Abstract
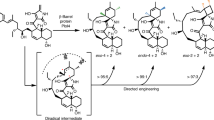
Cycloaddition reactions generate chemical complexity in a single step. Here we report the crystal structures of three homologous plant-derived cyclases involved in the biosynthesis of iboga and aspidosperma alkaloids. These enzymes act on the same substrate, named angryline, to generate three distinct scaffolds. Mutational analysis reveals how these highly similar enzymes control regio- and stereo-selectivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
O’Connor, S. E. & Maresh, J. J. Chemistry and biology of monoterpene indole alkaloid biosynthesis. Nat. Prod. Rep. 23, 532–472 (2006).
Caputi, L. et al. Missing enzymes in the biosynthesis of the anticancer drug vinblastine in Madagascar periwinkle. Science 60, 1235–1239 (2018).
Wenkert, E. Biosynthesis of indole alkaloids. The aspidosperma and iboga bases. J. Am. Chem. Soc. 84, 98–102 (1962).
Scott, A. I., Cherry, P. C. & Qureshi, A. A. Mechanisms of indole alkaloid biosynthesis. The Corynanthe-Strychnos relationship. J. Am. Chem. Soc. 91, 4932–4936 (1969).
Kutney, J. P., Ehret, C., Nelson, V. R. & Wigfield, D. C. Studies on indole alkaloid biosynthesis. II. J. Am. Chem. Soc. 90, 5929–5930 (1968).
Farrow, S. C. et al. Biosynthesis of an anti-addiction agent from the iboga plant. JACS 141, 12979–12983 (2019).
Scott, A. I. et al. Biosynthesis of the indole alkaloids. Acc. Chem. Res. 3, 151–157 (1970).
Qureshi, A. A. & Scott, A. I. Biogenetic-type synthesis of Iboga alkaloids: (±)-catharanthine. J. Chem. Soc. Chem. Commun. 16, 947–948 (1968).
Kuehne, M. E., Roland, D. M. & Haften, R. J. Studies in biomimetic alkaloid syntheses. 2. Synthesis of vincadifformine from tetrahydro-beta-carboline through a secodine intermediate. Org. Chem. 43, 3705–3710 (1978).
Marshall, S. D., Putterill, J. J., Plummer, K. M. & Newcomb, R. D. The carboxylesterase gene family from Arabidopsis thaliana. J. Mol. Evol. 57, 487–500 (2003).
Ileperuma, N. R. et al. High-resolution crystal structure of plant carboxylesterase AeCXE1, from Actinidia eriantha, and its complex with a high-affinity inhibitor paraoxon. Biochemistry 46, 1851–1859 (2007).
Andriamialisoa, R. Z., Langlois, N. & Langlois, Y. Preparation of 15-oxo-16-methoxycarbonyl-15, 20-dihydrocleavamine and coupling reaction with vindoline. Heterocycles 15, 245–250 (1981).
Langlois, N., Guéritte, F., Langlois, Y. & Potier, P. Application of a modification of the Polonovski reaction to the synthesis of vinblastine-type alkaloids. J. Am. Chem. Soc. 98, 7017–7024 (1976).
Buonora, P., Olsen, J.-C. & Oh, T. Recent developments in imino Diels–Alder reactions. Tetrahedron 57, 6099–6138 (2001).
Lichman, B. R., O’Connor, S. E. & Kries, H. Biocatalytic strategies towards [4+2] cycloadditions. Eur. J. Chem. 25, 6864–6877 (2019).
Tan, D. et al. Genome-mined diels–alderase catalyzes formation of the cis-octahydrodecalins of varicidin A and B. J. Am. Chem. Soc. 141, 769–773 (2019).
Jeon, B. S. et al. Investigation of the mechanism of the SpnF-catalyzed [4+2]-cycloaddition reaction in the biosynthesis of spinosyn A. Proc. Natl Acad. Sci. 114, 10408–10413 (2017).
Zhang, B. et al. Enzyme-catalysed [6+4] cycloadditions in the biosynthesis of natural products. Nature 568, 122–126 (2019).
Berrow, N. S. et al. A versatile ligation-independent cloning method suitable for high-throughput expression screening applications. Nucleic Acids Res. 35, e45 (2007).
Winter, G. et al. DIALS: implementation and evaluation of a new integration package. Acta Crystallogr. D. Biol. Crystallogr. 74, 85–97 (2018).
Kabsch, W. XDS. Acta Crystallogr. D. Biol. Crystallogr. 66, 125–132 (2010).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D. Biol. Crystallogr. 69, 1204–1214 (2013).
Potterton, L. et al. CCP4i2: the new graphical user interface to the CCP4 program suite. Acta Crystallogr. D. Biol. 74, 68–84 (2018).
Bunkoczi, G. & Read, R. J. Improvement of molecular-replacement models with Sculptor. Acta Crystallogr. D. Biol. Crystallogr. 67, 303–312 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D. Biol. Crystallogr. 53, 240–255 (1997).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D. Biol. Crystallogr. 62, 1002–1011 (2006).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nuc. Acids Res. 35, W375–W383 (2007).
McNicholas, S., Potterton, E., Wilson, K. S. & Noble, M. E. Presenting your structures: the CCP4mg molecular-graphics software. Acta Crystallogr. D. Biol. Crystallogr. 67, 386–394 (2011).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Acknowledgements
S.E.O. acknowledges ERC (788301). J.F. acknowledges financial support by the SMART BIOTECS alliance between the Technische Universität Braunschweig and the Leibniz Universität Hannover, supported by the Ministry for Science and Culture (MWK) of Lower Saxony, Germany. We acknowledge Diamond Light Source for access to beamline I03 under proposal MX13467 with support from the European Community’s Seventh Framework Program (No. FP7/2007–2013) under Grant Agreement No. 283570 (BioStruct-X).
Author information
Authors and Affiliations
Contributions
L.C. and S.E.O. conceived the project. D.M.L. managed all crystallography experiments. D.M.L., L.C., S.C.F. and C.E.M.S solved the crystal structures. L.C. and S.C.F. performed all biochemical experiments. K.B. and I.J.C.V. isolated substrates and products. J.F. solved the structure of the enzymatic substrates and developed the enzymatic mechanisms.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–4, Figs. 1–25 and Note.
Rights and permissions
About this article
Cite this article
Caputi, L., Franke, J., Bussey, K. et al. Structural basis of cycloaddition in biosynthesis of iboga and aspidosperma alkaloids. Nat Chem Biol 16, 383–386 (2020). https://doi.org/10.1038/s41589-019-0460-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0460-x