Abstract
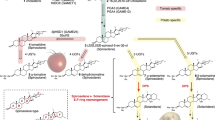
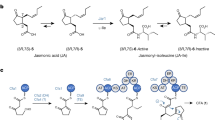
In plants, lineage-specific metabolites can be created by activities derived from the catalytic promiscuity of ancestral proteins, although examples of recruiting detoxification systems to biosynthetic pathways are scarce. The ubiquitous glyoxalase (GLX) system scavenges the cytotoxic methylglyoxal, in which GLXI isomerizes the α-hydroxy carbonyl in the methylglyoxal–glutathione adduct for subsequent hydrolysis. We show that GLXIs across kingdoms are more promiscuous than recognized previously and can act as aromatases without cofactors. In cotton, a specialized GLXI variant, SPG, has lost its GSH-binding sites and organelle-targeting signal, and evolved to aromatize cyclic sesquiterpenes bearing α-hydroxyketones to synthesize defense compounds in the cytosol. Notably, SPG is able to transform acetylated deoxynivalenol, the prevalent mycotoxin contaminating cereals and foods. We propose that detoxification enzymes are a valuable source of new catalytic functions and SPG, a standalone enzyme catalyzing complex reactions, has potential for toxin degradation, crop engineering and design of novel aromatics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that all relevant data supporting the findings of this study are available within the paper and its Supplementary Information. The FLNC reads and HiSeq transcriptomic reads generated in this study have been deposited in the NCBI SRA database under accession number PRJNA493958. Moreover, datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Code availability
All code used in this study is available from the corresponding author upon reasonable request.
References
Maeda, H. & Dudareva, N. The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu. Rev. Plant Biol. 63, 73–105 (2012).
Dixon, R. A. Natural products and plant disease resistance. Nature 411, 843–847 (2001).
Zhang, Y. et al. Multi-level engineering facilitates the production of phenylpropanoid compounds in tomato. Nat. Commun. 6, 8635 (2015).
Caputi, L. et al. Missing enzymes in the biosynthesis of the anticancer drug vinblastine in Madagascar periwinkle. Science 360, 1235–1239 (2018).
Weng, J. K., Philippe, R. N. & Noel, J. P. The rise of chemodiversity in plants. Science 336, 1667–1670 (2012).
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009).
Hovatta, I. et al. Glyoxalase 1 and glutathione reductase 1 regulate anxiety in mice. Nature 438, 662–666 (2005).
Rabbani, N. & Thornalley, P. J. The critical role of methylglyoxal and glyoxalase 1 in diabetic nephropathy. Diabetes 63, 50–52 (2014).
Morcos, M. et al. Glyoxalase-1 prevents mitochondrial protein modification and enhances lifespan in Caenorhabditis elegans. Aging Cell 7, 260–269 (2008).
Sankaranarayanan, S. et al. Glyoxalase goes green: the expanding roles of glyoxalase in plants. Int. J. Mol. Sci. 18, 898 (2017).
Sankaranarayanan, S., Jamshed, M. & Samuel, M. A. Degradation of glyoxalase I in Brassica napus stigma leads to self-incompatibility response. Nat. Plants 1, 15185 (2015).
Thornalley, P. J. Glyoxalase I–structure, function and a critical role in the enzymatic defence against glycation. Biochem. Soc. Trans. 31, 1343–1348 (2003).
Gadelha, I. C., Fonseca, N. B., Oloris, S. C., Melo, M. M. & Soto-Blanco, B. Gossypol toxicity from cottonseed products. ScientificWorldJournal 2014, 231635 (2014).
Keshmiri-Neghab, H. & Goliaei, B. Therapeutic potential of gossypol: an overview. Pharm. Biol. 52, 124–128 (2014).
Tian, X. et al. A gossypol biosynthetic intermediate disturbs plant defence response. Philos. Trans. R. Soc. Lond. B 374, 20180319 (2019).
Tian, X. et al. Characterization of gossypol biosynthetic pathway. Proc. Natl Acad. Sci. USA 115, E5410–E5418 (2018).
Knutsen, H. K. et al. Risks to human and animal health related to the presence of deoxynivalenol and its acetylated and modified forms in food and feed. EFSA J. 15, e04718 (2017).
Pestka, J. J. Deoxynivalenol: toxicity, mechanisms and animal health risks. Anim. Feed Sci. Technol. 137, 283–298 (2007).
Ma, D. et al. Genetic basis for glandular trichome formation in cotton. Nat. Commun. 7, 10456 (2016).
Wang, J. Y. et al. VdNEP, an elicitor from Verticillium dahliae, induces cotton plant wilting. Appl. Environ. Microbiol. 70, 4989–4995 (2004).
Lusas, E. W. & Jividen, G. M. Glandless cottonseed: a review of the first 25 years of processing and utilization research. J. Am. Oil Chem. Soc. 64, 839–854 (1987).
Khersonsky, O. & Tawfik, D. S. Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu. Rev. Biochem. 79, 471–505 (2010).
Gerlt, J. A. et al. Enzyme function initiative-enzyme similarity tool (EFI-EST): a web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 1854, 1019–1037 (2015).
Cameron, A. D., Olin, B., Ridderstrom, M., Mannervik, B. & Jones, T. A. Crystal structure of human glyoxalase I–evidence for gene duplication and 3D domain swapping. EMBO J. 16, 3386–3395 (1997).
Cameron, A. D. et al. Reaction mechanism of glyoxalase I explored by an X-ray crystallographic analysis of the human enzyme in complex with a transition state analogue. Biochemistry 38, 13480–13490 (1999).
Ridderstrom, M., Cameron, A. D., Jones, T. A. & Mannervik, B. Mutagenesis of residue 157 in the active site of human glyoxalase I. Biochem. J. 328, 231–235 (1997).
Paterson, A. H. et al. Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492, 423–427 (2012).
Li, F. et al. Genome sequence of cultivated upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 33, 524–530 (2015).
Zhang, T. et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537 (2015).
Schmitz, J. et al. Defense against reactive carbonyl species involves at least three subcellular compartments where individual components of the system respond to cellular sugar status. Plant Cell 29, 3234–3254 (2017).
Wu, S. et al. Redirection of cytosolic or plastidic isoprenoid precursors elevates terpene production in plants. Nat. Biotechnol. 24, 1441–1447 (2006).
Means, G. D. et al. Structural analysis of the gene encoding human aromatase cytochrome P-450, the enzyme responsible for estrogen biosynthesis. J. Biol. Chem. 264, 19385–19391 (1989).
Wu, X. et al. Biochemical characterization of TASSELSEED 2, an essential plant short-chain dehydrogenase/reductase with broad spectrum activities. FEBS J. 274, 1172–1182 (2007).
Sonawane, P. D. et al. Short-chain dehydrogenase/reductase governs steroidal specialized metabolites structural diversity and toxicity in the genus Solanum. Proc. Natl Acad. Sci. USA 115, E5419–E5428 (2018).
Ji, C., Fan, Y. & Zhao, L. Review on biological degradation of mycotoxins. Anim. Nutr. 2, 127–133 (2016).
Ibrahim, S. R. M. & Mohamed, G. A. Naturally occurring naphthalenes: chemistry, biosynthesis, structural elucidation, and biological activities. Phytochem. Rev. 15, 279–295 (2016).
Taura, F. et al. A novel class of plant type III polyketide synthase involved in orsellinic acid biosynthesis from Rhododendron dauricum. Front. Plant Sci. 7, 1452 (2016).
Tzin, V. & Galili, G. The biosynthetic pathways for shikimate and aromatic amino acids in Arabidopsis thaliana. Arabidopsis Book 8, e0132 (2010).
Sharifi, N. Minireview: androgen metabolism in castration-resistant prostate cancer. Mol. Endocrinol. 27, 708–714 (2013).
Brown, G. D. The biosynthesis of artemisinin (Qinghaosu) and the phytochemistry of Artemisia annua L. (Qinghao). Molecules 15, 7603–7698 (2010).
Czechowski, T. et al. Artemisia annua mutant impaired in artemisinin synthesis demonstrates importance of nonenzymatic conversion in terpenoid metabolism. Proc. Natl Acad. Sci. USA 113, 15150–15155 (2016).
Rabbani, N., Xue, M. & Thornalley, P. J. Activity, regulation, copy number and function in the glyoxalase system. Biochem. Soc. Trans. 42, 419–424 (2014).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Hackl, T., Hedrich, R., Schultz, J. & Forster, F. Proovread: large-scale high-accuracy PacBio correction through iterative short read consensus. Bioinformatics 30, 3004–3011 (2014).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. EdgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Shan, C. M. et al. Control of cotton fibre elongation by a homeodomain transcription factor GhHOX3. Nat. Commun. 5, 5519 (2014).
Luo, P., Wang, Y. H., Wang, G. D., Essenberg, M. & Chen, X. Y. Molecular cloning and functional identification of (+)-delta-cadinene-8-hydroxylase, a cytochrome P450 mono-oxygenase (CYP706B1) of cotton sesquiterpene biosynthesis. Plant J. 28, 95–104 (2001).
Pompon, D., Louerat, B., Bronine, A. & Urban, P. Yeast expression of animal and plant P450s in optimized redox environments. Methods Enzymol. 272, 51–64 (1996).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We thank W. Hu, S. Bu and Y. Liu for help with GC–MS, NMR and Q-TOF analyses and X. Hao, B. Xu, B. Yang, C. Shi, Y. Hu, Y. Li, L. Chen and K. Zhai for their kind and generous help. We also thank D. Nelson for naming the CYP protein. The research was supported by grants from the National Natural Science Foundation of China (Nos. 31788103, 31690092 to X.-Y.C. and No. 31700263 to J.X.L.), the Ministry of Agriculture of China (grant No. 2016ZX08010002-005 to L.J.W.), the Ministry of Science and Technology of China (grant No. 2016YFD0100500 to L.J.W.) and the Chinese Academy of Sciences (grant Nos. XDB11030000, QYZDY-SSW-SMC026 and 153D31KYSB20160074 to X.-Y.C.).
Author information
Authors and Affiliations
Contributions
J.-Q.H., X.F., X.T. and X.-Y.C. designed and managed the study. C.M., X.F., X.T., W.-M.S., Q.Z. and L.-J.W. discussed results and provided advice. J.-Q.H. isolated genes and characterized enzymes. J.-Q.H., X.T., J.-L.L., X.-X.G., R.A. and F.-Y.C. isolated compounds and performed LC–MS and GC–MS analyses. X.F. and Z.F. analyzed the NMR data. J.-Q.H., P.C. and Q.Z. performed bioinformatic analysis. J.-X.L. modeled the enzymes. X.-Y.C., J.-Q.H., X.F. and C.M. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary Tables 1 and 2, Figs. 1–9 and Note.
Rights and permissions
About this article
Cite this article
Huang, JQ., Fang, X., Tian, X. et al. Aromatization of natural products by a specialized detoxification enzyme. Nat Chem Biol 16, 250–256 (2020). https://doi.org/10.1038/s41589-019-0446-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0446-8
This article is cited by
-
Genome-wide identification and expression analyses of CYP450 genes in sweet potato (Ipomoea batatas L.)
BMC Genomics (2024)
-
Dirigent gene editing of gossypol enantiomers for toxicity-depleted cotton seeds
Nature Plants (2023)
-
Gossypol and related compounds are produced and accumulate in the aboveground parts of the cotton plant, independent of roots as the source
Planta (2023)
-
New function of gossypol, a natural product of cotton
Journal of Cotton Research (2022)
-
Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium
BMC Genomics (2020)