Abstract
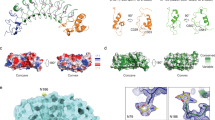
Polypeptide GalNAc-transferase T3 (GalNAc-T3) regulates fibroblast growth factor 23 (FGF23) by O-glycosylating Thr178 in a furin proprotein processing motif RHT178R↓S. FGF23 regulates phosphate homeostasis and deficiency in GALNT3 or FGF23 results in hyperphosphatemia and familial tumoral calcinosis. We explored the molecular mechanism for GalNAc-T3 glycosylation of FGF23 using engineered cell models and biophysical studies including kinetics, molecular dynamics and X-ray crystallography of GalNAc-T3 complexed to glycopeptide substrates. GalNAc-T3 uses a lectin domain mediated mechanism to glycosylate Thr178 requiring previous glycosylation at Thr171. Notably, Thr178 is a poor substrate site with limiting glycosylation due to substrate clashes leading to destabilization of the catalytic domain flexible loop. We suggest GalNAc-T3 specificity for FGF23 and its ability to control circulating levels of intact FGF23 is achieved by FGF23 being a poor substrate. GalNAc-T3’s structure further reveals the molecular bases for reported disease-causing mutations. Our findings provide an insight into how GalNAc-T isoenzymes achieve isoenzyme-specific nonredundant functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hurtado-Guerrero, R. Recent structural and mechanistic insights into protein O-GalNAc glycosylation. Biochem Soc. Trans. 44, 61–67 (2016).
Bennett, E. P. et al. Control of mucin-type O-glycosylation: a classification of the polypeptide GalNAc-transferase gene family. Glycobiology 22, 736–756 (2012).
de Las Rivas, M., Lira-Navarrete, E., Gerken, T. A. & Hurtado-Guerrero, R. Polypeptide GalNAc-Ts: from redundancy to specificity. Curr. Opin. Struct. Biol. 56, 87–96 (2019).
Lira-Navarrete, E. et al. Substrate-guided front-face reaction revealed by combined structural snapshots and metadynamics for the polypeptide N-acetylgalactosaminyltransferase 2. Angew. Chem. Int. Ed. Engl. 53, 8206–8210 (2014).
Gerken, T. A. et al. Emerging paradigms for the initiation of mucin-type protein O-glycosylation by the polypeptide GalNAc transferase family of glycosyltransferases. J. Biol. Chem. 286, 14493–14507 (2011).
Gerken, T. A. et al. The lectin domain of the polypeptide GalNAc transferase family of glycosyltransferases (ppGalNAc Ts) acts as a switch directing glycopeptide substrate glycosylation in an N- or C-terminal direction, further controlling mucin type O-glycosylation. J. Biol. Chem. 288, 19900–19914 (2013).
Revoredo, L. et al. Mucin-type O-glycosylation is controlled by short- and long-range glycopeptide substrate recognition that varies among members of the polypeptide GalNAc transferase family. Glycobiology 26, 360–376 (2016).
Lira-Navarrete, E. et al. Dynamic interplay between catalytic and lectin domains of GalNAc-transferases modulates protein O-glycosylation. Nat. Commun. 6, 6937 (2015).
Rivas, M. L. et al. The interdomain flexible linker of the polypeptide GalNAc transferases dictates their long-range glycosylation preferences. Nat. Commun. 8, 1959 (2017).
de Las Rivas, M. et al. Structural analysis of a GalNAc-T2 mutant reveals an induced-fit catalytic mechanism for GalNAc-Ts. Chemistry 24, 8382–8392 (2018).
Topaz, O. et al. Mutations in GALNT3, encoding a protein involved in O-linked glycosylation, cause familial tumoral calcinosis. Nat. Genet. 36, 579–581 (2004).
Kato, K. et al. Polypeptide GalNAc-transferase T3 and familial tumoral calcinosis. Secretion of fibroblast growth factor 23 requires O-glycosylation. J. Biol. Chem. 281, 18370–18377 (2006).
White, K. E. et al. Autosomal dominant hypophosphataemic rickets is associated with mutations in FGF23. Nat. Genet 26, 345–348 (2000).
Benet-Pages, A., Orlik, P., Strom, T. M. & Lorenz-Depiereux, B. An FGF23 missense mutation causes familial tumoral calcinosis with hyperphosphatemia. Hum. Mol. Genet. 14, 385–390 (2005).
Simpson, M. A. et al. Mutations in FAM20C also identified in non-lethal osteosclerotic bone dysplasia. Clin. Genet. 75, 271–276 (2009).
Tagliabracci, V. S. et al. Dynamic regulation of FGF23 by Fam20C phosphorylation, GalNAc-T3 glycosylation, and furin proteolysis. Proc. Natl Acad. Sci. USA 111, 5520–5525 (2014).
Urakawa, I. et al. Klotho converts canonical FGF receptor into a specific receptor for FGF23. Nature 444, 770–774 (2006).
Bennett, E. P., Hassan, H. & Clausen, H. cDNA cloning and expression of a novel human UDP-N-acetyl-alpha-d-galactosamine. Polypeptide N-acetylgalactosaminyltransferase, GalNAc-t3. J. Biol. Chem. 271, 17006–17012 (1996).
Kong, Y. et al. Probing polypeptide GalNAc-transferase isoform substrate specificities by in vitro analysis. Glycobiology 25, 55–65 (2015).
Schjoldager, K. T. et al. Deconstruction of O-glycosylation–GalNAc-T isoforms direct distinct subsets of the O-glycoproteome. EMBO Rep. 16, 1713–1722 (2015).
Khetarpal, S. A. et al. Loss of function of GALNT2 lowers high-density lipoproteins in humans, nonhuman primates, and rodents. Cell Metab. 24, 234–245 (2016).
Wang, S. et al. Site-specific O-glycosylation of members of the low-density lipoprotein receptor superfamily enhances ligand interactions. J. Biol. Chem. 293, 7408–7422 (2018).
Yoshimura, Y. et al. Elucidation of the sugar recognition ability of the lectin domain of UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase 3 by using unnatural glycopeptide substrates. Glycobiology 22, 429–438 (2012).
Frishberg, Y. et al. Hyperostosis-hyperphosphatemia syndrome: a congenital disorder of O-glycosylation associated with augmented processing of fibroblast growth factor 23. J. Bone Min. Res 22, 235–242 (2007).
Chen, G. et al. alpha-Klotho is a non-enzymatic molecular scaffold for FGF23 hormone signalling. Nature 553, 461–466 (2018).
Hagen, F. K., Hazes, B., Raffo, R., deSa, D. & Tabak, L. A. Structure-function analysis of the UDP-N-acetyl-d-galactosamine:polypeptide N-acetylgalactosaminyltransferase. Essential residues lie in a predicted active site cleft resembling a lactose repressor fold. J. Biol. Chem. 274, 6797–6803 (1999).
Kozarsky, K., Kingsley, D. & Krieger, M. Use of a mutant cell line to study the kinetics and function of O-linked glycosylation of low density lipoprotein receptors. Proc. Natl Acad. Sci. USA 85, 4335–4339 (1988).
Bar-Even, A. et al. The moderately efficient enzyme: evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 50, 4402–4410 (2011).
de Las Rivas, M. et al. Structural and mechanistic insights into the catalytic-domain-mediated short-range glycosylation preferences of GalNAc-T4. ACS Cent. Sci. 4, 1274–1290 (2018).
Lira-Navarrete, E. et al. Structural insights into the mechanism of protein O-fucosylation. PLoS ONE 6, e25365 (2011).
Ghirardello, M. et al. Glycomimetics targeting glycosyltransferases: synthetic, computational and structural studies of less-polar conjugates. Chemistry 22, 7215–7224 (2016).
Yamamoto, H. et al. Posttranslational processing of FGF23 in osteocytes during the osteoblast to osteocyte transition. Bone 84, 120–130 (2016).
Bennett, E. P. et al. Cloning and characterization of a close homologue of human UDP-N-acetyl-alpha-d-galactosamine: polypeptide N-acetylgalactosaminyltransferase-T3, designated GalNAc-T6. Evidence for genetic but not functional redundancy. J. Biol. Chem. 274, 25362–25370 (1999).
Joshi, H. J. et al. Glycosyltransferase genes that cause monogenic congenital disorders of glycosylation are distinct from glycosyltransferase genes associated with complex diseases. Glycobiology 28, 284–294 (2018).
Minisola, S. et al. Tumour-induced osteomalacia. Nat. Rev. Dis. Prim. 3, 17044 (2017).
Rafaelsen, S., Johansson, S., Raeder, H. & Bjerknes, R. Long-term clinical outcome and phenotypic variability in hyperphosphatemic familial tumoral calcinosis and hyperphosphatemic hyperostosis syndrome caused by a novel GALNT3 mutation; case report and review of the literature. BMC Genet. 15, 98 (2014).
Ramnitz, M. S., Gafni, R. I. & Collins, M. T. Hyperphosphatemic familial tumoral calcinosis. in GeneReviews (R) (eds Adams, M. P. et al.) https://www.ncbi.nlm.nih.gov/books/NBK476672/ (Univ. Wash., Seattle, 2018).
Takashi, Y. et al. Activation of unliganded FGF receptor by extracellular phosphate potentiates proteolytic protection of FGF23 by its O-glycosylation. Proc. Natl Acad. Sci. USA 116, 11418–11427 (2019).
Hintze, J. et al. Probing the contribution of individual polypeptide GalNAc-transferase isoforms to the O-glycoproteome by inducible expression in isogenic cell lines. J. Biol. Chem. 293, 19064–19077 (2018).
Wandall, H. H. et al. The lectin domains of polypeptide GalNAc-transferases exhibit carbohydrate-binding specificity for GalNAc: lectin binding to GalNAc-glycopeptide substrates is required for high density GalNAc-O-glycosylation. Glycobiology 17, 374–387 (2007).
Steentoft, C. et al. A validated collection of mouse monoclonal antibodies to human glycosyltransferases functioning in mucin-type O-glycosylation. Glycobiology 29, 645–656 (2019).
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Lonowski, L. A. et al. Genome editing using FACS enrichment of nuclease-expressing cells and indel detection by amplicon analysis. Nat. Protoc. 12, 581–603 (2017).
Wandall, H. H. et al. Substrate specificities of three members of the human UDP-N-acetyl-alpha-d-galactosamine: polypeptide N-acetylgalactosaminyltransferase family, GalNAc-T1, -T2, and -T3. J. Biol. Chem. 272, 23503–23514 (1997).
Kabsch, W. Xds. Acta Crystallogr. D. 66, 125–132 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D. 67, 235–242 (2011).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D. 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. 60, 2126–2132 (2004).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D. 67, 355–367 (2011).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Plattner, C., Hofener, M. & Sewald, N. One-pot azidochlorination of glycals. Org. Lett. 13, 545–547 (2011).
Horcas, I. et al. WSXM: a software for scanning probe microscopy and a tool for nanotechnology. Rev. Sci. Instrum. 78, 013705 (2007).
Lostao, A., Peleato, M. L., Gomez-Moreno, C. & Fillat, M. F. Oligomerization properties of FurA from the cyanobacterium Anabaena sp. PCC 7120: direct visualization by in situ atomic force microscopy under different redox conditions. Biochim. Biophys. Acta 1804, 1723–1729 (2010).
Sun, T., Lin, F. H., Campbell, R. L., Allingham, J. S. & Davies, P. L. An antifreeze protein folds with an interior network of more than 400 semi-clathrate waters. Science 343, 795–798 (2014).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Tria, G., Mertens, H. D., Kachala, M. & Svergun, D. I. Advanced ensemble modelling of flexible macromolecules using X-ray solution scattering. IUCr. J. 2, 207–217 (2015).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Bernado, P., Mylonas, E., Petoukhov, M. V., Blackledge, M. & Svergun, D. I. Structural characterization of flexible proteins using small-angle X-ray scattering. J. Am. Chem. Soc. 129, 5656–5664 (2007).
Acknowledgements
We thank the Diamond Light Source (Oxford) synchrotron beamline I24 (experiment nos. MX14739-6 and MX14739-11) and the SOLEIL synchrotron (Gif-sur-Yvette) SWING beamline (experiment nos. 99170088). We thank ARAID, MEC (grant no. CTQ2013-44367-C2-2-P, BFU2016-75633-P and RTI2018-099592-B-C21), the National Institutes of Health (grant no. GM113534 and instrument grant no. GM113534-01S), the Danish National Research Foundation (grant no. DNRF107), the FCT-Portugal (grant no. UID/Multi/04378/2013) and Gobierno de Aragón (grant nos. E34_R17, E35_17R and LMP58_18) with FEDER (grant no. 2014-2020) funds for ‘Building Europe from Aragón’ for financial support. I.C. thanks the Universidad de La Rioja for the FPI grant. F.M. and H.C. thank FCT-Portugal for IF Investigator (IF/00780/2015), PTDC/BIA-MIB/31028/2017 and UID/Multi/04378/2019 projects, and PTNMR (grant no. ROTEIRO/0031/2013 and PINFRA/22161/2016). P.B. acknowledges support from the Labex EpiGenMed, an ‘Investissements d’avenir’ program (grant no. ANR-10-LABX-12-01). The CBS (Montpellier) is a member of France-BioImaging (FBI, ANR-10-INBS-04-01) and the French Infrastructure for Integrated Structural Biology (FRISBI, ANR-10-INBS-05). The research leading to these results has also received funding from the FP7 (2007–2013) under BioStruct-X (grant agreement nos. 283570 and BIOSTRUCTX_5186). We also thank I. Echániz for technical support and K. Moremen from the University of Georgia, Complex Carbohydrate Research Center, for supplying the pGEn2-HsGalNAc-T6 and pGEn2-HsGalNAc-T12 plasmids.
Author information
Authors and Affiliations
Contributions
R.H.-G. designed the crystallization construct and solved the crystal structures. M.R. and R.H.-G. purified the enzymes, crystallized the complexes and refined the crystal structures. I.C. and F.C. synthesized the glycopeptides. F.C. performed the MD simulations. H.Coelho and F.M. performed and analyzed the NMR experiments. T.A.G. and E.J.P.D. performed the kinetic studies together with the Edman amino acid sequencing. Y.N., K.K. and R.M. performed the experiments in cells and did the MALDI–TOF MS mass spectrometry experiments. P.H. and A.L. performed the AFM studies. A.T. and P.B. performed the SAXS experiments. L.C.-L. conducted the expression and purification of GalNAc-T6 and T12 in HEK293 cells. L.H. identified the GalNAc-T3 mutations associated to disease. R.H.-G., T.A.G. and H.Clausen wrote the article with the other authors’ contributions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs 1–17 and Tables 1–6.
Supplementary Video 1
A 500 ns MD simulation of TgGalNAc-T3 complexed to UDP-Mn+2 and FGF23c in explicit water
Supplementary Video 2
A 500 ns MD simulation of HsGalNAc-T4 complexed to UDP-Mn+2 and FGF23c in explicit water
Supplementary Video 3
A 500 ns MD simulation of HsGalNAc-T6 complexed to UDP-Mn+2 and FGF23c in explicit water
Supplementary Video 4
A 500 ns MD simulation of HsGalNAc-T12 complexed to UDP-Mn+2 and FGF23c in explicit water
Rights and permissions
About this article
Cite this article
de las Rivas, M., Paul Daniel, E.J., Narimatsu, Y. et al. Molecular basis for fibroblast growth factor 23 O-glycosylation by GalNAc-T3. Nat Chem Biol 16, 351–360 (2020). https://doi.org/10.1038/s41589-019-0444-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0444-x
This article is cited by
-
Galectins use N-glycans of FGFs to capture growth factors at the cell surface and fine-tune their signaling
Cell Communication and Signaling (2023)
-
Regulation of FGF23 production and phosphate metabolism by bone–kidney interactions
Nature Reviews Nephrology (2023)
-
Discovery of a lectin domain that regulates enzyme activity in mouse N-acetylglucosaminyltransferase-IVa (MGAT4A)
Communications Biology (2022)
-
Structural basis for the synthesis of the core 1 structure by C1GalT1
Nature Communications (2022)
-
Polypeptide N-acetylgalactosamine transferase 3: a post-translational writer on human health
Journal of Molecular Medicine (2022)