Abstract
In proteins where conformational changes are functionally important, the number of accessible states and their dynamics are often difficult to establish. Here we describe a novel 19F-NMR spectroscopy approach to probe dynamics of large membrane proteins. We labeled a glutamate transporter homolog with a 19F probe via cysteine chemistry and with a Ni2+ ion via chelation by a di-histidine motif. We used distance-dependent enhancement of the longitudinal relaxation of 19F nuclei by the paramagnetic metal to assign the observed resonances. We identified one inward- and two outward-facing states of the transporter, in which the substrate-binding site is near the extracellular and intracellular solutions, respectively. We then resolved the structure of the unanticipated second outward-facing state by cryo-EM. Finally, we showed that the rates of the conformational exchange are accessible from measurements of the metal-enhanced longitudinal relaxation of 19F nuclei.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Atomic coordinates for the cryo-EM structures of the OFS and iOFS states have been deposited in the Protein Data Bank under accession codes 6UWF and 6UWL, respectively, and the corresponding cryo-EM maps have been deposited in the Electron Microscopy Data Bank under accession codes EMD-20922 and EMD-20923, respectively. The other data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Palmer, A. G. Enzyme dynamics from NMR spectroscopy. Acc. Chem. Res. 48, 457–465 (2015).
Pervushin, K., Riek, R., Wider, G. & Wuthrich, K. Attenuated T 2 relaxation by mutual cancellation of dipole–dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl Acad. Sci. USA 94, 12366–12371 (1997).
Jiang, Y. & Kalodimos, C. G. NMR studies of large proteins. J. Mol. Biol. 429, 2667–2676 (2017).
Danielson, M. A. & Falke, J. J. Use of 19F NMR to probe protein structure and conformational changes. Annu. Rev. Biophys. Biomol. Struct. 25, 163–195 (1996).
Susac, L., Eddy, M. T., Didenko, T., Stevens, R. C. & Wuthrich, K. A2A adenosine receptor functional states characterized by 19F-NMR. Proc. Natl Acad. Sci. USA 115, 12733–12738 (2018).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Di Pietrantonio, C., Pandey, A., Gould, J., Hasabnis, A. & Prosser, R. S. Understanding protein function through an ensemble description: characterization of functional states by 19F NMR. Methods Enzymol. 615, 103–130 (2019).
Yernool, D., Boudker, O., Folta-Stogniew, E. & Gouaux, E. Trimeric subunit stoichiometry of the glutamate transporters from Bacillus caldotenax and Bacillus stearothermophilus. Biochemistry 42, 12981–12988 (2003).
Danbolt, N. C. Glutamate uptake. Prog. Neurobiol. 65, 1–105 (2001).
Zerangue, N. & Kavanaugh, M. P. Flux coupling in a neuronal glutamate transporter. Nature 383, 634–637 (1996).
Levy, L. M., Warr, O. & Attwell, D. Stoichiometry of the glial glutamate transporter GLT-1 expressed inducibly in a Chinese hamster ovary cell line selected for low endogenous Na-dependent glutamate uptake. J. Neurosci. 18, 9620–9628 (1998).
Akyuz, N., Altman, R. B., Blanchard, S. C. & Boudker, O. Transport dynamics in a glutamate transporter homologue. Nature 502, 114–118 (2013).
Erkens, G. B., Hänelt, I., Goudsmits, J. M. H., Slotboom, D. J. & van Oijen, A. M. Unsynchronised subunit motion in single trimeric sodium-coupled aspartate transporters. Nature 502, 119–123 (2013).
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wuthrich, K. Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 335, 1106–1110 (2012).
Clore, G. M. & Iwahara, J. Theory, practice, and applications of paramagnetic relaxation enhancement for the characterization of transient low-population states of biological macromolecules and their complexes. Chem. Rev. 109, 4108–4139 (2009).
Bondarenko, V. et al. 19F paramagnetic relaxation-based NMR for quaternary structural restraints of ion channels. ACS Chem. Biol. 14, 2160–2165 (2019).
Solomon, I. Relaxation processes in a system of two spins. Phys. Rev. 99, 559–565 (1955).
Bloembergen, N. Proton relaxation times in paramagnetic solutions. J. Chem. Phys. 27, 572–573 (1957).
Matei, E. & Gronenborn, A. M. 19F paramagnetic relaxation enhancement: a valuable tool for distance measurements in proteins. Angew. Chem. Int. Ed. 55, 150–154 (2016).
Hull, W. E. & Sykes, B. D. Fluorotyrosine alkaline phosphatase: internal mobility of individual tyrosines and the role of chemical shift anisotropy as a 19F nuclear spin relaxation mechanism in proteins. J. Mol. Biol. 98, 121–153 (1975).
Hull, W. E. & Sykes, B. D. Dipolar nuclear spin relaxation of 19F in multispin systems. Application to 19F labeled proteins. J. Chem. Phys. 63, 867–880 (1975).
Gerig, J. T. Fluorine-proton Overhauser effects in fluorine-labeled macromolecular systems. J. Am. Chem. Soc. 99, 1721–1725 (1977).
Todd, R. J., Van Dam, M. E., Casimiro, D., Haymore, B. L. & Arnold, F. H. Cu(ii)-binding properties of a cytochrome c with a synthetic metal-binding site: His-X3-His in an α-helix. Proteins 10, 156–161 (1991).
Didenko, T., Liu, J. J., Horst, R., Stevens, R. C. & Wuthrich, K. Fluorine-19 NMR of integral membrane proteins illustrated with studies of GPCRs. Curr. Opin. Struct. Biol. 23, 740–747 (2013).
McIlwain, B. C., Vandenberg, R. J. & Ryan, R. M. Characterization of the inward- and outward-facing substrate binding sites of the prokaryotic aspartate transporter, GltPh. Biochemistry 55, 6801–6810 (2016).
Akyuz, N. et al. Transport domain unlocking sets the uptake rate of an aspartate transporter. Nature 518, 68–73 (2015).
Ruan, Y. et al. Direct visualization of glutamate transporter elevator mechanism by high-speed AFM. Proc. Natl Acad. Sci. USA 114, 1584–1588 (2017).
Georgieva, E. R., Borbat, P. P., Ginter, C., Freed, J. H. & Boudker, O. Conformational ensemble of the sodium-coupled aspartate transporter. Nat. Struct. Mol. Biol. 20, 215–221 (2013).
Reyes, N., Oh, S. & Boudker, O. Binding thermodynamics of a glutamate transporter homolog. Nat. Struct. Mol. Biol. 20, 634–640 (2013).
Boudker, O., Ryan, R. M., Yernool, D., Shimamoto, K. & Gouaux, E. Coupling substrate and ion binding to extracellular gate of a sodium-dependent aspartate transporter. Nature 445, 387–393 (2007).
Suh, S.-S., Haymore, B. L. & Arnold, F. H. Characterization of His-X3-His sites in α-helices of synthetic metal-binding bovine somatotropin. Protein Eng. Des. Sel. 4, 301–305 (1991).
Verdon, G. & Boudker, O. Crystal structure of an asymmetric trimer of a bacterial glutamate transporter homolog. Nat. Struct. Mol. Biol. 19, 355–357 (2012).
Machtens, J. P. et al. Mechanisms of anion conduction by coupled glutamate transporters. Cell 160, 542–553 (2015).
Cavanagh, J., Fairbrother, W. J., Palmer, A. G., Rance, M. & Skelton, N. J. in Protein NMR Spectroscopy 2nd edn 333–404 (Academic Press, 2007).
Mayer, M. & Meyer, B. Characterization of ligand binding by saturation transfer difference NMR spectroscopy. Angew. Chem. Int. Ed. 38, 1784–1788 (1999).
Spoerner, M. et al. Conformational states of human rat sarcoma (Ras) protein complexed with its natural ligand GTP and their role for effector interaction and GTP hydrolysis. J. Biol. Chem. 285, 39768–39778 (2010).
Reyes, N., Ginter, C. & Boudker, O. Transport mechanism of a bacterial homologue of glutamate transporters. Nature 462, 880–885 (2009).
Ye, L., Van Eps, N., Zimmer, M., Ernst, O. P. & Prosser, R. S. Activation of the A2A adenosine G-protein-coupled receptor by conformational selection. Nature 533, 265–268 (2016).
Yernool, D., Boudker, O., Jin, Y. & Gouaux, E. Structure of a glutamate transporter homologue from Pyrococcus horikoshii. Nature 431, 811–818 (2004).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Jo, S., Lim, J. B., Klauda, J. B. & Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58 (2009).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Shi, L., Quick, M., Zhao, Y., Weinstein, H. & Javitch, J. A. The mechanism of a neurotransmitter:sodium symporter—inward release of Na+ and substrate is triggered by substrate in a second binding site. Mol. Cell 30, 667–677 (2008).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
McGibbon, RobertT. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Ritchie, T. K. et al. Reconstitution of membrane proteins in phospholipid bilayer nanodiscs. Methods Enzymol. 464, 211–231 (2009).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, 42166 (2018).
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Afonine, P. V. et al. phenix.model_vs_data: a high-level tool for the calculation of crystallographic model and data statistics. J. Appl. Crystallogr. 43, 669–676 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We thank X. Yao for helpful discussions, M. Goger and S. Bhattacharya for help with setting up NMR and M.A. Cuendet for help with setting up initial MD simulations. We thank C. Xu and K. Song at the UMass cryo-EM facility for help with electron microscopy data collection. We also thank W. Eng for assistance with protein expression. This work was supported by NIH grants R37NS085318 and R01NS064357 (O.B.), R37AG019391 (D.E.) and S10OD016320 (C.B.). O.B., D.E. and C.B. are members of the New York Structural Biology Center (NYSBC), which is supported in part by NIH grant P41 GM118302 (CoMD/NMR, Center on Macromolecular Dynamics by NMR Spectroscopy), ORIP/NIH facility improvement grant CO6RR015495 and NIH grant S10OD018509. The coordinates of the structures and the density maps have been submitted to the Protein Data Bank under the accession codes of 6UWF and 6UWL.
Author information
Authors and Affiliations
Contributions
Y.H., D.E. and O.B. designed the experiments. Y.H. and G.L. performed the NMR experiments. Y.H. and O.B analyzed the data. C.B. assisted with NMR experimental design, data collection and analysis. X.W. performed cryo-EM imaging, analyzed data and refined molecular models. G.H.M.H. performed transport activity assays. A.M.R. performed molecular dynamics simulations. Y.H., X.W., A.M.R., O.B., D.E., H.W. and C.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Protein purity, and 3H L-asp uptake and simulated 19F-Ni2+ distance distribution.
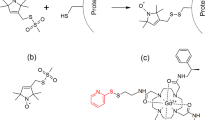
a, Scheme for site-specifically introducing 19F label into M385C GltPh mutant. b, Representative size exclusion chromatography elution profile of M385C-TET GltPh. More than 3 independent samples were repeated with similar results. c, SDS-PAGE gel imaged by Coommasie blue staining (middle) and fluorescence (right) and of M385C GltPh labeled with fluorescein-5-malaimide before (lane 1) and after (lane 2) labeling with TFET. Protein samples were incubated with 10-fold excess of fluorescein-5-maleimide for 4 h prior to analysis. Two independent samples were prepared and yielded similar results. d. Michaelis-Menten kinetics of 3H L-Asp uptake for wide type GltPh (black circle), M385C-TET (red square), and dHis/M385C-TET GltPh (blue triangle). Data shown are means ± s.d. (N = 3 biological replicates). e, Distance probability distributions between 19F and Ni2+ calculated from 100 ns of the molecular dynamics simulation trajectories. To mimic experimental conditions M385 was mutated to NMR probe TET, residues 215 and 219 were mutated to histidine, and Zn2+ ion was constrained between these histidines (see Online Methods for details). The distance distributions were calculated for all three protomers and shown curves are averaged values from three protomers in the OFS (left) and the IFS (right). DTDP: 2,2’-dithiodipyridine; TFET: trifluoroethanethiol.
Extended Data Fig. 2 Specific Ni2+ binding to dHis/M385C-TET GltPh mutant.
1D 19F-NMR spectra of M385C-TET GltPh (a) and dHis/M385C-TET GltPh (b) without (up) and with (bottom) 3 molar equivalents of Ni2+ ions. Spectra were recorded at 293 K in the presence of NaCL and L-asp. Raw data are black, fits are magenta and deconvoluted peaks are blue. Note: the dHis/M385C-TET spectra are the same as the ones shown in Fig. 2a and Fig. 3a in the main text.
Extended Data Fig. 3 Cryo-EM data processing.
a, Angular distribution of particles contributing to the final reconstitution. Number of views at each angular orientation is represented by length and color of cylinders where red indicates more views. b, Final maps after Relion post-processing colored according to local resolution estimation using ResMap. c, Fourier shell correlation (FSC) curves indicating the resolution at the 0.143 threshold of final masked (black) and unmasked (orange) maps of GltPh OFS (left) and iOFS (right). d, FSC curves from cross-validation of refined GltPh OFS (left) and iOFS (right) models compared to the masked half-map 1 (Orange traces: FSCwork, used during validation refinement), masked half-map 2 (Blue traces: FSCfree, not used during validation refinement), and the masked summed map (Black traces: FSCsum). e, Data processing flow chart for GltPh reconstituted into nanodisc in the presence of NaCl and L-asp.
Extended Data Fig. 4 2D 19F EXSY spectrum of dHis/M385C-TET GltPh.
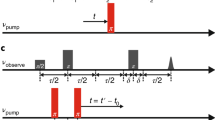
Spectrum was recorded with mixing time of 0.4 s in the presence of 200 mM Na+ and 10 µM L-asp at 298 K.
Supplementary information
Supplementary Information
Supplementary Note, Fig. 1 and Tables 1–2.
Source data
Source Data Extended Data Fig. 1
Unprocessed SDS-PAGE gels.
Rights and permissions
About this article
Cite this article
Huang, Y., Wang, X., Lv, G. et al. Use of paramagnetic 19F NMR to monitor domain movement in a glutamate transporter homolog. Nat Chem Biol 16, 1006–1012 (2020). https://doi.org/10.1038/s41589-020-0561-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0561-6
This article is cited by
-
Expanding the membrane-protein NMR toolkit
Nature Chemical Biology (2020)
-
Millisecond dynamics of an unlabeled amino acid transporter
Nature Communications (2020)
-
A suite of 19F based relaxation dispersion experiments to assess biomolecular motions
Journal of Biomolecular NMR (2020)