Abstract
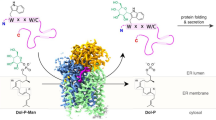
Tryptophan C-mannosylation is an unusual co-translational protein modification performed by metazoans and apicomplexan protists. The prevalence and biological functions of this modification are poorly understood, with progress in the field hampered by a dearth of convenient tools for installing and detecting the modification. Here, we engineer a yeast system to produce a diverse array of proteins with and without tryptophan C-mannosylation and interrogate the modification’s influence on protein stability and function. This system also enabled mutagenesis studies to identify residues of the glycosyltransferase and its protein substrates that are crucial for catalysis. The collection of modified proteins accrued during this work facilitated the generation and thorough characterization of monoclonal antibodies against tryptophan C-mannosylation. These antibodies empowered proteomic analyses of the brain C-glycome by enriching for peptides possessing tryptophan C-mannosylation. This study revealed many new modification sites on proteins throughout the secretory pathway with both conventional and non-canonical consensus sequences.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All relevant data are available from the authors upon request. Structure coordinates have been deposited in the Protein Data Bank (https://www.rcsb.org/) under accession code 6PLH. Proteomics data are available via ProteomeXchange60 (http://www.proteomexchange.org/) with identifiers PXD018401, PXD020336 and PXD021612. Some data were sourced from the UniProt (https://www.uniprot.org/) database (accession numbers D3YXG0, P11680, P59511, Q3UHD1, Q3UPR9, Q62217, Q69ZU6, Q6P4U0, Q80TI0, Q8VDX6, Q8VCC9). Source data are provided with this paper.
References
Spiro, R. G. Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12, 43r–56r (2002).
de Beer, T., Vliegenthart, J. F., Löffler, A. & Hofsteenge, J. The hexopyranosyl residue that is C-glycosidically linked to the side chain of tryptophan-7 in human RNase Us is α-mannopyranose. Biochemistry 34, 11785–11789 (1995).
Hofsteenge, J. et al. New type of linkage between a carbohydrate and a protein: C-glycosylation of a specific tryptophan residue in human RNase Us. Biochemistry 33, 13524–13530 (1994).
Shcherbakova, A., Tiemann, B., Buettner, F. F. & Bakker, H. Distinct C-mannosylation of netrin receptor thrombospondin type 1 repeats by mammalian DPY19L1 and DPY19L3. Proc. Natl Acad. Sci. USA 114, 2574–2579 (2017).
Doucey, M. A., Hess, D., Cacan, R. & Hofsteenge, J. Protein C-mannosylation is enzyme-catalysed and uses dolichyl-phosphate-mannose as a precursor. Mol. Biol. Cell 9, 291–300 (1998).
Buettner, F. F., Ashikov, A., Tiemann, B., Lehle, L. & Bakker, H. C. elegans DPY-19 is a C-mannosyltransferase glycosylating thrombospondin repeats. Mol. Cell 50, 295–302 (2013).
Pronker, M. F. et al. Structural basis of myelin-associated glycoprotein adhesion and signalling. Nat. Commun. 7, 13584 (2016).
Goto, Y. et al. C-mannosylation of human hyaluronidase 1: possible roles for secretion and enzymatic activity. Int. J. Oncol. 45, 344–350 (2014).
Okamoto, S. et al. Regulation of secretion and enzymatic activity of lipoprotein lipase by C-mannosylation. Biochem. Biophys. Res. Commun. 486, 558–563 (2017).
Falzarano, D. et al. Ebola sGP—the first viral glycoprotein shown to be C-mannosylated. Virology 368, 83–90 (2007).
Hofsteenge, J., Blommers, M., Hess, D., Furmanek, A. & Miroshnichenko, O. The four terminal components of the complement system are C-mannosylated on multiple tryptophan residues. J. Biol. Chem. 274, 32786–32794 (1999).
Lovelace, L. L., Cooper, C. L., Sodetz, J. M. & Lebioda, L. Structure of human C8 protein provides mechanistic insight into membrane pore formation by complement. J. Biol. Chem. 286, 17585–17592 (2011).
Hofsteenge, J. et al. C-mannosylation and O-fucosylation of the thrombospondin type 1 module. J. Biol. Chem. 276, 6485–6498 (2001).
Hartmann, S. & Hofsteenge, J. Properdin, the positive regulator of complement, is highly C-mannosylated. J. Biol. Chem. 275, 28569–28574 (2000).
Sorvillo, N. et al. Identification of N-linked glycosylation and putative O-fucosylation, C-mannosylation sites in plasma derived ADAMTS13. J. Thromb. Haemost. 12, 670–679 (2014).
Furmanek, A., Hess, D., Rogniaux, H. & Hofsteenge, J. The WSAWS motif is C-hexosylated in a soluble form of the erythropoietin receptor. Biochemistry 42, 8452–8458 (2003).
Hamming, O. J. et al. Crystal structure of interleukin-21 receptor (IL-21R) bound to IL-21 reveals that sugar chain interacting with WSXWS motif is integral part of IL-21R. J. Biol. Chem. 287, 9454–9460 (2012).
Sasazawa, Y., Sato, N., Suzuki, T., Dohmae, N. & Simizu, S. C-Mannosylation of thrombopoietin receptor (c-Mpl) regulates thrombopoietin-dependent JAK–STAT signaling. Biochem. Biophys. Res. Commun. 468, 262–268 (2015).
Doucey, M. A., Hess, D., Blommers, M. J. & Hofsteenge, J. Recombinant human interleukin-12 is the second example of a C-mannosylated protein. Glycobiology 9, 435–441 (1999).
Wang, L. W., Leonhard-Melief, C., Haltiwanger, R. S. & Apte, S. S. Post-translational modification of thrombospondin type-1 repeats in ADAMTS-like 1/punctin-1 by C-mannosylation of tryptophan. J. Biol. Chem. 284, 30004–30015 (2009).
Niwa, Y., Suzuki, T., Dohmae, N. & Simizu, S. Identification of DPY19L3 as the C-mannosyltransferase of R-spondin1 in human cells. Mol. Biol. Cell 27, 744–756 (2016).
Morishita, S., Suzuki, T., Niwa, Y., Dohmae, N. & Simizu, S. Dpy-19 like 3-mediated C-mannosylation and expression levels of RPE-spondin in human tumor cell lines. Oncol. Lett. 14, 2537–2544 (2017).
Fujiwara, M. et al. C-mannosylation of R-spondin3 regulates its secretion and activity of Wnt/β-catenin signaling in cells. FEBS Lett. 590, 2639–2649 (2016).
Li, Y. et al. Structure of the F-spondin domain of mindin, an integrin ligand and pattern recognition molecule. EMBO J. 28, 286–297 (2009).
Ling, J., Su, J., Ma, Z. & Ruan, C. The WXXW motif in the TSR1 of ADAMTS13 is important for its secretion and proteolytic activity. Thromb. Res. 131, 529–534 (2013).
Shcherbakova, A. et al. C-mannosylation supports folding and enhances stability of thrombospondin repeats. eLife 8, e52978 (2019).
Olsen, J. G. & Kragelund, B. B. Who climbs the tryptophan ladder? On the structure and function of the WSXWS motif in cytokine receptors and thrombospondin repeats. Cytokine Growth Factor Rev. 25, 337–341 (2014).
Craven, T. W., Cho, M. K., Traaseth, N. J., Bonneau, R. & Kirshenbaum, K. A miniature protein stabilized by a cation-π interaction network. J. Am. Chem. Soc. 138, 1543–1550 (2016).
Hoppe, C. M. et al. Apicomplexan C-mannosyltransferases modify thrombospondin type I-containing adhesins of the TRAP family. Glycobiology 28, 333–343 (2018).
Niwa, Y. et al. Topological analysis of DPY19L3, a human C-mannosyltransferase. FEBS J. 285, 1162–1174 (2018).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Krieg, J. et al. Recognition signal for C-mannosylation of Trp-7 in RNase 2 consists of sequence Trp-x-x-Trp. Mol. Biol. Cell 9, 301–309 (1998).
Albuquerque-Wendt, A., Hütte, H. J., Buettner, F. F. R., Routier, F. H. & Bakker, H. Membrane topological model of glycosyltransferases of the GT-C superfamily. Int. J. Mol. Sci. 20, 4842 (2019).
Ihara, Y. et al. Increased expression of protein C-mannosylation in the aortic vessels of diabetic Zucker rats. Glycobiology 15, 383–392 (2005).
Mou, Y. et al. Engineering improved antiphosphotyrosine antibodies based on an immunoconvergent binding motif. J. Am. Chem. Soc. 140, 16615–16624 (2018).
Piechotta, A. et al. Structural and functional analyses of pyroglutamate-amyloid-β-specific antibodies as a basis for Alzheimer immunotherapy. J. Biol. Chem. 292, 12713–12724 (2017).
Munte, C. E. et al. C-mannosylation in the hypertrehalosaemic hormone from the stick insect Carausius morosus. FEBS J. 275, 1163–1173 (2008).
Jonker, H. R. A. et al. NMR spectroscopic characterization of the C-mannose conformation in a thrombospondin repeat using a selective labeling approach. Angew. Chem. Int. Ed. Engl. 59, 20659–20665 (2020).
Sekula, P. et al. From discovery to translation: characterization of C-mannosyltryptophan and pseudouridine as markers of kidney function. Sci. Rep. 7, 17400 (2017).
Manya, H. et al. The muscular dystrophy gene TMEM5 encodes a ribitol β1,4-xylosyltransferase required for the functional glycosylation of dystroglycan. J. Biol. Chem. 291, 24618–24627 (2016).
Jae, L. T. et al. Deciphering the glycosylome of dystroglycanopathies using haploid screens for Lassa virus entry. Science 340, 479–483 (2013).
Sandhu, J. et al. Aster proteins facilitate nonvesicular plasma membrane to ER cholesterol transport in mammalian cells. Cell 175, 514–529 (2018).
Laraia, L. et al. The cholesterol transfer protein GRAMD1A regulates autophagosome biogenesis. Nat. Chem. Biol. 15, 710–720 (2019).
De Wachter, C., Van Landuyt, L. & Callewaert, N. Engineering of yeast glycoprotein expression. in Advances in Biochemical Engineering/Biotechnology 1–43 (Springer, 2018); https://doi.org/10.1007/10_2018_69
Park, D. et al. BAI1 is an engulfment receptor for apoptotic cells upstream of the ELMO/Dock180/Rac module. Nature 450, 430–434 (2007).
Das, S. et al. Brain angiogenesis inhibitor 1 (BAI1) is a pattern recognition receptor that mediates macrophage binding and engulfment of Gram-negative bacteria. Proc. Natl Acad. Sci. USA 108, 2136–2141 (2011).
He, Y. W. et al. The extracellular matrix protein mindin is a pattern-recognition molecule for microbial pathogens. Nat. Immunol. 5, 88–97 (2004).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Chen, Y., Zhang, J., Xing, G. & Zhao, Y. Mascot-derived false positive peptide identifications revealed by manual analysis of tandem mass spectra. J. Proteome Res. 8, 3141–3147 (2009).
Tripathy, D. R., Dinda, A. K. & Dasgupta, S. A simple assay for the ribonuclease activity of ribonucleases in the presence of ethidium bromide. Anal. Biochem. 437, 126–129 (2013).
Adamczyk, M., Gebler, J. C. & Wu, J. Papain digestion of different mouse IgG subclasses as studied by electrospray mass spectrometry. J. Immunol. Methods 237, 95–104 (2000).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Storoni, L. C., McCoy, A. J. & Read, R. J. Likelihood-enhanced fast rotation functions. Acta Crystallogr. D Biol. Crystallogr. 60, 432–438 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell Proteom. 13, 2513–2526 (2014).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank K. Wycherley, P. Masendycz and K. Mackwell at the Walter and Eliza Hall Institute Monoclonal Antibody Facility (WEHI-MAF) for their assistance in creating the mAbs reported in this paper. We also thank the beamline staff at the Australian Synchrotron for help with X-ray data collection as well as J. Newman and B. Marshall at the Commonwealth Scientific and Industrial Research Organisation (CSIRO) Collaborative Crystallisation Centre (C3) for assistance in protein crystallization. This research was undertaken, in part, using the MX2 beamline at the Australian Synchrotron, part of ANSTO, and made use of the Australian Cancer Research Foundation (ACRF) detector. E.D.G.-B. would like to acknowledge support from the Walter and Eliza Hall Institute of Medical Research, National Health and Medical Research Council of Australia (NHMRC) project grants GNT1139546 and GNT1139549, the ACRF, a Victorian State Government Operational Infrastructure support grant and the support of the Brian M. Davis Charitable Foundation Centenary Fellowship.
Author information
Authors and Affiliations
Contributions
A.J. performed all molecular biology, protein expression, biochemical assays and immunoprecipitations; M.A.J. performed the structural biology experiments; S.S. and R.M. assisted with the radioassays; R.W.B. and P.E.C. performed SPR; A.W.L. assisted with mAb sequencing; S.C. assisted with animal work; N.E.S. performed all MS experiments; E.D.G.-B. conceived the project; A.J. and E.D.G.-B. cowrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Expression of CeDPY19 mutants in P. pastoris.
Western blots demonstrating expression of all twelve CeDPY19 mutants in P. pastoris (n = 1 independent replicates).
Extended Data Fig. 2 Probing mAb selectivity by western blot.
Representative western blots using the collection of proteins with and without tryptophan C-mannosylation to map the epitope preferences of mAbs 10E9, 9E7, 6G4 and 7G12 (one of n = 3 independent replicates).
Extended Data Fig. 3 Determining mAb affinity by SPR.
Representative SPR sensorgrams used to determine the affinity of mAbs 6G4 and 7G12.
Extended Data Fig. 4 Stereo views of the glycopeptide in the 5G12:glycopeptide complex.
Two stereo views of the glycopeptide antigen with tryptophan C-mannosylation that is bound to 5G12 with a Fo-Fc omit map contoured at 1.5σ.
Extended Data Fig. 5 Detection of modified proteins in mouse brain extract.
A representative western blot of protein extracted from mouse brain using the 5G12 mAb (one of n = 3 independent replicates).
Extended Data Fig. 6 Localisation of the tryptophan C-mannosylation site in Rxylt1.
MS/MS spectra demonstrating the location of tryptophan C-mannosylation for Rxylt1, as well as a sequence alignment showing conservation of this site across the Euteleostomi clade.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Tables 1–5.
Supplementary Data 1
Unprocessed western blots for Supplementary Fig. 1c.
Supplementary Data 2
Unprocessed gels for Supplementary Fig. 3b.
Supplementary Data 3
Unprocessed gels for Supplementary Fig. 14b.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 3
Unprocessed western blots and gels.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
John, A., Järvå, M.A., Shah, S. et al. Yeast- and antibody-based tools for studying tryptophan C-mannosylation. Nat Chem Biol 17, 428–437 (2021). https://doi.org/10.1038/s41589-020-00727-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00727-w
This article is cited by
-
Oxidative cyclization reagents reveal tryptophan cation–π interactions
Nature (2024)
-
Structure, sequon recognition and mechanism of tryptophan C-mannosyltransferase
Nature Chemical Biology (2023)
-
Recent trends in glycoproteomics by characterization of intact glycopeptides
Analytical and Bioanalytical Chemistry (2023)
-
Glycoproteomics
Nature Reviews Methods Primers (2022)
-
Tryptophan C-mannosylation is critical for Plasmodium falciparum transmission
Nature Communications (2022)