Abstract
The plant cuticle is the final barrier for volatile organic compounds (VOCs) to cross for release to the atmosphere, yet its role in the emission process is poorly understood. Here, using a combination of reverse-genetic and chemical approaches, we demonstrate that the cuticle imposes substantial resistance to VOC mass transfer, acting as a sink/concentrator for VOCs and hence protecting cells from the potentially toxic internal accumulation of these hydrophobic compounds. Reduction in cuticle thickness has differential effects on individual VOCs depending on their volatility, and leads to their internal cellular redistribution, a shift in mass transfer resistance sources and altered VOC synthesis. These results reveal that the cuticle is not simply a passive diffusion barrier for VOCs to cross, but plays the aforementioned complex roles in the emission process as an integral member of the overall VOC network.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
We declare that all the data supporting the funding of this study are available within the paper and its supplementary information files or from the corresponding author upon reasonable request. P. axillaris genomic data were obtained from http://solgenomics.net using the P. axillaris v.1.6.2 genome database. Source data are provided with this paper.
References
Zhao, D. F. et al. Environmental conditions regulate the impact of plants on cloud formation. Nat. Commun. 8, 14067 (2017).
Guenther, A. B. et al. The model of emissions of gases and aerosols from nature version 2.1 (MEGAN2.1): an extended and updated framework for modeling biogenic emissions. Geosci. Model Dev. 5, 1471–1492 (2012).
Dudareva, N., Negre, F., Nagegowda, D. A. & Orlova, I. Plant volatiles: recent advances and future perspectives. Crc. Crit. Rev. Plant Sci. 25, 417–440 (2006).
Dudareva, N., Klempien, A., Muhlemann, J. K. & Kaplan, I. Biosynthesis, function and metabolic engineering of plant volatile organic compounds. N. Phytol. 198, 16–32 (2013).
Staudt, M. & Bertin, N. Light and temperature dependence of the emission of cyclic and acyclic monoterpenes from holm oak (Quercus ilex L.) leaves. Plant, Cell Environ. 21, 385–395 (1998).
Penuelas, J. & Llusia, J. Plant VOC emissions: making use of the unavoidable. Trends Ecol. Evol. 19, 402–404 (2004).
Gouinguené, S. P. & Turlings, T. C. J. The effects of abiotic factors on induced volatile emissions in corn plants. Plant Physiol. 129, 1296–1307 (2002).
Lynch J. H., Pichersky, E. & Dudareva, N. F. in Biology of Plant Volatiles (eds. Pichersky, E. & Dudareva, N.) 147–164 (CRC Press, 2020).
Widhalm, J. R., Jaini, R., Morgan, J. A. & Dudareva, N. Rethinking how volatiles are released from plant cells. Trends Plant Sci. 20, 545–550 (2015).
Adebesin, F. et al. Emission of volatile organic compounds from petunia flowers is facilitated by an ABC transporter. Science 356, 1386–1388 (2017).
Goodwin, S. M. et al. Cuticle characteristics and volatile emissions of petals in Antirrhinum majus. Physiol. Plant. 117, 435–443 (2003).
Jeffree, C. E. in Annual Plant Reviews Vol. 23: Biology of the Plant Cuticle (eds. M. Riederer & C. Muller) 11–125 (Blackwell, 2006).
Martin, L. B. B. & Rose, J. K. C. There’s more than one way to skin a fruit: formation and functions of fruit cuticles. J. Exp. Bot. 65, 4639–4651 (2014).
Bernard, A. & Joubès, J. Arabidopsis cuticular waxes: advances in synthesis, export and regulation. Prog. Lipid Res. 52, 110–129 (2013).
Yeats, T. H. & Rose, J. K. C. The formation and function of plant cuticles. Plant Physiol. 163, 5–20 (2013).
Jenks, M. A., Eigenbrode, S. D. & Lemieux, B. Cuticular waxes of Arabidopsis. Arabidopsis Book 1, e0016 (2002).
Li-Beisson, Y. et al. Acyl-lipid metabolism. Arabidopsis Book 11, e0161 (2013).
Kerstiens, G. Water transport in plant cuticles: an update. J. Exp. Bot. 57, 2493–2499 (2006).
Howells, D. J., Hambrook, J. L. & Allenby, E. A. Uptake of some volatile alkyl methylphosphonofluoridates from the vapour phase by wheat plants. Pestic. Sci. 7, 349–354 (1976).
Sabljic, A., Guesten, H., Schoenherr, J. & Riederer, M. Modeling plant uptake of airborne organic chemicals. 1. Plant cuticle/water partitioning and molecular connectivity. Environ. Sci. Technol. 24, 1321–1326 (1990).
Umlauf, G., Hauk, H. & Reissinger, M. The distribution of semivolatile organic compounds in conifer needles following gas phase contamination. Chemosphere 28, 1689–1699 (1994).
Merk, S. & Riederer, M. Sorption of volatile C1 to C6 alkanols in plant cuticles. J. Exp. Bot. 48, 1095–1104 (1997).
Platts, J. A. & Abraham, M. H. Partition of volatile organic compounds from air and from water into plant cuticular matrix: an LFER analysis. Environ. Sci. Technol. 34, 318–323 (2000).
Boatright, J. et al. Understanding in vivo benzenoid metabolism in petunia petal tissue. Plant Physiol. 135, 1993–2011 (2004).
Verdonk, J. C. et al. Regulation of floral scent production in petunia revealed by targeted metabolomics. Phytochemistry 62, 997–1008 (2003).
Colquhoun, T. A. et al. Petunia floral volatile benzenoid/phenylpropanoid genes are regulated in a similar manner. Phytochemistry 71, 158–167 (2010).
Javelle, M. et al. Overexpression of the epidermis-specific homeodomain-leucine zipper IV transcription factor OUTER CELL LAYER1 in maize identifies target genes involved in lipid metabolism and cuticle biosynthesis. Plant Physiol. 154, 273–286 (2010).
Pighin, J. A. et al. Plant cuticular lipid export requires an ABC transporter. Science 306, 702–704 (2004).
Bird, D. et al. Characterization of Arabidopsis ABCG11/WBC11, an ATP binding cassette (ABC) transporter that is required for cuticular lipid secretion. Plant J. 52, 485–498 (2007).
Luo, B., Xue, X.-Y., Hu, W.-L., Wang, L.-J. & Chen, X.-Y. An ABC transporter gene of Arabidopsis thaliana, AtWBC11, is involved in cuticle development and prevention of organ fusion. Plant Cell Physiol. 48, 1790–1802 (2007).
McFarlane, H. E., Shin, J. J. H., Bird, D. A. & Samuels, A. L. Arabidopsis ABCG transporters, which are required for export of diverse cuticular lipids, dimerize in different combinations. Plant Cell 22, 3066–3075 (2010).
Panikashvili, D., Shi, J. X., Schreiber, L. & Aharoni, A. The Arabidopsis ABCG13 transporter is required for flower cuticle secretion and patterning of the petal epidermis. N. Phytol. 190, 113–124 (2011).
Panikashvili, D. et al. The Arabidopsis DSO/ABCG11 transporter affects cutin metabolism in reproductive organs and suberin in roots. Mol. Plant 3, 563–575 (2010).
Bombarely, A. et al. Insight into the evolution of the Solanaceae from the parental genomes of Petunia hybrida. Nat. Plants 2, 16074 (2016).
Widhalm, J. R. et al. Identification of a plastidial phenylalanine exporter that influences flux distribution through the phenylalanine biosynthetic network. Nat. Commun. 6, 8142 (2015).
Azuma, M. et al. A petal-specific InMYB1 promoter from Japanese morning glory: a useful tool for molecular breeding of floricultural crops. Plant Biotechnol. J. 14, 354–363 (2016).
Tanaka, T., Tanaka, H., Machida, C., Watanabe, M. & Machida, Y. A new method for rapid visualization of defects in leaf cuticle reveals five intrinsic patterns of surface defects in Arabidopsis. Plant J. 37, 139–146 (2004).
Rolny, N., Costa, L., Carrión, C. & Guiamet, J. J. Is the electrolyte leakage assay an unequivocal test of membrane deterioration during leaf senescence? Plant Physiol. Biochem. 49, 1220–1227 (2011).
Kolosova, N., Gorenstein, N., Kish, C. M. & Dudareva, N. Regulation of circadian methyl benzoate emission in diurnally and nocturnally emitting plants. Plant Cell 13, 2333–2347 (2001).
Parrish, S., Fleenor, J., Xu, S., Mello, C. & Fire, A. Functional anatomy of a dsRNA trigger. Mol. Cell 6, 1077–1087 (2000).
Segal, G., Song, R. & Messing, J. A new opaque variant of maize by a single dominant RNA-interference-inducing transgene. Genetics 165, 387–397 (2003).
Oshima, Y. et al. Novel vector systems to accelerate functional analysis of transcription factors using chimeric repressor gene-silencing technology (CRES-T). Plant Biotechnol. 28, 201–210 (2011).
Horsch, R. B. et al. A simple and general method for transferring genes into plants. Science 227, 1229–1231 (1985).
Klempien, A. et al. Contribution of CoA ligases to benzenoid biosynthesis in petunia flowers. Plant Cell 24, 2015–2030 (2012).
Qualley, A. V., Widhalm, J. R., Adebesin, F., Kish, C. M. & Dudareva, N. Completion of the core ß-oxidative pathway of benzoic acid biosynthesis in plants. Proc. Natl Acad. Sci. USA 109, 16383–16388 (2012).
Qian, Y. et al. Completion of the cytosolic post-chorismate phenylalanine biosynthetic pathway in plants. Nat. Commun. 10, 15 (2019).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Abràmoff, M. D., Magalhães, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Orlova, I. et al. Reduction of benzenoid synthesis in petunia flowers reveals multiple pathways to benzoic acid and enhancement in auxin transport. Plant Cell 18, 3458–3475 (2006).
Li-Beisson, Y. et al. Nanoridges that characterize the surface morphology of flowers require the synthesis of cutin polyester. Proc. Natl Acad. Sci. USA 106, 22008–22013 (2009).
Wang, Z.-Y., Xiong, L., Li, W., Zhu, J.-K. & Zhu, J. The plant cuticle is required for osmotic stress regulation of abscisic acid biosynthesis and osmotic stress tolerance in Arabidopsis. Plant Cell 23, 1971–1984 (2011).
Zhang, J.-Y. et al. Overexpression of WXP1, a putative Medicago truncatula AP2 domain-containing transcription factor gene, increases cuticular wax accumulation and enhances drought tolerance in transgenic alfalfa (Medicago sativa). Plant J. 42, 689–707 (2005).
Lynch, J. H. et al. Modulation of auxin formation by the cytosolic phenylalanine biosynthetic pathway. Nat. Chem. Biol. 16, 850–856 (2020).
Maeda, H. et al. RNAi suppression of arogenate dehydratase1 reveals that phenylalanine is synthesized predominantly via the arogenate pathway in petunia petals. Plant Cell 22, 832–849 (2010).
Boachon, B. et al. Natural fumigation as a mechanism for volatile transport between flower organs. Nat. Chem. Biol. 15, 583–588 (2019).
Arnon, D. I. Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol. 24, 1–15 (1949).
Yanagisawa, M. et al. Patterning mechanisms of cytoskeletal and cell wall systems during leaf trichome morphogenesis. Nat. Plants 1, 15014 (2015).
Press, W. H. Numerical Recipes: The Art of Scientific Computing (Cambridge Univ. Press, 2007).
Jaini, R., Wang, P., Dudareva, N., Chapple, C. & Morgan, J. A. Targeted metabolomics of the phenylpropanoid pathway in Arabidopsis thaliana using reversed phase liquid chromatography coupled with tandem mass spectrometry. Phytochem. Anal. 28, 267–276 (2017).
Acknowledgements
This work was supported by grant from the National Science Foundation no. IOS-1655438 to N.D. and J.A.M. and by the USDA National Institute of Food and Agriculture Hatch Project no. 177845 to N.D. We acknowledge the use of the imaging facilities of the Bindley Bioscience Center, a core facility of the NIH-funded Indiana Clinical and Translational Sciences Institute, for collection of confocal microscopy images. We thank Y. Oshima (National Institute of Advanced Industrial Science and Technology, Japan) for providing pDONOR_P4PIR-InMYB1pro and R4pGWB5_stop_HSP vectors. We thank R. Seiler and L. Mueller for technical assistance on SEM and TEM, respectively.
Author information
Authors and Affiliations
Contributions
B.B., J.A.M. and N.D. conceived the project and designed research. P.L. performed analysis of flower phenotypes, protein levels, seed production and germination rates, VOC emission and their internal pools, internal cellular VOC distribution over a 15 h period, toluidine and propidium iodide staining experiments, expression analysis and feeding experiments. S.R. performed wax profiling, VOC distribution and metabolic flux analyses, dewaxed experiments, water loss determination and analyzed rhythmicity of VOC production and accumulation. B.B. generated PhABCG12-RNAi lines and performed initial expression and metabolic profiling. J.H.L. performed analysis of sucrose and starch levels as well as flux through the shikimate pathway. A.D. run and analyzed samples for shikimate pathway flux experiments. S.M. analyzed respiration rate. P.L., S.R, B.B., J.H.L., A.D., S.M., J.A.M. and N.D. analyzed data. N.D. wrote the manuscript with contribution from all authors. All authors read and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effect of PhABCG12 downregulation on cuticle composition of 2-day-old petunia petals.
a, Wax composition by major constituent classes: fatty acids, primary alcohols, n-alkanes, and alkyl esters in 2-day-old flowers of wild type (WT), EV control and transgenic lines 7, 8, and 9 collected at 10 PM. Relative amount of each wax constituent is presented as a percentage of the total in their respective compound class. X axis represents the chain length (number of carbon atoms) for each constituent. Inset graphs show total amount of each class (µg/g DW). Data are means ± s.e.m. (n = 5 biological replicates). P values were determined by two-way ANOVA with the Tukey’s multiple comparisons test relative to the corresponding WT (black) and EV (blue) controls. b, Relative abundances of major constituent classes: fatty acids, primary alcohols, n-alkanes, and alkyl esters (shown in a) in wax of 2-day-old flowers of WT, EV control and transgenic lines 7, 8, and 9 collected at 10 PM. Data are means ± s.e.m. (n = 5 biological replicates).
Extended Data Fig. 2 Effect of PhABCG12 downregulation on petunia flower phenotype.
a, Representative 2-day-old flowers from wild type (WT), empty vector control (EV) and 3 independent PhABCG12-RNAi lines (7, 8 and 9). b, Corolla diameter, (c) flower weight and (d) corolla weight from WT, empty vector (EV)-transformed flowers and 3 independent PhABCG12-RNAi lines (7, 8 and 9). Data are means ± s.e.m. In b and c, n = 20 biological replicates for WT, n = 18 biological replicates for EV; and n = 9, 9 and 12 biological replicates for PhABCG12 lines 7, 8, and 9, respectively; in d, n = 19 biological replicates for WT, n = 14 biological replicates for EV; and n = 9, 7 and 9 biological replicates for PhABCG12 lines 7, 8, and 9, respectively. P values were determined by two-way ANOVA with the Tukey’s multiple comparisons test relative to the WT (black) and EV (blue) controls. Scale bar in a, 1 cm.
Extended Data Fig. 3 Histogram showing cuticle thickness distribution in wild-type and PhABCG12 petunia flowers.
Cuticle thickness probability distribution in epidermal cells of 2-day-old wild type (WT) and PhABCG12-RNAi line 9 flowers. To be consistent cuticle thickness was measured only in conical cells. The distribution is presented as the probability of different cuticle thickness values over 675 measurements. Data are means ± s.e.m. (n = 675 measurements). Measurements were taken from discreet locations in a minimum of 11 cells per genotype.
Extended Data Fig. 4 Effect of PhABCG12 downregulation on emissions and internal pools of individual benzenoid/phenylpropanoid volatiles in 2-day-old petunia flowers.
Emission rates (a) and internal pools (b) of individual VOCs in wild type (WT), empty vector (EV) control and three independent PhABCG12-RNAi lines (7, 8 and 9). Data are means ± s.e.m. (in a n = 6 biological replicates for all samples; and in b, n = 8 biological replicates for WT, n = 6 biological replicates for EV, PhABCG12-RNAi lines 7 and 8; and n = 9 biological replicates for PhABCG12-RNAi line 9). P values were determined by two-way ANOVA with the Tukey’s multiple comparisons test relative to the corresponding WT (black) and EV (blue) controls.
Extended Data Fig. 5 Metabolic flux analysis of benzenoid and phenylpropanoid VOCs from petunia flowers supplied with 150 mM 13C6-phenylalanine.
Metabolic flux maps of 2-day-old wild-type (a) and PhABCG12-9 (b) petunia petals fed with 150 mM 13C6-Phe. Fluxes were obtained from time-course measurements of label incorporation in the endogenous pools and headspace collections of Phe-derived VOCs. Arrow thickness reflects the relative value of fluxes normalized to the incoming flux (turnover of 13C6-Phe). BAlc, benzyl alcohol; BAld, benzaldehyde; BB, benzylbenzoate; Eug, eugenol; IEug, isoeugenol; MB, methylbenzoate; 2-PE, 2-phenylethanol; PEB, phenylethylbenzoate; and PhAld, phenylacetaldehyde. The total labeled biosynthetic (c) and emission (d) fluxes shown in (a) and (b). Flux values are given in nmol·g FW−1·h−1. Data are the mean ± s.e.m. of total biosynthetic and emission fluxes calculated from the regression of time-dependent VOC pools and emission collections. Regression was performed using all data points simultaneously from biological replicates to obtain flux distribution values. Standard error of calculated fluxes were determined based on the propagation of measurement standard errors (n = 3 biological replicates) and the model regression error. P value was determined by unpaired two-tailed Student’s t-test relative to wild type.
Extended Data Fig. 6 Pool sizes and isotopic abundances of total endogenous (internal pools) and exogenous (emitted) VOCs in control and PhABCG12-9 line petunia petals fed with 150 mM 13C6-Phe.
Total internal pools (a) and emissions (b) of VOCs from wild-type (WT) and PhABCG12-9 petunia corollas supplied with 150 mM 13C6-Phe for 4 h. Isotopic labeling of internal pools (c) and emitted volatiles (d) over 4 h, from 6 PM till 10 PM. Data are means ± s.e.m. (n = 3 biological replicates).
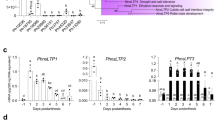
Extended Data Fig. 7 Effect of PhABCG12 downregulation on emission and biosynthetic fluxes of individual benzenoid/phenylpropanoid VOCs.
Emission and biosynthetic fluxes for individual benzenoid/phenylpropanoid VOCs in 2-day-old wild-type (WT) (a) and PhABCG12-9 (b) petunia flowers. BAlc, benzyl alcohol; IEug, isoeugenol; 2-PE, 2-phenylethanol; PhAld, phenylacetaldehyde. Data are means ± s.e.m. (n = 3 biological replicates).
Extended Data Fig. 8 Effect of PhABCG12 downregulation on cellular distribution of individual benzenoid/phenylpropanoid VOCs in 2-day-old petunia flowers.
a, Distribution of individual VOCs in wild-type (WT), empty vector control (EV) and PhABCG12-RNAi lines 7, 8 and 9. VOCs are shown in order of increasing volatility (left to right). Data are means ± s.e.m. (n = 4 biological replicates). P values were determined by unpaired two-tailed Student’s t-test relative to corresponding wild type. b, c, Relationships between distribution of individual VOCs in the cuticle and their corresponding (b) vapor pressure and (c) octanol-water partitioning coefficient. Data are means ± s.e.m. (n = 4 biological replicates). In b and c, Pearson correlation coefficients (R2) and corresponding P values are shown. BAlc, benzyl alcohol; BAld, benzaldehyde; BB, benzylbenzoate; IEug, isoeugenol; MB, methylbenzoate; 2-PE, 2-phenylethanol; PhAld, phenylacetaldehyde.
Extended Data Fig. 9 Total internal pools, emissions and distribution in cuticle of benzenoid/phenylpropanoid VOCs in 2-day-old petunia flowers around peak of emission.
Total internal pools (a), emissions (b) and distribution in cuticle (c) of VOCs collected from wild-type (WT) and PhABCG12-9 flowers over 15 h period. Data are means ± s.e.m. (n = 3 biological replicates in a and b; and n = 6 biological replicates in c). In c, PhABCG12-9 VOC distribution in cuticle was statistically different (P < 0.0001) from WT based on two-way ANOVA.
Extended Data Fig. 10 Analysis of metabolic potential of petunia flowers after dewaxing.
a, PAL activity detected in nondewaxed and dewaxed 2-day-old wild-type flowers in the beginning (6 PM) and the end (10 PM) of scent collection period. Data are means ± s.e.m. (n = 3 biological replicates). PAL activity was not statistically different between nondewaxed and dewaxed petals (P = 0.1476 and 0.2824 for 6 PM and 10 PM, respectively, determined by two-tailed Student’s t-test). b, Volatile emission from nondewaxed and dewaxed wild-type flowers upon feeding different (0 - 150 mM) Phe concentrations. c, VOC biosynthetic fluxes in nondewaxed and dewaxed wild-type flowers fed with 150 mM Phe. d, Internal VOC pools in nondewaxed and dewaxed wild-type flowers upon feeding different (0 -150 mM) Phe concentrations. e, Fold-change in VOC internal pools from 6 PM to 10 PM in nondewaxed and dewaxed wild-type flowers upon feeding different (0 - 150 mM) Phe concentrations. P values were determined by unpaired two-tailed Student’s t-test relative to corresponding non-dewaxed samples. Data in b - e are means ± s.e.m. (n = 4 biological replicates).
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Tables 1–5.
Source data
Source Data Fig. 1
Statistical Source Data for Fig.1
Source Data Fig. 2
Statistical Source Data for Fig.2
Source Data Fig. 3
Statistical Source Data for Fig.3
Source Data Fig. 4
Statistical Source Data for Fig.4
Source Data Fig. 5
Statistical Source Data for Fig.5
Source Data Extended Data Fig. 1
Statistical Source Data for Extended Data Fig.1
Source Data Extended Data Fig. 2
Statistical Source Data for Extended Data Fig.2
Source Data Extended Data Fig. 3
Statistical Source Data for Extended Data Fig.3
Source Data Extended Data Fig. 4
Statistical Source Data for Extended Data Fig.4
Source Data Extended Data Fig. 5
Statistical Source Data for Extended Data Fig.5
Source Data Extended Data Fig. 6
Statistical Source Data for Extended Data Fig.6
Source Data Extended Data Fig. 7
Statistical Source Data for Extended Data Fig.7
Source Data Extended Data Fig. 8
Statistical Source Data for Extended Data Fig.8
Source Data Extended Data Fig. 9
Statistical Source Data for Extended Data Fig.9
Source Data Extended Data Fig. 10
Statistical Source Data for Extended Data Fig.10
Rights and permissions
About this article
Cite this article
Liao, P., Ray, S., Boachon, B. et al. Cuticle thickness affects dynamics of volatile emission from petunia flowers. Nat Chem Biol 17, 138–145 (2021). https://doi.org/10.1038/s41589-020-00670-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00670-w
This article is cited by
-
Novel lignin-based extracellular barrier in glandular trichome
Nature Plants (2024)
-
Emission of floral volatiles is facilitated by cell-wall non-specific lipid transfer proteins
Nature Communications (2023)
-
A peroxisomal heterodimeric enzyme is involved in benzaldehyde synthesis in plants
Nature Communications (2022)
-
To protect and emit beauty
Nature Chemical Biology (2021)
-
Adaptive mechanisms of plant specialized metabolism connecting chemistry to function
Nature Chemical Biology (2021)