Abstract
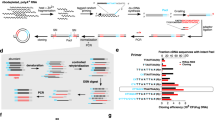
Secreted polypeptides are a fundamental axis of intercellular and endocrine communication. However, a global understanding of the composition and dynamics of cellular secretomes in intact mammalian organisms has been lacking. Here, we introduce a proximity biotinylation strategy that enables labeling, detection and enrichment of secreted polypeptides in a cell type-selective manner in mice. We generate a proteomic atlas of hepatocyte, myocyte, pericyte and myeloid cell secretomes by direct purification of biotinylated secreted proteins from blood plasma. Our secretome dataset validates known cell type–protein pairs, reveals secreted polypeptides that distinguish between cell types and identifies new cellular sources for classical plasma proteins. Lastly, we uncover a dynamic and previously undescribed nutrient-dependent reprogramming of the hepatocyte secretome characterized by the increased unconventional secretion of the cytosolic enzyme betaine–homocysteine S-methyltransferase (BHMT). This secretome profiling strategy enables dynamic and cell type-specific dissection of the plasma proteome and the secreted polypeptides that mediate intercellular signaling.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that data supporting the findings of this study are available within the paper and its Supplementary Information files. Figs. 3, 5 and 6 have associated raw data provided in Supplementary Tables 1–4. All proteomic data generated here are publicly available on ProteomeXchange under project accession no. PXD021602. Source data are provided with this paper.
References
Rorsman, P. & Braun, M. Regulation of insulin secretion in human pancreatic islets. Annu. Rev. Physiol. 75, 155–179 (2013).
Gaceb, A., Barbariga, M., Özen, I. & Paul, G. The pericyte secretome: potential impact on regeneration. Biochimie 155, 16–25 (2018).
Lim, J. M., Wollaston-Hayden, E. E., Teo, C. F., Hausman, D. & Wells, L. Quantitative secretome and glycome of primary human adipocytes during insulin resistance. Clin. Proteomics 11, 20 (2014).
Rabouille, C. Pathways of unconventional protein secretion. Trends Cell Biol. 27, 230–240 (2017).
Eichelbaum, K., Winter, M., Diaz, M. B., Herzig, S. & Krijgsveld, J. Selective enrichment of newly synthesized proteins for quantitative secretome analysis. Nat. Biotechnol. 30, 984–990 (2012).
Yang, A. C. et al. Multiple click-selective tRNA synthetases expand mammalian cell-specific proteomics. J. Am. Chem. Soc. 140, 7046–7051 (2018).
Shin, J. et al. Comparative analysis of differentially secreted proteins in serum-free and serum-containing media by using BONCAT and pulsed SILAC. Sci. Rep. 9, 3096 (2019).
Eichelbaum, K. & Krijgsveld, J. Combining pulsed SILAC labeling and click-chemistry for quantitative secretome analysis. Methods Mol. Biol. 1174, 101–114 (2014).
Witzke, K. E. et al. Quantitative secretome analysis of activated Jurkat cells using click chemistry-based enrichment of secreted glycoproteins. J. Proteome Res. 16, 137–146 (2017).
Kim, D. I. et al. An improved smaller biotin ligase for BioID proximity labeling. Mol. Biol. Cell 27, 1188–1196 (2016).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
May, D. G., Scott, K. L., Campos, A. R. & Roux, K. J. Comparative application of BioID and TurboID for protein-proximity biotinylation. Cells 9, 1070 (2020).
Octeau, J. C. et al. An optical neuron–astrocyte proximity assay at synaptic distance scales. Neuron 98, 49–66 (2018).
Long, J. Z. et al. The secreted enzyme PM20D1 regulates lipidated amino acid uncouplers of mitochondria. Cell 166, 424–435 (2016).
Long, J. Z. et al. Ablation of PM20D1 reveals N-acyl amino acid control of metabolism and nociception. Proc. Natl Acad. Sci. USA 115, E6937–E6945 (2018).
Kim, J. T. et al. Cooperative enzymatic control of N-acyl amino acids by PM20D1 and FAAH. eLife 9, e55211 (2020).
Jackson, A. et al. Heat shock induces the release of fibroblast growth factor 1 from NIH 3T3 cells. Proc. Natl Acad. Sci. USA 89, 10691–10695 (1992).
Yan, Z., Yan, H. & Ou, H. Human thyroxine binding globulin (TBG) promoter directs efficient and sustaining transgene expression in liver-specific pattern. Gene 506, 289–294 (2012).
Uezu, A. et al. Identification of an elaborate complex mediating postsynaptic inhibition. Science 353, 1123–1129 (2016).
Pályi-Krekk, Z. et al. EGFR and ErbB2 are functionally coupled to CD44 and regulate shedding, internalization and motogenic effect of CD44. Cancer Lett. 263, 231–242 (2008).
Wu, C. et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 10, R130 (2009).
Samuel, V. T. Fructose induced lipogenesis: from sugar to fat to insulin resistance. Trends Endocrinol. Metab. 22, 60–65 (2011).
Softic, S. et al. Divergent effects of glucose and fructose on hepatic lipogenesis and insulin signaling. J. Clin. Invest. 127, 4059–4074 (2017).
Softic, S. et al. Dietary sugars alter hepatic fatty acid oxidation via transcriptional and post-translational modifications of mitochondrial proteins. Cell Metab. 30, 735–753 (2019).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Teng, Y. W., Mehedint, M. G., Garrow, T. A. & Zeisel, S. H. Deletion of betaine–homocysteine S-methyltransferase in mice perturbs choline and 1-carbon metabolism, resulting in fatty liver and hepatocellular carcinomas. J. Biol. Chem. 286, 36258–36267 (2011).
Qin, S. et al. Identification of organ-enriched protein biomarkers of acute liver injury by targeted quantitative proteomics of blood in acetaminophen- and carbon-tetrachloride-treated mouse models and acetaminophen overdose patients. J. Proteome Res. 7, 3724–3740 (2016).
Wang, B. et al. Construction and analysis of compact muscle-specific promoters for AAV vectors. Gene Ther. 15, 1489–1499 (2008).
Schnütgen, F. et al. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nat. Biotechnol. 21, 562–565 (2003).
Canli, Ö. et al. Myeloid cell-derived reactive oxygen species induce epithelial mutagenesis. Cancer Cell 32, 869–883 (2017).
Cuervo, H. et al. PDGFRβ-P2A-CreERT2 mice: a genetic tool to target pericytes in angiogenesis. Angiogenesis 20, 655–662 (2017).
Postic, C. et al. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic β cell-specific gene knock-outs using Cre recombinase. J. Biol. Chem. 274, 305–315 (1999).
Zimmers, T. A. et al. Induction of cachexia in mice by systemically administered myostatin. Science 296, 1486–1488 (2002).
McPherron, A. C., Lawler, A. M. & Lee, S. J. Regulation of skeletal muscle mass in mice by a new TGF-β superfamily member. Nature 387, 83–90 (1997).
Hu, E., Liang, P. & Spiegelman, B. M. AdipoQ is a novel adipose-specific gene dysregulated in obesity. J Biol. Chem. 271, 10697–10703 (1996).
Scherer, P. E., Williams, S., Fogliano, M., Baldini, G. & Lodish, H. F. A novel serum protein similar to C1q, produced exclusively in adipocytes. J. Biol. Chem. 270, 26746–26749 (1995).
Delaigle, A. M., Senou, M., Guiot, Y., Many, M. C. & Brichard, S. M. Induction of adiponectin in skeletal muscle of type 2 diabetic mice: in vivo and in vitro studies. Diabetologia 49, 1311–1323 (2006).
Piñeiro, R. et al. Adiponectin is synthesized and secreted by human and murine cardiomyocytes. FEBS Lett. 579, 5163–5169 (2005).
Desiere, F. et al. The PeptideAtlas project. Nucleic Acids Res. 34, D655–D658 (2006).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Mouchiroud, M. et al. The hepatokine Tsukushi is released in response to NAFLD and impacts cholesterol homeostasis. JCI Insight 4, e129492 (2019).
Barb, D., Bril, F., Kalavalapalli, S. & Cusi, K. Plasma fibroblast growth factor 21 is associated with severity of nonalcoholic steatohepatitis in patients with obesity and type 2 diabetes. J. Clin. Endocrinol. Metab. 104, 3327–3336 (2019).
Xiong, X. et al. Mapping the molecular signatures of diet-induced NASH and its regulation by the hepatokine Tsukushi. Mol. Metab. 20, 128–137 (2019).
Krahmer, N. et al. Organellar proteomics and phospho-proteomics reveal subcellular reorganization in diet-induced hepatic steatosis. Dev. Cell 47, 205–221 (2018).
Hailemariam, M. et al. S-Trap, an ultrafast sample-preparation approach for shotgun proteomics. J. Proteome Res. 17, 2917–2924 (2018).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Michaud, S. A. et al. Molecular phenotyping of laboratory mouse strains using 500 multiple reaction monitoring mass spectrometry plasma assays. Commun. Biol. 1, 78 (2018).
Acknowledgements
We thank members of the Long, Bertozzi, Svensson and Abu-Remaileh labs for helpful discussions. We gratefully acknowledge the staff at the Penn Vector Core for the production of AAVs. This work was supported by the US National Institutes of Health (DK105203 and DK124265 to J.Z.L. and K00CA21245403 to N.M.R.) and the Stanford Diabetes Research Center (P30DK116074).
Author information
Authors and Affiliations
Contributions
W.W. generated all constructs used for this study and performed all mouse experiments. N.M.R. performed all proteomic analyses. A.C.Y. and S.M.T. performed histological and immunofluorescence analyses of livers. J.T.K., V.L.L. and M.G.-C. assisted in tissue collection and processing for mouse experiments. C.R.B. and J.Z.L. supervised the work. W.W., N.M.R. and J.Z.L. conceived experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13
Supplementary Table 1
LC–MS/MS proteomic analysis of streptavidin-purified plasma proteins from mice transduced with AAV-Tbg-Mem, AAV-Tbg-Cyto or AAV-Tbg-ER viruses or control mice. Statistical comparisons were made using ANOVA and Student’s two-way t-test.
Supplementary Table 2
LC–MS/MS proteomic analysis of streptavidin-purified liver proteins from mice transduced with AAV-Tbg-Mem, AAV-Tbg-Cyto or AAV-Tbg-ER viruses.
Supplementary Table 3
LC–MS/MS proteomic analysis of streptavidin-purified plasma from control mice (chow) without viral transduction or mice transduced with AAV-Tbg-Cyto after 2 weeks of chow or HFHS diet. Statistical comparisons were made using ANOVA and Student’s two-way t-test.
Supplementary Table 4
LC–MS/MS proteomic analysis of streptavidin-purified plasma from mice. The following cell types correspond to the following mouse genotypes and viruses: hepatocytes (albumin-Cre, AAV-FLEx), myocytes (WT mice, AAV-tMCK), pericytes (Pdgfrb-creERT2, AAV-FLEx), myeloid cells (LysM-creERT2, AAV-FLEx). Statistical comparisons were made using ANOVA.
Source data
Source Data Fig. 1
Uncropped gel images for Fig. 1.
Source Data Fig. 2
Uncropped gel images for Fig. 2.
Source Data Fig. 4
Uncropped gel images for Fig. 4.
Source Data Fig. 5
Uncropped gel images for Fig. 5.
Rights and permissions
About this article
Cite this article
Wei, W., Riley, N.M., Yang, A.C. et al. Cell type-selective secretome profiling in vivo. Nat Chem Biol 17, 326–334 (2021). https://doi.org/10.1038/s41589-020-00698-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00698-y
This article is cited by
-
Uncovering the mechanisms underlying pear leaf apoplast protein-mediated resistance against Colletotrichum fructicola through transcriptome and proteome profiling
Phytopathology Research (2024)
-
Cell-selective proteomics segregates pancreatic cancer subtypes by extracellular proteins in tumors and circulation
Nature Communications (2023)
-
Imaging and AI based chromatin biomarkers for diagnosis and therapy evaluation from liquid biopsies
npj Precision Oncology (2023)
-
Genetically encoded photocatalytic protein labeling enables spatially-resolved profiling of intracellular proteome
Nature Communications (2023)
-
EGFR signaling promotes nuclear translocation of plasma membrane protein TSPAN8 to enhance tumor progression via STAT3-mediated transcription
Cell Research (2022)