation reveals the oocyte DNA damage checkpoint pathway. Science 343, 533–536 (2014).
Tuppi, M. et al. Oocyte DNA damage quality control requires consecutive interplay of CHK2 and CK1 to activate p63. Nat. Struct. Mol. Biol. 25, 261–269 (2018).
Cesaro, L. & Pinna, L. A. The generation of phosphoserine stretches in phosphoproteins: mechanism and significance. Mol. Biosyst. 11, 2666–2679 (2015).
Knippschild, U. et al. The CK1 family: contribution to cellular stress response and its role in carcinogenesis. Front Oncol. 4, 96 (2014).
Schittek, B. & Sinnberg, T. Biological functions of casein kinase 1 isoforms and putative roles in tumorigenesis. Mol. Cancer 13, 231 (2014).
Coutandin, D. et al. Quality control in oocytes by p63 is based on a spring-loaded activation mechanism on the molecular and cellular level. eLife 5, e13909 (2016).
Prehoda, K. E., Scott, J. A., Mullins, R. D. & Lim, W. A. Integration of multiple signals through cooperative regulation of the N-WASP-Arp2/3 complex. Science 290, 801–806 (2000).
Chen, L., Glover, J. N., Hogan, P. G., Rao, A. & Harrison, S. C. Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 392, 42–48 (1998).
Rinaldi, V. D., Hsieh, K., Munroe, R., Bolcun-Filas, E. M. & Schimenti, J. C. Pharmacological inhibition of the DNA damage checkpoint prevents radiation-induced oocyte death. Genetics 206, 1823–1828 (2017).
Carmell, M. A. et al. A widely employed germ cell marker is an ancient disordered protein with reproductive functions in diverse eukaryotes. eLife 5, e19993 (2016).
Keller, P. J., Schmidt, A. D., Wittbrodt, J. & Stelzer, E. H. Digital scanned laser light-sheet fluorescence microscopy (DSLM) of zebrafish and Drosophila embryonic development. Cold Spring Harb. Protoc. 2011, 1235–1243 (2011).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Hotte, K. et al. Ultra-thin fluorocarbon foils optimise multiscale imaging of three-dimensional native and optically cleared specimens. Sci. Rep. 9, 17292 (2019).
Ferrell, J. E. Jr & Ha, S. H. Ultrasensitivity part III: cascades, bistable switches and oscillators. Trends Biochem. Sci. 39, 612–618 (2014).
Ferrell, J. E. Jr & Ha, S. H. Ultrasensitivity part II: multisite phosphorylation, stoichiometric inhibitors and positive feedback. Trends Biochem. Sci. 39, 556–569 (2014).
Ferrell, J. E. Jr & Ha, S. H. Ultrasensitivity part I: Michaelian responses and zero-order ultrasensitivity. Trends Biochem. Sci. 39, 496–503 (2014).
Ferrell, J. E. Jr & Bhatt, R. R. Mechanistic studies of the dual phosphorylation of mitogen-activated protein kinase. J. Biol. Chem. 272, 19008–19016 (1997).
Serber, Z. et al. A C-terminal inhibitory domain controls the activity of p63 by an intramolecular mechanism. Mol. Cell Biol. 22, 8601–8611 (2002).
Straub, W. E. et al. The C-terminus of p63 contains multiple regulatory elements with different functions. Cell Death Dis. 1, e5 (2010).
Selenko, P. et al. In situ observation of protein phosphorylation by high-resolution NMR spectroscopy. Nat. Struct. Mol. Biol. 15, 321–329 (2008).
Cordier, F. et al. Ordered phosphorylation events in two independent cascades of the PTEN C-tail revealed by NMR. J. Am. Chem. Soc. 134, 20533–20543 (2012).
Narasimamurthy, R. et al. CK1ẟε protein kinase primes the PER2 circadian phosphoswitch. Proc. Natl Acad. Sci. USA 115, 5986–5991 (2018).
Mylona, A. et al. Opposing effects of Elk-1 multisite phosphorylation shape its response to ERK activation. Science 354, 233–237 (2016).
Leroy, A. et al. Spectroscopic studies of GSK3β phosphorylation of the neuronal Tau protein and its interaction with the N-terminal domain of apolipoprotein E. J. Biol. Chem. 285, 33435–33444 (2010).
Philpott, J. M. Casein kinase 1 dynamics underlie substrate selectivity and the PER2 circadian phosphoswitch. eLife 9, e52343 (2020).
Theillet, F. X. et al. Sensitivity-enhanced 13C-NMR for monitoring multisite phosphorylation at physiological temperature and pH. Angew. Chem. Int. Ed. 59, 10411–10415 (2020).
Favier, A. & Brutscher, B. Recovering lost magnetization: polarization enhancement in biomolecular NMR. J. Biomol. NMR 49, 9–15 (2011).
Zhao, B., Li, L., Tumaneng, K., Wang, C. Y. & Guan, K. L. A coordinated phosphorylation by Lats and CK1 regulates YAP stability through SCF(β-TRCP). Genes Dev. 24, 72–85 (2010).
Shinohara, Y. et al. Temperature-sensitive substrate and product binding underlie temperature-compensated phosphorylation in the clock. Mol. Cell 67, 783–798 (2017).
Rossi, V. et al. LH prevents cisplatin-induced apoptosis in oocytes and preserves female fertility in mouse. Cell Death Differ. 24, 72–82 (2017).
Pampaloni, F., Stelzer, E. H. K. & Mattheyer, C. Capillary cell, arrangement and method for accommodating, positioning and examining a microscopic specimen. US patent 20150211981A1 (2015).
Lohr, F., Gebel, J., Henrich, E., Hein, C. & Dotsch, V. Towards complete polypeptide backbone NH assignment via combinatorial labeling. J. Magn. Reson. 302, 50–63 (2019).
Gil, S. et al. NMR spectroscopic studies of intrinsically disordered proteins at near-physiological conditions. Angew. Chem. Int. Ed. 52, 11808–11812 (2013).
Bermel, W. et al. Complete assignment of heteronuclear protein resonances by protonless NMR spectroscopy. Angew. Chem. Int. Ed. 44, 3089–3092 (2005).
McIntosh, L. P. et al. Detection and assignment of phosphoserine and phosphothreonine residues by 13C-31P spin-echo difference NMR spectroscopy. J. Biomol. NMR 43, 31–37 (2009).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
McCoy, A. J. Acknowledging errors: advanced molecular replacement with phaser. Methods Mol. Biol. 1607, 421–453 (2017).
Long, A. M., Zhao, H. & Huang, X. Structural basis for the potent and selective inhibition of casein kinase 1 epsilon. J. Med. Chem. 55, 10307–10311 (2012).
Emsley, P. Tools for ligand validation in COOT. Acta Crystallogr. D Struct. Biol. 73, 203–210 (2017).
Skubak, P., Murshudov, G. N. & Pannu, N. S. Direct incorporation of experimental phase information in model refinement. Acta Crystallogr. D Biol. Crystallogr. 60, 2196–2201 (2004).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 27–28 (1996).
Bekker, H. et al. GROMACS—a parallel computer for molecular-dynamics simulations. Phys. Comput. 92, 252–256 (1993).
Best, R. B. & Hummer, G. Optimized molecular dynamics force fields applied to the helix-coil transition of polypeptides. J. Phys. Chem. B 113, 9004–9015 (2009).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Mamatkulov, S. & Schwierz, N. Force fields for monovalent and divalent metal cations in TIP3P water based on thermodynamic and kinetic properties. J. Chem. Phys. 148, 074504 (2018).
Meagher, K. L., Redman, L. T. & Carlson, H. A. Development of polyphosphate parameters for use with the AMBER force field. J. Comput. Chem. 24, 1016–1025 (2003).
Homeyer, N., Horn, A. H. C., Lanig, H. & Sticht, H. AMBER force-field parameters for phosphorylated amino acids in different protonation states: phosphoserine, phosphothreonine, phosphotyrosine and phosphohistidine. J. Mol. Model. 12, 281–289 (2006).
Greis, K. D. et al. MALDI-TOF MS as a label-free approach to rapid inhibitor screening. J. Am. Soc. Mass Spectrom. 17, 815–822 (2006).
Heap, R. E. et al. Identifying inhibitors of inflammation: a novel high-throughput MALDI-TOF screening assay for salt-inducible kinases (SIKs). SLAS Discov. 22, 1193–1202 (2017).
Acknowledgements
We thank I. Theofel, S. Young and E. Chih-Chao Liang for their review of and input into this manuscript. The research was funded by the DFG (DO 545/18-1), the Centre for Biomolecular Magnetic Resonance (BMRZ) and the Cluster of Excellence Frankfurt (Macromolecular Complexes). M.T. was supported by a Fellowship from the fund of the German Chemical Industry. L.S. and G.H. were supported by the Max Planck Society. F.P., K.H. and E.H.K.S. thank the EU Horizon2020 project LSFM4LIFE (grant no. 668350-2) and the ZonMw-BMBF joint sponsored project ‘The Onconoid Hub’ (grant no. 114027003) for funding. The Structural Genomics Consortium is a registered charity (no. 1097737) that receives funds from the Canadian Institutes for Health Research, the Canadian Foundation for Innovation, Genome Canada through the Ontario Genomics Institute, GlaxoSmithKline, Karolinska Institute, the Knut and Alice Wallenberg Foundation, the Ontario Innovation Trust, the Ontario Ministry for Research and Innovation, Merck & Co., the Novartis Research Foundation, the Swedish Agency for Innovation Systems, the Swedish Foundation for Strategic Research and the Wellcome Trust.
Author information
Authors and Affiliations
Contributions
J.G. and M.T. performed NMR and kinetic assays in vitro and in ovaries. A.C. crystallized and solved the structures of the kinase–peptide complexes. K.H. and F.P. performed microscopy experiments and quantitative semi-automated segmentation. L.S. carried out MD simulations. F.L. performed NMR experiments. N.G. measured tetramerization kinetics. F.F., E.H. and J.M. expressed and purified proteins. M.S. measured phosphorylation kinetics. R.L. helped to analyze the data. M.T., G.H., E.H.K.S., S.K. and V.D. designed experiments and analyzed data. J.G., M.T., A.C. and V.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phosphorylation kinetics of the PAD peptide.
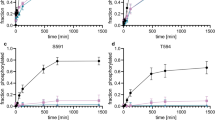
Phosphorylation kinetics of the PAD peptide. a, Overlay of HSQC spectra of unphosphorylated (cyan) and MK2 phosphorylated PAD peptide (red). A large chemical shift change in S582 can be observed as a result of phosphorylation. b, Phosphorylation kinetics of S582 pre-phosphorylated PAD peptide (250 µM) and 12.5 nM CK1 kinase. The phosphorylation of S585 (red) is faster compared to the S588 phosphorylation (yellow) to the extent that the pS582/pS585 intermediate state (cyan) is populated to >1000x of the kinase concentration. c, Phosphorylation kinetics of S582, S585, and S588 pre-phosphorylated PAD peptide (250 µM) with 2.5 µM CK1 kinase. The phosphorylation of S591 (red) is faster than T594 (yellow). The difference is smaller, but the concentration of the intermediate state is at least 4x larger than the kinase concentration. d, Overlay of [15 N, 1H]-HSQC spectra of a pS582/S585A PAD peptide (red) and the same peptide after 500 min of exposure to CK1 (blue, 1:2000 kinase:substrate ratio, 25 °C). Only partial phosphorylation of a single residue in the vicinity of S582 is visible, indicated by the splitting of the signal, accounting for 45% of the population of S582. e, f, Overlay of [15 N, 1H]-HSQC spectra of a pS582/S588A PAD peptide (red) and the same peptide after 500 min of exposure to CK1 and phosphorylation kinetics, demonstrating that S585 and T586 are the only residues phosphorylated by CK1 when the second phosphorylation site for CK1 is eliminated. g, Mutation of T586 to alanine does not change the biphasic behavior of the CK1 phosphorylation of the PAD peptide. All experiments were repeated twice and reproducible. Only one replicate is shown in each case.
Extended Data Fig. 2 Phosphorylation kinetics of PAD peptide mutants.
Phosphorylation kinetics of PAD peptide mutants. a, Phosphorylation kinetics of selected PAD mutants showing a reduction in the kinetic difference between S588 and S591. b, Measured kinetic constants for the phosphorylation of S588 (k2) and S591 (k3) in wild type and different mutant PAD peptides as well as the constants for the second and third phosphorylation event in the YAP1 peptide. Measurements were repeated twice under identical conditions. Individual data points are shown. The height of the bar represents the mean value and single measurements are indicated. The phosphorylation kinetics of YAP was measured once.
Extended Data Fig. 3 Crystallographic details.
Crystallographic details. a–c, Binding of the PAD peptides within CK1δ in the crystal structures. |Fo | -|Fc| omitted electron density map contoured at three-sigma for the bound PAD peptides. d–f, Detailed interactions at the N termini of the PAD peptides within the kinase.
Extended Data Fig. 4 MD simulation and kinetics with CK1 mutants.
MD simulation and kinetics with CK1 mutants. a, MD simulation of CK1 in complex with a shorter PAD-3P peptide (ACE-TPpSSApSTVpSVGSSETRG-NME) with N-terminal acetyl and C-terminal methylamino capping groups showing similar results as the longer peptide (Fig. 5a, b). b, Snapshot at 1 µs, zooming in on the C-terminal region of the shorter p63 peptide. CK1 is shown as a transparent electrostatic surface (blue/red for positive/negative charge) and the p63 peptide is represented as a cyan cartoon. The residues E593, Arg127 and Lys154 are highlighted. The minimum distances between E593 and the basic residues are indicated. c, Phosphorylation kinetics of wild type and selected CK1δ mutants showing a reduction in the kinetic difference between S588 and S591 compared to the wild type kinase for Lys171Glu and Lys154Glu mutants. For Arg127Glu the kinetics is, however, slower. CK1 amino acids are labeled in three-letter code and PAD residues are labeled in one-letter code. All kinetic experiments involving kinase mutants were measured once.
Extended Data Fig. 5 Measurements of binding affinity and enzyme kinetics.
Measurements of binding affinity and enzyme kinetics. a, Three independent KD measurements for each phosphorylation state of the peptide are shown (see also Fig. 6a and Supplementary Table 3). Values given in Supplementary Table 3 represent mean + /-SD. b, KM/vmax measurements for different phosphorylation reactions were performed in biological triplicates for S585 as well as S588 and duplicates for S591. In all subpanels single experiments are shown individually (see also Fig. 6b and Supplementary Tables 4 and 5). Values given in Supplementary Table 5 represent mean + /-SD. c–e, Linearization curves to account for nonlinear ionization behavior of PAD peptides with different phosphorylation states. Linearization curves were determined once.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Tables 1–5.
Rights and permissions
About this article
Cite this article
Gebel, J., Tuppi, M., Chaikuad, A. et al. p63 uses a switch-like mechanism to set the threshold for induction of apoptosis. Nat Chem Biol 16, 1078–1086 (2020). https://doi.org/10.1038/s41589-020-0600-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0600-3
This article is cited by
-
Fuzzy interactions between the auto-phosphorylated C-terminus and the kinase domain of CK1δ inhibits activation of TAp63α
Scientific Reports (2023)
-
Molecular basis for integrin adhesion receptor binding to p21-activated kinase 4 (PAK4)
Communications Biology (2022)
-
Structural diversity of p63 and p73 isoforms
Cell Death & Differentiation (2022)
-
Designed Ankyrin Repeat Proteins as a tool box for analyzing p63
Cell Death & Differentiation (2022)
-
Oocyte Casein kinase 1α deletion causes defects in primordial follicle formation and oocyte loss by impairing oocyte meiosis and enhancing autophagy in developing mouse ovary
Cell Death Discovery (2022)