Abstract
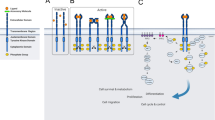
Receptor tyrosine kinases (RTKs) are transmembrane receptors of great clinical interest due to their role in disease. Historically, therapeutics targeting RTKs have been identified using in vitro kinase assays. Due to frequent development of drug resistance, however, there is a need to identify more diverse compounds that inhibit mutated but not wild-type RTKs. Here, we describe MaMTH-DS (mammalian membrane two-hybrid drug screening), a live-cell platform for high-throughput identification of small molecules targeting functional protein–protein interactions of RTKs. We applied MaMTH-DS to an oncogenic epidermal growth factor receptor (EGFR) mutant resistant to the latest generation of clinically approved tyrosine kinase inhibitors (TKIs). We identified four mutant-specific compounds, including two that would not have been detected by conventional in vitro kinase assays. One of these targets mutant EGFR via a new mechanism of action, distinct from classical TKI inhibition. Our results demonstrate how MaMTH-DS is a powerful complement to traditional drug screening approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings presented in this study are available in the paper and its Supplementary Information files.
Code availability
All R code used in the analysis of the presented drug screening data is available from the authors on request.
References
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Volinsky, N. & Kholodenko, B. N. Complexity of receptor tyrosine kinase signal processing. Cold Spring Harb. Perspect. Biol. 5, a009043–a009043 (2013).
Jia, Y., Quinn, C. M., Kwak, S. & Talanian, R. V. Current in vitro kinase assay technologies: the quest for a universal format. Curr. Drug Discov. Technol. 5, 59–69 (2008).
Petschnigg, J. et al. The mammalian-membrane two-hybrid assay (MaMTH) for probing membrane-protein interactions in human cells. Nat. Methods 11, 585–592 (2014).
Sokolina, K. et al. Systematic protein–protein interaction mapping for clinically relevant human GPCRs. Mol. Syst. Biol. 13, 918 (2017).
Snider, J. et al. Mapping the functional yeast ABC transporter interactome. Nat. Chem. Biol. 9, 565–572 (2013).
Stagljar, I., Korostensky, C., Johnsson, N. & te Heesen, S. A genetic system based on split-ubiquitin for the analysis of interactions between membrane proteins in vivo. Proc. Natl Acad. Sci. USA 95, 5187–5192 (1998).
Yao, Z. et al. A global analysis of the receptor tyrosine kinase-protein phosphatase interactome. Mol. Cell 65, 347–360 (2017).
Petschnigg, J. et al. Systematic identification of oncogenic EGFR interaction partners. J. Mol. Biol. 429, 280–294 (2017).
Robbins, A. K. & Horlick, R. A. Macrophage scavenger receptor confers an adherent phenotype to cells in culture. Biotechniques 25, 240–244 (1998).
Zheng, Y. et al. Temporal regulation of EGF signalling networks by the scaffold protein Shc1. Nature 499, 166–171 (2013).
Saucier, C. et al. The Shc adaptor protein is critical for VEGF induction by Met/HGF and ErbB2 receptors and for early onset of tumor angiogenesis. Proc. Natl Acad. Sci. USA 101, 2345–2350 (2004).
Goździk-Spychalska, J. et al. C-MET inhibitors in the treatment of lung cancer. Curr. Treat. Options Oncol. 15, 670–682 (2014).
Hagel, M. et al. First selective small molecule inhibitor of FGFR4 for the treatment of hepatocellular carcinomas with an activated FGFR4 signaling pathway. Cancer Discov. 5, 424–437 (2015).
Eder, J. P. et al. A phase I study of foretinib, a multi-targeted inhibitor of c-Met and vascular endothelial growth factor receptor 2. Clin. Cancer Res. 16, 3507–3516 (2010).
Qian, F. et al. Inhibition of tumor cell growth, invasion, and metastasis by EXEL-2880 (XL880, GSK1363089), a novel inhibitor of HGF and VEGF receptor tyrosine kinases. Cancer Res. 69, 8009–8016 (2009).
Zhang, S. et al. The potent ALK inhibitor brigatinib (AP26113) overcomes mechanisms of resistance to first- and second-generation ALK inhibitors in preclinical models. Clin. Cancer Res. 22, 5527–5538 (2016).
Walter, A. O. et al. Discovery of a mutant-selective covalent inhibitor of EGFR that overcomes T790M-mediated resistance in NSCLC. Cancer Discov. 3, 1404–1415 (2013).
Jiang, T. & Zhou, C. Clinical activity of the mutant-selective EGFR inhibitor AZD9291 in patients with EGFR inhibitor-resistant non-small cell lung cancer. Transl. Lung Cancer Res. 3, 370–372 (2014).
Thress, K. S. et al. Acquired EGFR C797S mutation mediates resistance to AZD9291 in non-small cell lung cancer harboring EGFR T790M. Nat. Med. 21, 560–562 (2015).
Zhang, J.-H., Chung, T. D. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Brideau, C., Gunter, B., Pikounis, B. & Liaw, A. Improved statistical methods for hit selection in high-throughput screening. J. Biomol. Screen. 8, 634–647 (2003).
Lee, H. J. et al. Noncovalent wild-type-sparing inhibitors of EGFR T790M. Cancer Discov. 3, 168–181 (2013).
M., L. Midostaurin approved for FLT3-mutated AML. Blood 129, 3403–3406 (2017).
Galanis, A. et al. Crenolanib is a potent inhibitor of FLT3 with activity against resistance-conferring point mutants. Blood 123, 94–100 (2014).
Antar, A. et al. Inhibition of FLT3 in AML: a focus on sorafenib. Bone Marrow Transplant. 52, 344–351 (2017).
Cortes, J. et al. Quizartinib, an FLT3 inhibitor, as monotherapy in patients with relapsed or refractory acute myeloid leukaemia: an open-label, multicentre, single-arm, phase 2 trial. Lancet Oncol. 19, 889–903 (2018).
Perl, A. E. et al. Selective inhibition of FLT3 by gilteritinib in relapsed or refractory acute myeloid leukaemia: a multicentre, first-in-human, open-label, phase 1–2 study. Lancet Oncol. 18, 1061–1075 (2017).
Zabludoff, S. D. et al. AZD7762, a novel checkpoint kinase inhibitor, drives checkpoint abrogation and potentiates DNA-targeted therapies. Mol. Cancer Ther. 7, 2955–2966 (2008).
Sausville, E. et al. Phase I dose-escalation study of AZD7762, a checkpoint kinase inhibitor, in combination with gemcitabine in US patients with advanced solid tumors. Cancer Chemother. Pharmacol. 73, 539–549 (2014).
Kim, S. N. et al. 7-Diethylamino-3(2′-benzoxazolyl)-coumarin is a novel microtubule inhibitor with antimitotic activity in multidrug resistant cancer cells. Biochem. Pharmacol. 77, 1773–1779 (2009).
Mellman, I. & Yarden, Y. Endocytosis and cancer. Cold Spring Harb. Perspect. Biol. 5, a016949 (2013).
Villaseñor, R., Nonaka, H., Del Conte-Zerial, P., Kalaidzidis, Y. & Zerial, M. Regulation of EGFR signal transduction by analogue-to-digital conversion in endosomes. eLife 4, e06156 (2015).
Collinet, C. et al. Systems survey of endocytosis by multiparametric image analysis. Nature 464, 243–249 (2010).
Rink, J., Ghigo, E., Kalaidzidis, Y. & Zerial, M. Rab conversion as a mechanism of progression from early to late endosomes. Cell 122, 735–749 (2005).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659–661 (2011).
R: a language and environment for statistical computing (R Core Team, 2017).
Box, G. E. P. & Cox, D. R. An analysis of transformations. R. Stat. Soc. 26, 211–252 (1964).
Boutros, M., Brás, L. P. & Huber, W. Analysis of cell-based RNAi screens. Genome Biol. 7, R66 (2006).
Aher, A. et al. CLASP suppresses microtubule catastrophes through a single TOG domain. Dev. Cell 46, 40–58.e8 (2018).
Mohan, R. et al. End-binding proteins sensitize microtubules to the action of microtubule-targeting agents. Proc. Natl Acad. Sci. USA 110, 8900–8905 (2013).
Jost, M. et al. Combined CRISPRi/a-based chemical genetic screens reveal that rigosertib is a microtubule-destabilizing agent. Mol. Cell 68, 210–223 (2017).
Acknowledgements
We thank L. Riley for valuable discussions and editing of this manuscript. We also thank P.A. Jänne and M.J. Eck (both at Dana Farber Cancer Institute, Harvard Medical School, Boston, MA, USA) for providing us with the PC9-T790M and PC9-C797S cells used in our study, and the Center for Information Services and High Performance Computing of the TU Dresden for providing support and computational resources. This research was supported by funding from Consortium Québécois sur la Découverte du Médicament (CQDM) (Explore) and OCE (no. 23929). In addition, work in the Stagljar laboratory is supported by the Canadian Cancer Society Research Institute (no. 703889), Genome Canada via Ontario Genomics (grant nos, 9427 and 9428), Ontario Research fund (grant nos. ORF/DIG-501411 and RE08-009), CQDM (Quantum Leap) and Brain Canada (Quantum Leap, and Cancer Research Society (grant no. 23235)). J.H.P.’s work in the laboratory of Mark Lemmon at Yale was supported by National Institutes of Health grant nos. R01-CA198164 and R35-GM122485. G.P. acknowledges the support of the Ontario Institute for Cancer Research and its funding from the Government of Ontario.
Author information
Authors and Affiliations
Contributions
P.S. performed preliminary MaMTH testing and validation/functional analysis of identified compounds, managed the project and wrote the manuscript. J.S. generated MaMTH reporter constructs and cells lines, performed MaMTH-DS screening and data analysis, managed the project and wrote the manuscript. Y.K. and M.Z. performed and analyzed the high-throughput EGFR microscopy data. K.W., L.D., A.L., N.V., A.V., S.P., F.A., Z.Y. and V.W. performed construct generation, preliminary MaMTH validation and were involved in the validation/functional analysis of compounds. N.R., N.A.P., Y.W., A.S. and M.S.T. generated and validated organoids and provided cell lines. A.R. and A.A. performed and analyzed microtubule dynamics data. L.E.W.G., A.D. and J.W. performed MaMTH-DS screening. B.T., A.A., M.P., B.J., R.M., D.U. and R.A. performed medicinal chemistry and assisted technically throughout the whole project. G.P. performed computational chemistry, molecular docking and hit expansion. J.H.P. assisted with computational modeling and undertook some biochemical studies. R.M.S., M.J., M.S., M.F.M., N.L. and F.A.S. provided clinical expertise and support. All authors discussed the results and commented on the manuscript. I.S. initiated and supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
I.S., P.S. and J.S. (in conjunction with the University of Toronto) are listed as inventors on a patent (publication no. 20190091205) for the use of EMI1 (and structurally related analogs), midostaurin, gilteritinib and AZD7762 (and structurally related analogs) in the treatment of mutant EGFR-mediated NSCLC.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–32, Tables 1 and 2 and Notes 1 and 2
Supplementary Dataset 1
Collection of compounds used in MaMTH-DS pilot screening.
Supplementary Dataset 2
Results of MaMTH-DS screening of EGFR L858R/T790M/C797S in the presence of ShcI (round 1).
Supplementary Dataset 3
Results of MaMTH-DS screening of EGFR L858R/T790M/C797S in the presence of ShcI (round 2).
Rights and permissions
About this article
Cite this article
Saraon, P., Snider, J., Kalaidzidis, Y. et al. A drug discovery platform to identify compounds that inhibit EGFR triple mutants. Nat Chem Biol 16, 577–586 (2020). https://doi.org/10.1038/s41589-020-0484-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0484-2
This article is cited by
-
The diversification of methods for studying cell–cell interactions and communication
Nature Reviews Genetics (2024)
-
Programmable synthetic receptors: the next-generation of cell and gene therapies
Signal Transduction and Targeted Therapy (2024)
-
Receptor tyrosine kinases and cancer: oncogenic mechanisms and therapeutic approaches
Oncogene (2021)
-
A homogeneous split-luciferase assay for rapid and sensitive detection of anti-SARS CoV-2 antibodies
Nature Communications (2021)