Abstract
Autophagy is an essential cellular process that removes harmful protein species, and autophagy upregulation may be able to protect against neurodegeneration and various pathogens. Here, we have identified the essential protein VCP/p97 (VCP, valosin-containing protein) as a novel regulator of autophagosome biogenesis, where VCP regulates autophagy induction in two ways, both dependent on Beclin-1. Utilizing small-molecule inhibitors of VCP ATPase activity, we show that VCP stabilizes Beclin-1 levels by promoting the deubiquitinase activity of ataxin-3 towards Beclin-1. VCP also regulates the assembly and activity of the Beclin-1-containing phosphatidylinositol-3-kinase (PI3K) complex I, thus regulating the production of PI(3)P, a key signaling lipid responsible for the recruitment of downstream autophagy factors. A decreased level of VCP, or inhibition of its ATPase activity, impairs starvation-induced production of PI(3)P and limits downstream recruitment of WIPI2, ATG16L and LC3, thereby decreasing autophagosome formation, illustrating an important role for VCP in early autophagy initiation.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All relevant data are included in the paper or its Supplementary Information. Source data are provided with this paper. Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding author.
References
Rubinsztein, D. C., Codogno, P. & Levine, B. Autophagy modulation as a potential therapeutic target for diverse diseases. Nat. Rev. Drug Discov. 11, 709–730 (2012).
Bento, C. F. et al. Mammalian autophagy: How does it work? Annu. Rev. Biochem. 85, 685–713 (2016).
Obara, K., Noda, T., Niimi, K. & Ohsumi, Y. Transport of phosphatidylinositol 3-phosphate into the vacuole via autophagic membranes in Saccharomyces cerevisiae. Genes Cells 13, 537–547 (2008).
Puri, C., Vicinanza, M. & Rubinsztein, D. C. Phagophores evolve from recycling endosomes. Autophagy 14, 1475–1477 (2018).
Hill, S. M., Wrobel, L. & Rubinsztein, D. C. Post-translational modifications of Beclin 1 provide multiple strategies for autophagy regulation. Cell Death Differ. https://doi.org/10.1038/s41418-018-0254-9 (2018).
Mizushima, N., Sugita, H., Yoshimori, T. & Ohsumi, Y. A new protein conjugation system in human. The counterpart of the yeast Apg12p conjugation system essential for autophagy. J. Biol. Chem. 273, 33889–33892 (1998).
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
van den Boom, J. & Meyer, H. VCP/p97-mediated unfolding as a principle in protein homeostasis and signaling. Mol. Cell 69, 182–194 (2018).
Tresse, E. et al. VCP/p97 is essential for maturation of ubiquitin-containing autophagosomes and this function is impaired by mutations that cause IBMPFD. Autophagy 6, 217–227 (2010).
Ju, J. S., Miller, S. E., Hanson, P. I. & Weihl, C. C. Impaired protein aggregate handling and clearance underlie the pathogenesis of p97/VCP-associated disease. J. Biol. Chem. 283, 30289–30299 (2008).
Ju, J. S. et al. Valosin-containing protein (VCP) is required for autophagy and is disrupted in VCP disease. J. Cell Biol. 187, 875–888 (2009).
Papadopoulos, C. et al. VCP/p97 cooperates with YOD1, UBXD1 and PLAA to drive clearance of ruptured lysosomes by autophagy. EMBO J. 36, 135–150 (2017).
Ramanathan, H. N. & Ye, Y. The p97 ATPase associates with EEA1 to regulate the size of early endosomes. Cell Res. 22, 346–359 (2012).
Ritz, D. et al. Endolysosomal sorting of ubiquitylated caveolin-1 is regulated by VCP and UBXD1 and impaired by VCP disease mutations. Nat. Cell Biol. 13, 1116–1123 (2011).
Chou, T. F. et al. Reversible inhibitor of p97, DBeQ, impairs both ubiquitin-dependent and autophagic protein clearance pathways. Proc. Natl Acad. Sci. USA 108, 4834–4839 (2011).
Birgisdottir, A. B. & Johansen, T. Autophagy and endocytosis—interconnections and interdependencies. J. Cell Sci. (2020); https://doi.org/10.1242/jcs.228114
Pleasure, I. T., Black, M. M. & Keen, J. H. Valosin-containing protein, VCP, is an ubiquitous clathrin-binding protein. Nature 365, 459–462 (1993).
Ashkenazi, A. et al. Polyglutamine tracts regulate beclin 1-dependent autophagy. Nature 545, 108–111 (2017).
Hirabayashi, M. et al. VCP/p97 in abnormal protein aggregates, cytoplasmic vacuoles, and cell death, phenotypes relevant to neurodegeneration. Cell Death Differ. 8, 977–984 (2001).
Doss-Pepe, E. W., Stenroos, E. S., Johnson, W. G. & Madura, K. Ataxin-3 interactions with rad23 and valosin-containing protein and its associations with ubiquitin chains and the proteasome are consistent with a role in ubiquitin-mediated proteolysis. Mol. Cell. Biol. 23, 6469–6483 (2003).
Wang, Q., Li, L. & Ye, Y. Regulation of retrotranslocation by p97-associated deubiquitinating enzyme ataxin-3. J. Cell Biol. 174, 963–971 (2006).
Laco, M. N., Cortes, L., Travis, S. M., Paulson, H. L. & Rego, A. C. Valosin-containing protein (VCP/p97) is an activator of wild-type ataxin-3. PLoS ONE 7, e43563 (2012).
Sun, Q. et al. Identification of Barkor as a mammalian autophagy-specific factor for Beclin 1 and class III phosphatidylinositol 3-kinase. Proc. Natl Acad. Sci. USA 105, 19211–19216 (2008).
Song, C., Wang, Q. & Li, C. C. Characterization of the aggregation-prevention activity of p97/valosin-containing protein. Biochemistry 46, 14889–14898 (2007).
Magnaghi, P. et al. Covalent and allosteric inhibitors of the ATPase VCP/p97 induce cancer cell death. Nat. Chem. Biol. 9, 548–556 (2013).
Zhou, H. J. et al. Discovery of a first-in-class, potent, selective and orally bioavailable inhibitor of the p97 AAA ATPase (CB-5083). J. Med. Chem. 58, 9480–9497 (2015).
Ravikumar, B., Imarisio, S., Sarkar, S., O’Kane, C. J. & Rubinsztein, D. C. Rab5 modulates aggregation and toxicity of mutant huntingtin through macroautophagy in cell and fly models of Huntington disease. J. Cell Sci. 121, 1649–1660 (2008).
Fu, Q. et al. Valosin-containing protein (VCP) promotes the growth, invasion and metastasis of colorectal cancer through activation of STAT3 signaling. Mol. Cell. Biochem. 418, 189–198 (2016).
Wojcik, C., Yano, M. & DeMartino, G. N. RNA interference of valosin-containing protein (VCP/p97) reveals multiple cellular roles linked to ubiquitin/proteasome-dependent proteolysis. J. Cell Sci. 117, 281–292 (2004).
Dooley, H. C. et al. WIPI2 links LC3 conjugation with PI3P, autophagosome formation and pathogen clearance by recruiting Atg12-5-16L1. Mol. Cell 55, 238–252 (2014).
Herzog, L. K. et al. The Machado–Joseph disease deubiquitylase ataxin-3 interacts with LC3C/GABARAP and promotes autophagy. Aging Cell 19, e13051 (2020).
Boeddrich, A. et al. An arginine/lysine-rich motif is crucial for VCP/p97-mediated modulation of ataxin-3 fibrillogenesis. EMBO J. 25, 1547–1558 (2006).
Funderburk, S. F., Wang, Q. J. & Yue, Z. The Beclin 1-VPS34 complex—at the crossroads of autophagy and beyond. Trends Cell Biol. 20, 355–362 (2010).
Itakura, E., Kishi, C., Inoue, K. & Mizushima, N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol. Biol. Cell 19, 5360–5372 (2008).
Liang, C. et al. Beclin1-binding UVRAG targets the class C VPS complex to coordinate autophagosome maturation and endocytic trafficking. Nat. Cell Biol. 10, 776–787 (2008).
Matsunaga, K. et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat. Cell Biol. 11, 385–396 (2009).
Zhong, Y. et al. Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat. Cell Biol. 11, 468–476 (2009).
Itakura, E. & Mizushima, N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy 6, 764–776 (2010).
Bruderer, R. M., Brasseur, C. & Meyer, H. H. The AAA ATPase p97/VCP interacts with its alternative co-factors, Ufd1-Npl4 and p47, through a common bipartite binding mechanism. J. Biol. Chem. 279, 49609–49616 (2004).
Chou, T. F. et al. Specific inhibition of p97/VCP ATPase and kinetic analysis demonstrate interaction between D1 and D2 ATPase domains. J. Mol. Biol. 426, 2886–2899 (2014).
Tang, W. K., Odzorig, T., Jin, W. & Xia, D. Structural basis of p97 inhibition by the site-selective anticancer compound CB-5083. Mol. Pharmacol. 95, 286–293 (2019).
Shiozawa, K. et al. The common phospholipid-binding activity of the N-terminal domains of PEX1 and VCP/p97. FEBS J. 273, 4959–4971 (2006).
Pecheur, E. I. et al. Phospholipid species act as modulators in p97/p47-mediated fusion of Golgi membranes. Biochemistry 41, 9813–9823 (2002).
Nascimbeni, A. C., Codogno, P. & Morel, E. Phosphatidylinositol-3-phosphate in the regulation of autophagy membrane dynamics. FEBS J. 284, 1267–1278 (2017).
Chan, N. et al. Valosin-containing protein mutation and Parkinson’s disease. Parkinsonism Relat. Disord. 18, 107–109 (2012).
Johnson, J. O. et al. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68, 857–864 (2010).
Darwich, N. F. et al. Autosomal dominant VCP hypomorph mutation impairs disaggregation of PHF-tau. Science 370, eaay8826 (2020); https://doi.org/10.1126/science.aay8826
Ghosh, D. K., Roy, A. & Ranjan, A. The ATPase VCP/p97 functions as a disaggregase against toxic huntingtin-exon1 aggregates. FEBS Lett. 592, 2680–2692 (2018).
Wang, B. et al. ULK1 and ULK2 regulate stress granule disassembly through phosphorylation and activation of VCP/p97. Mol. Cell 74, 742–757 (2019).
Arhzaouy, K. et al. VCP maintains lysosomal homeostasis and TFEB activity in differentiated skeletal muscle. Autophagy 15, 1082–1099 (2019).
Chan, N. C. et al. Degradation of the deubiquitinating enzyme USP33 is mediated by p97 and the ubiquitin ligase HERC2. J. Biol. Chem. 289, 19789–19798 (2014).
Steffen, J. et al. Rapid degradation of mutant SLC25A46 by the ubiquitin-proteasome system results in MFN1/2-mediated hyperfusion of mitochondria. Mol. Biol. Cell 28, 600–612 (2017).
McLelland, G. L. et al. Mfn2 ubiquitination by PINK1/parkin gates the p97-dependent release of ER from mitochondria to drive mitophagy. eLife https://doi.org/10.7554/eLife.32866 (2018).
Singh, A. N. et al. The p97–Ataxin 3 complex regulates homeostasis of the DNA damage response E3 ubiquitin ligase RNF8. EMBO J. 38, e102361 (2019).
Ganji, R., Mukkavalli, S., Somanji, F. & Raman, M. The VCP–UBXN1 complex mediates triage of ubiquitylated cytosolic proteins bound to the BAG6 complex. Mol. Cell. Biol. https://doi.org/10.1128/MCB.00154-18 (2018).
Nguyen, T. V. et al. p97/VCP promotes degradation of CRBN substrate glutamine synthetase and neosubstrates. Proc. Natl Acad. Sci. USA 114, 3565–3571 (2017).
Hammond, G. R., Schiavo, G. & Irvine, R. F. Immunocytochemical techniques reveal multiple, distinct cellular pools of PtdIns4P and PtdIns(4,5)P2. Biochem. J. 422, 23–35 (2009).
Acknowledgements
We thank Z. Yue for sharing the pFLAG-ATG14L construct and R. Schröder and C. Clemen for sharing pGEX-VCP-GST. We thank P. Widlund and S. Qamar for advice and technical support for the protein purification and in vitro methods. We thank F. Siddiqi and C. Karabiyik for assistance with brain dissection and primary cultures, G. Runwal for cloning the UFD1LΔVCP construct and R. Antrobus at CIMR proteomics facility for mass spectrometry analysis. We are grateful for funding from the UK Dementia Research Institute (funded by the MRC, Alzheimer’s Research UK and the Alzheimer’s Society) (UKDRI-2002 to D.C.R.), The Tau Consortium, Alzheimer’s Research UK, an anonymous donation to the Cambridge Centre for Parkinson-Plus, AstraZeneca, the Swedish Natural Research Council (VR) (reference 2016–06605; to S.M.H.) and from the European Molecular Biology Organisation (EMBO long-term fellowships, ALTF 1024-2016 and ALTF 135-2016, to S.M.H. and L.W., respectively).
Author information
Authors and Affiliations
Contributions
D.C.R. conceptualized and supervised the project. Experiments were designed, performed and analyzed by S.M.H., L.W., A.A. and M.F.-E., and A.A. performed the initial experiments on the VCP and Beclin-1 interaction, and analyzed the effect of VCP knockdown and inhibition on LC3 levels. L.W. and M.F.-E. performed immunofluorescence analysis of early autophagy factors WIPI2, ATG16L and LC3. S.M.H. performed all the in cell and in vitro binding studies, PI(3)P detection, in vitro deubiquitination, analysis of protein levels and summarized and compiled all data for the manuscript. R.W.B. and K.T. contributed to the design and interpretation of experiments. S.M.H. and D.C.R. wrote the manuscript with input from L.W. and all the other authors.
Corresponding author
Ethics declarations
Competing interests
K.T. and R.W.B. were employees of AstraZeneca when the experiments were performed and are shareholders of AstraZeneca. R.W.B. is currently employed by Cerevance Ltd. D.C.R. is a consultant for Aladdin Healthcare Technologies Ltd and Nido Biosciences. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 VCP interacts with Beclin-1 in vivo and in vitro.
a, Schematic overview of Beclin-1 protein domains; with the N-terminal domain in dark gray, coiled-coiled domain (CCD) in blue, and the beta-alpha repeated, autophagy-specific (BARA) domain in light grey. Depicted below is the different truncation mutants used for binding studies (all with N-terminal FLAG tag). b, In vitro binding of FLAG-tagged Beclin-1 truncation mutants to VCP-GST crosslinked to beads. Empty Glutathione beads and mutated VCP interactor UFD1L (Δaa215–241; UFD1LΔVCP), were used as a negative control in binding experiments. Ratio of bound protein bound to VCP-GST was quantified by detection of FLAG signal in bound fraction divided by input signal and normalized to the full-length control; n=5, UFD1LΔVCP p= 0.0031. c, Representative gel of in vitro binding experiment quantified in (b). d, DBeQ impairs autophagic flux and autophagosome formation during starvation (HBSS, 4 h) in primary neurons, as demonstrated by LC3-II levels in conditions without and with 1 h pre-treatment with Bafilomycin A1 (BafA; 400 nM); n=3, -BafA p = 0.00105, +BafA p = 0.00005. b, d, Bar graphs data presented as normalized mean ± SD, **p < 0.005, ***p < 0.0005, unpaired two-tailed Students t-test. See also main Fig. 1.
Extended Data Fig. 2 VCP regulates early autophagy initiation.
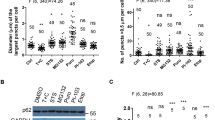
a, LC3 puncta formation upon treatment with VCP inhibitors. HeLa cells were pre-treated with BafA (2 h), before transition to HBSS (+ BafA), with addition of DMSO, 10 μM NMS873 or 5 μM CB-5083 for 3 h. Number of LC3 puncta per cell normalized to DMEM control; n=3, number of cells counted: 50-60 per condition, one-way ANOVA with Dunnett’s correction for multiple comparisons, DMEM vs HBSS+DMSO p = 0.049, HBSS+DMSO vs HBSS+NMS873 p = 0.009, HBSS+DMSO vs HBSS+CB-5083 p = 0.012. b, Size of LC3 puncta during starvation and VCP inhibition; n=3. c, Representative images of LC3 puncta formation, as quantified in (a) and (b). d, Representative images for PI(3)P puncta formation upon starvation (HBSS 1 h) in HeLa cells, with and without treatment with the VPS34 inhibitor Wm (1μM) as quantified in main Fig. 1f. e, Representative images for PI(3)P puncta formation upon starvation (HBSS 1 h) in HeLa cells, with and without treatment with 10 μM NMS873 or 5 μM CB-5083 compared to DMSO in basal media (DMEM) as quantified in (f, g). f, Quantification of number of PI(3)P puncta per cell during starvation and VCP inhibition normalized to DMEM control; n=4, number of cells counted: 40-50 per condition, one-way ANOVA with Dunnett’s correction for multiple comparisons: DMEM vs HBSS p= 0.003, HBSS vs HBSS+NMS873 p < 0,001, HBSS vs HBSS+2 µM CB-5083 p= 0.016, HBSS vs HBSS+5 µM CB-5083 p < 0,001, HBSS vs HBSS+IN1 p= 0.002. g, Quantification of size of PI(3)P puncta per cell from experiment in (f); one-way ANOVA with Dunnett’s correction for multiple comparisons: HBSS vs HBSS+NMS873 p= 0.033, HBSS vs HBSS+2 µM CB-5083 p= 0.049, HBSS vs HBSS+5 µM CB-5083 p=0.049, HBSS vs HBSS+IN1 p= 0.026. h, ATG5-ATG12 conjugation in HeLa cells treated with siRNA to knockdown expression of VCP; n=3, unpaired two-tailed Student t-test, VCP siRNA 11 p= 0.0009, VCP siRNA 12 P= 0.0007. a, b, f-h, Data in bar graphs presented as normalized mean ± SD. *p<0.05, **p<0.005, ***p < 0.0005. Scale bar = 10 μm. See also main Fig. 1.
Extended Data Fig. 3 Knockdown of VCP affects autophagy initiation.
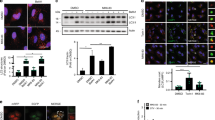
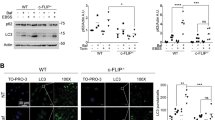
a, Knockdown of VCP impairs PI(3)P production upon starvation. HeLa cells treated with control siRNA (ctrl) or siRNA targeting VCP were starved (HBSS 1 h) 48 h post transfection and stained for nuclei (DAPI, blue), VCP levels (green) and PI(3)P production (red). b, Quantified data for experiment in (a), shown as normalized mean ± SD, n= 3, number of cells counted: 40-50 per condition, unpaired two-tailed Students t-test, DMEM vs HBSS p= 0.03855, HBSS vs kd VCP p= 0.04527. c, LC3-II levels upon VCP knockdown in the absence and presence of BafA (400 nM). Efficiency of VCP kd tested in separate blot, before cells were divided into -BafA and +BafA conditions. d, Quantified data for LC3-II levels for experiment in (c), shown as normalized mean ± SD, n= 4, -BafA VCP siRNA 11 p= 0.00058, -BafA VCP siRNA 12 p= 0.00046. *p<0.05, **p < 0.005, ***p < 0.0005. Scale bar = 10 uM. See also main Fig. 1.
Extended Data Fig. 4 VCP regulates the recruitment of early autophagy markers during starvation and mTOR inhibition.
a, Immunofluorescent analysis of WIPI2 puncta formation upon starvation with VCP inhibition. HeLa cells in basal media (DMEM) compared to cells subjected to starvation (HBSS 2 h) in combination with DMSO, 10 μM NMS873 or 5 μM CB-5083. Median area of WIPI2 signal per cell normalized to control; n=5, number of cells counted: 30-40 per condition, DMEM+DMSO vs HBSS+DMSO p=0.0076, HBSS+DMSO vs HBSS+NMS873 p=0.0223, HBSS+DMSO vs HBSS+CB-5083 p=0.0253. b, Immunofluorescent analysis of ATG16 puncta formation upon starvation during VCP inhibition. HBSS 2 h, in combination with DMSO, 10 μM NMS873 or 5 μM CB-5083. Median area of ATG16 per cell normalized to control; n=4, number of cells counted: 30-40 per condition, DMEM+DMSO vs HBSS+DMSO p=0.0036, HBSS+DMSO vs HBSS+NMS873 p= 0.0033, HBSS+DMSO vs HBSS+CB-5083 p=0.0006. c, Immunofluorescent analysis of LC3 puncta induction upon mTOR inhibition by Torin 1. HeLa cells pre-treated with BafA (400 nM) together with DMSO, 10 μM NMS873 or 5 μM CB-5083 for 1 h, were subjected to Torin 1 (1 μM) treatment for 4 h. Median area of LC3 per cell normalized to control; n=3, number of cells counted: 30-40 per condition, DMSO vs Torin 1 p= 0.0021, Torin 1 vs Torin 1+NMS873 p= 0.0035, Torin 1 vs Torin 1+CB-5083 p= 0.0034. (a-c) Bar graphs data presented as normalized mean ± SD, *p<0.05, **p < 0.005, unpaired two-tailed students t-test. d, Representative images of WIPI2 puncta formation upon starvation, as quantified in (a). e, Representative images of ATG16 puncta formation, as quantified in (b). f, LC3 puncta formation upon Torin 1 treatment, as quantified in (c). Scale bar = 10 μm. See also main Fig. 2.
Extended Data Fig. 5 VCP regulates levels of Beclin-1 via Ataxin-3.
a, Beclin-1 levels in HeLa cells after knockdown of VCP compared to control cells transfected with non-targeting siRNA; n=3, VCP siRNA 11 p= 0.00158, VCP siRNA 12 p<0.0001. b, Beclin-1 levels in HeLa cells upon treatment with DMSO, 10 µM NMS873 or 5 µM CB-5083 for 5 h; n=7, NMS873 p= 0.00013, CB-5083 p= 0.00319. (c) Ataxin-3 levels in HeLa cells upon treatment with DMSO, 10 µM NMS873 or 5 µM CB-5083 for 5 h; n=3, NMS873 p= 0.0002, CB-5083 p= 0.0055. a-c, Bar graphs represent Beclin-1 or Ataxin-3 to loading control (Actin or GAPDH) ratios displayed as mean ± SD normalized to control, *p<0.05, **p < 0.005, ***p < 0.0005, unpaired two-tailed students t-test. d, FLAG-tagged Ataxin-3 or mutated version Ataxin-3 ΔVCP were expressed together with VCP-HA, followed by FLAG immunoprecipitation. e, Immunoprecipitation of FLAG proteins from cells expressing empty FLAG, FLAG-tagged Ataxin-3 or Ataxin-3 ΔVCP. See also main Fig. 3.
Extended Data Fig. 6 VCP regulates Ataxin-3 deubiquitinase activity towards Beclin-1.
a, b, Representative experiment from main Fig. 3f: LC3 puncta per cell in HeLa cells depleted of Ataxin-3 and reconstituted with wild-type Ataxin-3 or Ataxin-3ΔVCP, n=60-80 cells analyzed per condition. Scale bar = 10 μm. c, Representative gel image of in vitro deubiquitination assay, quantified in main Fig. 3g. See also main Fig. 3.
Extended Data Fig. 7 VCP interacts with PI3K complexes in vitro.
a, Immunoprecipitation of Ataxin-3-FLAG in control and VCP knockdown cells, overexpressing FLAG-Ataxin-3 and ATG14L. IP ratios are normalized to input to account for differences in input protein levels. b, Protein-stained gel with individually purified FLAG-tagged PI3K components. c, Protein-stained gel of VCP-GST purified from E. coli and crosslinked to glutathione agarose beads. d, In vitro immunoprecipitation with VCP-GST beads and individually purified PI3K components: VPS34, ATG14L, Beclin-1 and UFD1LΔVCP (Δaa215-241) as a negative control. Experiment was repeated 6 times.
Extended Data Fig. 8 VCP interacts with PI3K complex I.
a, FLAG-tagged PI3K complexes purified from HEK293 cells. Protein-stained gel (left) and FLAG-probed western blot (right). b, Quantification of immunoprecipitation of endogenous ATG14L in control cells and in VCP knockdown cells. IP ratios have been normalized to input ratios. Representative gel in (c); n=3, unpaired two-tailed student t-test, bar graph data presented as normalized mean ± SD, kd VCP Beclin-1 p=0.0127, kd VCP VPS34 p=0.0041.d, VCP levels in input samples from endogenous immunoprecipitation of ATG14L with short and long exposure (SE and LE). VCP upper band denoted by red arrow. e, Short and long exposures of VCP levels from VCP kd experiments. Quantifications shown below blots, normalized to levels in control sample. f, Purification of FLAG-VPS15 and FLAG-VPS34 from 293Expi cells. Purified proteins are visualized with protein stain (instant blue; left lane) and by western blot analysis using FLAG antibody (middle lane). The same membrane was cut and probed with individual antibodies for VPS15, VPS34, ATG14L and Beclin-1 (right lane). See also main Figs. 4 and 5.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 8
Unprocessed western blots and/or gels.
Rights and permissions
About this article
Cite this article
Hill, S.M., Wrobel, L., Ashkenazi, A. et al. VCP/p97 regulates Beclin-1-dependent autophagy initiation. Nat Chem Biol 17, 448–455 (2021). https://doi.org/10.1038/s41589-020-00726-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00726-x
This article is cited by
-
Crosstalk of cell death pathways unveils an autophagy-related gene AOC3 as a critical prognostic marker in colorectal cancer
Communications Biology (2024)
-
Seasonal modulation of the testis transcriptome reveals insights into hibernation and reproductive adaptation in Onychostoma macrolepis
Fish Physiology and Biochemistry (2024)
-
PRKAA2, MTOR, and TFEB in the regulation of lysosomal damage response and autophagy
Journal of Molecular Medicine (2024)
-
Cardioprotective effect of crude polysaccharide fermented by Trametes Sanguinea Lyoyd on doxorubicin-induced myocardial injury mice
BMC Pharmacology and Toxicology (2023)
-
Valosin containing protein (VCP): initiator, modifier, and potential drug target for neurodegenerative diseases
Molecular Neurodegeneration (2023)