Abstract
The DNA guanine quadruplexes (G4) play important roles in multiple cellular processes, including DNA replication, transcription and maintenance of genome stability. Here, we showed that Yin and Yang 1 (YY1) can bind directly to G4 structures. ChIP–seq results revealed that YY1-binding sites overlap extensively with G4 structure loci in chromatin. We also observed that the dimerization of YY1 and its binding with G4 structures contribute to YY1-mediated long-range DNA looping. Displacement of YY1 from G4 structure sites disrupts substantially the YY1-mediated DNA looping. Moreover, treatment with G4-stabilizing ligands modulates the expression of not only those genes with G4 structures in their promoters, but also those associated with distal G4 structures that are brought to close proximity via YY1-mediated DNA looping. Together, we identified YY1 as a DNA G4-binding protein, and revealed that YY1-mediated long-range DNA looping requires its dimerization and occurs, in part, through its recognition of G4 structure.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The ChIP–seq, HiChIP–seq and RNA-seq data generated in this study have been deposited into the NCBI GEO database (for ChIP–seq and HiChIP–seq data: GEO accession number GSE128106; for RNA-seq: GEO accession number GSE142075). The ChIP–seq data for Ishikawa and SK-N-SH cells were obtained from NCBI GEO database with accession numbers of GSM1010753 and GSM1010897, respectively43. The two G4 ChIP–seq datasets were obtained from NCBI GEO database with accession numbers of GSE99205 and GSE107690 (refs. 3,7). The human hg19 reference genome was downloaded from https://hgdownload.soe.ucsc.edu/goldenPath/hg19/bigZips. Source data are provided with this paper.
Code availability
Custom codes used in this work are available from https://github.com/linliucr/UCR_code.
References
Bochman, M. L., Paeschke, K. & Zakian, V. A. DNA secondary structures: stability and function of G-quadruplex structures. Nat. Rev. Genet. 13, 770–780 (2012).
Hansel-Hertsch, R., Di Antonio, M. & Balasubramanian, S. DNA G-quadruplexes in the human genome: detection, functions and therapeutic potential. Nat. Rev. Mol. Cell Biol. 18, 279–284 (2017).
Mao, S. Q. et al. DNA G-quadruplex structures mold the DNA methylome. Nat. Struct. Mol. Biol. 25, 951–957 (2018).
Huppert, J. L. & Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 33, 2908–2916 (2005).
Biffi, G., Tannahill, D., McCafferty, J. & Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 5, 182–186 (2013).
Henderson, A. et al. Detection of G-quadruplex DNA in mammalian cells. Nucleic Acids Res. 42, 860–869 (2014).
Hansel-Hertsch, R., Spiegel, J., Marsico, G., Tannahill, D. & Balasubramanian, S. Genome-wide mapping of endogenous G-quadruplex DNA structures by chromatin immunoprecipitation and high-throughput sequencing. Nat. Protoc. 13, 551–564 (2018).
Soldatenkov, V. A., Vetcher, A. A., Duka, T. & Ladame, S. First evidence of a functional interaction between DNA quadruplexes and poly(ADP-ribose) polymerase-1. ACS Chem. Biol. 3, 214–219 (2008).
Gonzalez, V., Guo, K., Hurley, L. & Sun, D. Identification and characterization of nucleolin as a c-myc G-quadruplex-binding protein. J. Biol. Chem. 284, 23622–23635 (2009).
Paramasivam, M. et al. Protein hnRNP A1 and its derivative Up1 unfold quadruplex DNA in the human KRAS promoter: implications for transcription. Nucleic Acids Res. 37, 2841–2853 (2009).
Williams, P., Li, L., Dong, X. & Wang, Y. Identification of SLIRP as a G quadruplex-binding protein. J. Am. Chem. Soc. 139, 12426–12429 (2017).
Gordon, S., Akopyan, G., Garban, H. & Bonavida, B. Transcription factor YY1: structure, function, and therapeutic implications in cancer biology. Oncogene 25, 1125–1142 (2006).
Deng, Z., Cao, P., Wan, M. M. & Sui, G. Yin Yang 1: a multifaceted protein beyond a transcription factor. Transcription 1, 81–84 (2010).
Wu, S. et al. A YY1–INO80 complex regulates genomic stability through homologous recombination-based repair. Nat. Struct. Mol. Biol. 14, 1165–1172 (2007).
Weintraub, A. S. et al. YY1 is a structural regulator of enhancer-promoter loops. Cell 171, 1573–1588.e28 (2017).
Maru, Y., Afar, D. E., Witte, O. N. & Shibuya, M. The dimerization property of glutathione S-transferase partially reactivates Bcr-Abl lacking the oligomerization domain. J. Biol. Chem. 271, 15353–15357 (1996).
Han, F. X., Wheelhouse, R. T. & Hurley, L. H. Interactions of TMPyP4 and TMPyP2 with quadruplex DNA. Structural basis for the differential effects on telomerase inhibition. J. Am. Chem. Soc. 121, 3561–3570 (1999).
Rodriguez, R. et al. A novel small molecule that alters shelterin integrity and triggers a DNA-damage response at telomeres. J. Am. Chem. Soc. 130, 15758–15759 (2008).
Houbaviy, H. B., Usheva, A., Shenk, T. & Burley, S. K. Cocrystal structure of YY1 bound to the adeno-associated virus P5 initiator. Proc. Natl Acad. Sci. USA 93, 13577–13582 (1996).
Whitfield, T. W. et al. Functional analysis of transcription factor binding sites in human promoters. Genome Biol. 13, R50 (2012).
Chambers, V. S. et al. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat. Biotechnol. 33, 877–881 (2015).
Muller, S., Kumari, S., Rodriguez, R. & Balasubramanian, S. Small-molecule-mediated G-quadruplex isolation from human cells. Nat. Chem. 2, 1095–1098 (2010).
Siddiqui-Jain, A., Grand, C. L., Bearss, D. J. & Hurley, L. H. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc. Natl Acad. Sci. USA 99, 11593–11598 (2002).
Lee, G. R. Role of YY1 in long-range chromosomal interactions regulating Th2 cytokine expression. Transcription 5, e27976 (2014).
Liu, H. et al. Yin Yang 1 is a critical regulator of B-cell development. Genes Dev. 21, 1179–1189 (2007).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Mohaghegh, P., Karow, J. K., Brosh, R. M. Jr., Bohr, V. A. & Hickson, I. D. The Bloom’s and Werner’s syndrome proteins are DNA structure-specific helicases. Nucleic Acids Res. 29, 2843–2849 (2001).
Lopez-Perrote, A. et al. Structure of Yin Yang 1 oligomers that cooperate with RuvBL1-RuvBL2 ATPases. J. Biol. Chem. 289, 22614–22629 (2014).
Shi, Y., Seto, E., Chang, L. S. & Shenk, T. Transcriptional repression by YY1, a human GLI-Kruppel-related protein, and relief of repression by adenovirus E1A protein. Cell 67, 377–388 (1991).
Merkenschlager, M. & Nora, E. P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 17, 17–43 (2016).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Parelho, V. et al. Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 132, 422–433 (2008).
Khachigian, L. M. The Yin and Yang of YY1 in tumor growth and suppression. Int. J. Cancer 143, 460–465 (2018).
Shi, J., Hao, A., Zhang, Q. & Sui, G. The role of YY1 in oncogenesis and its potential as a drug target in cancer therapies. Curr. Cancer Drug Targets 15, 145–157 (2015).
Rhodes, D. & Lipps, H. J. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 43, 8627–8637 (2015).
Favicchio, R., Dragan, A. I., Kneale, G. G. & Read, C. M. Fluorescence spectroscopy and anisotropy in the analysis of DNA–protein interactions. Methods Mol. Biol. 543, 589–611 (2009).
Li, L., Miao, W., Huang, M., Williams, P. & Wang, Y. Integrated genomic and proteomic analyses reveal novel mechanisms of the methyltransferase SETD2 in renal cell carcinoma development. Mol. Cell. Proteom. 18, 437–447 (2019).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP–seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Matsuda, K. et al. ChIP–seq analysis of genomic binding regions of five major transcription factors highlights a central role for ZIC2 in the mouse epiblast stem cell gene regulatory network. Development 144, 1948–1958 (2017).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997v2 (2013).
Wolff, J. et al. Galaxy HiCExplorer: a web server for reproducible Hi-C data analysis, quality control and visualization. Nucleic Acids Res. 46, W11–W16 (2018).
Gertz, J. et al. Distinct properties of cell-type-specific and shared transcription factor binding sites. Mol. Cell 52, 25–36 (2013).
Acknowledgements
This work was supported by the National Institutes of Health (R35 ES031707 to Y.W. and R35 GM119721 to J.S.). M.H. was supported in part by an NRSA T32 Institutional Research Training Grant (ES018827).
Author information
Authors and Affiliations
Contributions
L.L. and Y.W. conceived the project. P.W., W.M. and M.H. performed the G-quadruplex pull-down experiments and analyzed the mass spectrometry data. L.L., M.Y.W., Z.G. and W.M. performed the plasmid construction and cell culture experiments. L.L. and Z.G. performed the in vitro binding assay. L.L. performed the ChIP–seq, HiChIP–seq, RNA-seq and relevant data analysis. W.R. and J.S. assisted with the protein expression and purification. L.L. and P.W. analyzed the data. L.L. and Y.W. wrote the manuscript, which was reviewed and commented by all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
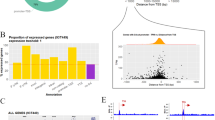
Extended Data Fig. 1 Proteome-wide identification of G quadruplex-binding proteins.
Volcano plots showing the quantification results of G quadruplex-binding proteins identified from SILAC-based interaction screening. The −Log10(P value) was plotted against the log2(ratio G4/M4). YY1 is labeled in red.
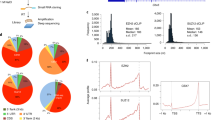
Extended Data Fig. 2 Circular dichroism (CD) spectra of the three G4 sequences in the absence or presence of tag-free YY1 protein.
a-c, CD spectra of cMYC G4 (a), YY1 (b) and cMYC G4-YY1 complex (c). d-f, Comparison of CD spectra for G4 probes in the presence or absence of YY1 protein. The CD spectra for the G4 probes in the presence of YY1 protein were obtained by subtracting the CD spectrum of the YY1 protein from the composite spectra of the protein-G4 DNA complexes.
Extended Data Fig. 3 Zinc finger domain of YY1 is essential, but not sufficient for the specific binding toward G4 structure.
a, A schematic diagram depicting the domain structure of YY1 protein. b-e, Fluorescence anisotropy for monitoring the bindings of mutant (YY1C360S) and truncated (YY11-382, YY1231-414, and YY1293-414) YY1 proteins with hTEL G4 and the corresponding M4 DNA probes. The data represent the mean ± S.E.M. of results obtained from 3 independent experiments.
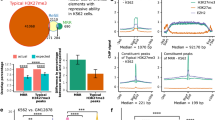
Extended Data Fig. 4 Statistical analysis of YY1 ChIP-Seq and BG4 ChIP-Seq data.
a, The ChIP-seq signal enrichment of YY1 ChIP-Seq and BG4 ChIP-Seq based on the overlapped peaks for the two datasets. The YY1 ChIP-Seq and BG4 ChIP-Seq average signal enrichments are plotted against the BG4 overlapped peaks. b, Analysis of peak width distribution for YY1 ChIP-seq and BG4 ChIP-seq data.
Extended Data Fig. 5 Unwinding of G4 with the overexpression of BLM helicase disrupts the YY1-mediated DNA looping.
a-b, Reads enrichment of two regions from BG4 ChIP-Seq and YY1 ChIP-Seq. The long-range DNA interactions monitored in HiChIP-PCR experiment are labeled with red arches, and the regions monitored in BG4 ChIP PCR experiments are indicated with blue triangles. c-d, BG4 ChIP and YY1 ChIP enrichments at the two sites were markedly diminished after ectopic overexpression of BLM helicase (BLM-O.E.). e, YY1-mediated DNA looping is disrupted by overexpression of BLM helicase. HiChIP-PCR quantification results of YY1-mediated DNA looping in HEK293T cells with and without the overexpression of BLM. Shown in (c-e) are mean ± S.E.M. of results obtained from 3 independent experiments. The p values were calculated by using two-tailed, unpaired Student’s t test: **, p < 0.01; ***, p < 0.001.
Extended Data Fig. 6 Analysis of YY1 dimerization and binding stoichiometry of the YY1-G4 DNA complex.
a-b, Gel filtration chromatography revealed the dimerization of YY1 and truncated YY1231-414, but not YY1293-414. c-d, The binding stoichiometry of YY1:G4 DNA was analyzed using EMSA. The quantification results showed the YY1-bound fraction of TAMRA-G4. The stoichiometry in binding of YY1 to G4 DNA matches with the theoretical curve in 1:1 binding stoichiometry. The data represent mean ± S.E.M. from 4 independent experiments.
Extended Data Fig. 7 YY1 promotes interactions between DNA elements containing its consensus sequence motifs, G4 DNA, or both.
a, A scheme depicting the in vitro proximity ligation assay for assessing the ability of YY1 to enhance DNA-DNA interactions involving G4 structures and/or YY1 consensus motifs. b, qPCR quantification results of the proximity ligation products formed between motifs, between G4 structures, and between motif and G4 structure in the presence or absence of YY1 protein. c, qPCR quantification results revealed the inability of YY1 in promoting the ligation between M4 (that is mutated sequence of G4 that can no longer fold into G4 structure) and G4, motif, or M4. d, qPCR quantification results of the proximity ligation products formed between G4 structures in the absence or presence of PDS or TMPyP4. The data represent mean ± S.E.M. of results from three independent experiments.
Extended Data Fig. 8 Dimerization of YY1 promotes long-range DNA looping.
a, SDS-PAGE for monitoring the purified recombinant truncated forms of YY1 that is incapable of dimerization, but able to discriminate G4 structure from single-stranded DNA. b, Gel filtration chromatography revealed that YY1∆231-290 (calculated monomer MW: 38.4 kDa) exists as a monomer, and GST-YY1∆231-290 is present as a dimer (calculated monomer MW: 66.3 kDa). c-d, Fluorescence anisotropy for monitoring the binding of YY1∆231-290 and GST-YY1∆231-290 protein with G4 structure and the corresponding mutated sequence (M4) derived from the MYC promoter. e, Proximity ligation assay showing that YY1 and GST-YY1∆231-290, but not YY1∆231-290, is capable of promoting ligation between G4 DNA sequences.
Extended Data Fig. 9 Regulation of gene expression by YY1-promoter G4 interactions.
a-c, Quantification of mRNA expression levels of MYC (a), SLC25A28 (b) and TMEM145 (c) genes after shRNA-mediated knockdown of YY1 and/or PDS/TMPyP4 treatment. Top of each panel shows read enrichments from BG4 ChIP-Seq and YY1 ChIP-Seq. The data represent mean ± S.E.M. of results from three independent experiments.
Extended Data Fig. 10 YY1-G4 binding participates in transcription regulation of EHD3 genes through long-range DNA looping.
a, Read enrichments obtained from BG4 ChIP-seq and YY1 ChIP-Seq experiments. b, Normalized interaction frequency between the two sites linked with a red arch in a in HEK293T cells with or without PDS/TMPyP4 treatment, as captured by YY1 HiChIP. c, Quantification results for the mRNA expression levels of EHD3 genes in HEK293T cells after knockdown of YY1 and/or with PDS/TMPyP4 treatment. The data represent mean ± S.E.M. of results from three independent experiments.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2 and Figs. 1–13.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Li, L., Williams, P., Ren, W. et al. YY1 interacts with guanine quadruplexes to regulate DNA looping and gene expression. Nat Chem Biol 17, 161–168 (2021). https://doi.org/10.1038/s41589-020-00695-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00695-1
This article is cited by
-
G-quadruplexes promote the motility in MAZ phase-separated condensates to activate CCND1 expression and contribute to hepatocarcinogenesis
Nature Communications (2024)
-
Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
Nature Communications (2024)
-
Spatiotemporal and global profiling of DNA–protein interactions enables discovery of low-affinity transcription factors
Nature Chemistry (2023)
-
The ALS/FTD-related C9orf72 hexanucleotide repeat expansion forms RNA condensates through multimolecular G-quadruplexes
Nature Communications (2023)
-
Interaction between non-coding RNAs, mRNAs and G-quadruplexes
Cancer Cell International (2022)