Abstract
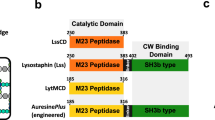
Lysostaphin is a bacteriolytic enzyme targeting peptidoglycan, the essential component of the bacterial cell envelope. It displays a very potent and specific activity toward staphylococci, including methicillin-resistant Staphylococcus aureus. Lysostaphin causes rapid cell lysis and disrupts biofilms, and is therefore a therapeutic agent of choice to eradicate staphylococcal infections. The C-terminal SH3b domain of lysostaphin recognizes peptidoglycans containing a pentaglycine crossbridge and has been proposed to drive the preferential digestion of staphylococcal cell walls. Here we elucidate the molecular mechanism underpinning recognition of staphylococcal peptidoglycan by the lysostaphin SH3b domain. We show that the pentaglycine crossbridge and the peptide stem are recognized by two independent binding sites located on opposite sides of the SH3b domain, thereby inducing a clustering of SH3b domains. We propose that this unusual binding mechanism allows synergistic and structurally dynamic recognition of S. aureus peptidoglycan and underpins the potent bacteriolytic activity of this enzyme.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Structural data have been deposited in the PDB with coordinate accession numbers 6RK4 (high-resolution set) and 6RJE (home source set). All other data generated or analyzed during this study are included in this article and its supplementary information or are available from the corresponding authors upon request.
References
Schindler, C. A. & Schuhardt, V. T. Lysostaphin: a new bacteriolytic agent for the Staphylococcus. Proc. Natl Acad. Sci. USA 51, 414–421 (1964).
Schindler, C. A. & Schuhardt, V. T. Purification and properties of Lysostaphin—a lytic agent for Staphylococcus aureus. Biochim. Biophys. Acta 97, 242–250 (1965).
Thumm, G. & Götz, F. Studies on prolysostaphin processing and characterization of the lysostaphin immunity factor (Lif) of Staphylococcus simulans biovar staphylolyticus. Mol. Microbiol 23, 1251–1265 (1997).
Wu, J. A., Kusuma, C., Mond, J. J. & Kokai-Kun, J. F. Lysostaphin disrupts Staphylococcus aureus and Staphylococcus epidermidis biofilms on artificial surfaces. Antimicrob. Agents Chemother. 47, 3407–3414 (2003).
Climo, M. W., Patron, R. L., Goldstein, B. P. & Archer, G. L. Lysostaphin treatment of experimental methicillin-resistant Staphylococcus aureus aortic valve endocarditis. Antimicrob. Agents Chemother. 42, 1355–1360 (1998).
Dajcs, J. J. et al. Lysostaphin treatment of methicillin-resistant Staphylococcus aureus keratitis in the rabbit. Invest Ophthalmol. Vis. Sci. 41, 1432–1437 (2000).
Johnson, C. T. et al. Hydrogel delivery of lysostaphin eliminates orthopedic implant infection by Staphylococcus aureus and supports fracture healing. Proc. Natl Acad. Sci. USA 115, E4960–E4969 (2018).
Kiri, N., Archer, G. & Climo, M. W. Combinations of lysostaphin with β-lactams are synergistic against oxacillin-resistant Staphylococcus epidermidis. Antimicrob. Agents Chemother. 46, 2017–2020 (2002).
Kokai-Kun, J. F., Chanturiya, T. & Mond, J. J. Lysostaphin as a treatment for systemic Staphylococcus aureus infection in a mouse model. J. Antimicrob. Chemother. 60, 1051–1059 (2007).
Kokai-Kun, J. F., Walsh, S. M., Chanturiya, T. & Mond, J. J. Lysostaphin cream eradicates Staphylococcus aureus nasal colonization in a cotton rat model. Antimicrob. Agents Chemother. 47, 1589–1597 (2003).
Satishkumar, R. et al. Evaluation of the antimicrobial activity of lysostaphin-coated hernia repair meshes. Antimicrob. Agents Chemother. 55, 4379–4385 (2011).
Blazanovic, K. et al. Structure-based redesign of lysostaphin yields potent antistaphylococcal enzymes that evade immune cell surveillance. Mol. Ther. Methods Clin. Dev. 2, 15021 (2015).
Zhao, H. et al. Depletion of T cell epitopes in lysostaphin mitigates anti-drug antibody response and enhances antibacterial efficacy in vivo. Chem. Biol. 22, 629–639 (2015).
Liu, Y. et al. Immunomimetic designer cells protect mice from MRSA infection. Cell 174, 259–270 (2018).
Raz, A., Serrano, A., Thaker, M., Alston, T. & Fischetti, V. A. Lysostaphin lysibody leads to effective opsonization and killing of methicillin-resistant Staphylococcus aureus in a murine model. Antimicrob. Agents Chemother. 62, e01056-18 (2018).
Wall, R. J. et al. Genetically enhanced cows resist intramammary Staphylococcus aureus infection. Nat. Biotechnol. 23, 445–451 (2005).
Wittekind, M. & Schuch, R. Cell wall hydrolases and antibiotics: exploiting synergy to create efficacious new antimicrobial treatments. Curr. Opin. Microbiol. 33, 18–24 (2016).
Baba, T. & Schneewind, O. Target cell specificity of a bacteriocin molecule: a C-terminal signal directs lysostaphin to the cell wall of Staphylococcus aureus. EMBO J. 15, 4789–4797 (1996).
Lu, J. Z., Fujiwara, T., Komatsuzawa, H., Sugai, M. & Sakon, J. Cell wall-targeting domain of glycylglycine endopeptidase distinguishes among peptidoglycan cross-bridges. J. Biol. Chem. 281, 549–558 (2006).
Mitkowski, P. et al. Structural bases of peptidoglycan recognition by lysostaphin SH3b domain. Sci. Rep. 9, 5965 (2019).
Gründling, A. & Schneewind, O. Cross-linked peptidoglycan mediates lysostaphin binding to the cell wall envelope of Staphylococcus aureus. J. Bacteriol. 188, 2463–2472 (2006).
Tamai, E. et al. X-ray structure of a novel endolysin encoded by episomal phage phiSM101 of Clostridium perfringens. Mol. Microbiol 92, 326–337 (2014).
Shaner, N. C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Williamson, M. P How Proteins Work. (Garland Science, 2011). .
Zhou, H. X. Quantitative relation between intermolecular and intramolecular binding of pro-rich peptides to SH3 domains. Biophys. J. 91, 3170–3181 (2006).
Bolam, D. N. et al. Pseudomonas cellulose-binding domains mediate their effects by increasing enzyme substrate proximity. Biochem. J. 331, 775–781 (1998).
Gill, J. et al. The type II and X cellulose-binding domains of Pseudomonas xylanase A potentiate catalytic activity against complex substrates by a common mechanism. Biochem. J. 342, 473–480 (1999).
Nagy, T. et al. Characterization of a double dockerin from the cellulosome of the anaerobic fungus Piromyces equi. J. Mol. Biol. 373, 612–622 (2007).
Raghothama, S. et al. Characterization of a cellulosome dockerin domain from the anaerobic fungus Piromyces equi. Nat. Struct. Biol. 8, 775–778 (2001).
Nega, M. et al. Secretome analysis revealed adaptive and non-adaptive responses of the Staphylococcus carnosus femB mutant. Proteomics 15, 1268–1279 (2015).
Gally, D. & Archibald, A. R. Cell wall assembly in Staphylococcus aureus: proposed absence of secondary crosslinking reactions. J. Gen. Microbiol 139, 1907–1913 (1993).
Gu, J. et al. Structural and biochemical characterization reveals LysGH15 as an unprecedented “EF-hand-like” calcium-binding phage lysin. PLoS Pathog. 10, e1004109 (2014).
Meroueh, S. O. et al. Three-dimensional structure of the bacterial cell wall peptidoglycan. Proc. Natl Acad. Sci. USA 103, 4404–4409 (2006).
Tossavainen, H. et al. Structural and functional insights into lysostaphin–substrate interaction. Front. Mol. Biosci. 5, 60 (2018).
Francius, G., Domenech, O., Mingeot-Leclercq, M. P. & Dufrêne, Y. F. Direct observation of Staphylococcus aureus cell wall digestion by lysostaphin. J. Bacteriol. 190, 7904–7909 (2008).
Jagielska, E., Chojnacka, O. & Sabala, I. LytM fusion with SH3b-like domain expands its activity to physiological conditions. Micro. Drug Resist. 22, 461–469 (2016).
Sabala, I. et al. Crystal structure of the antimicrobial peptidase lysostaphin from Staphylococcus simulans. FEBS J. 281, 4112–4122 (2014).
Mesnage, S., Chau, F., Dubost, L. & Arthur, M. Role of N-acetylglucosaminidase and N-acetylmuramidase activities in Enterococcus faecalis peptidoglycan metabolism. J. Biol. Chem. 283, 19845–19853 (2008).
Mesnage, S. et al. Molecular basis for bacterial peptidoglycan recognition by LysM domains. Nat. Commun. 5, 4269 (2014).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl Crystallogr. 40, 658–674 (2007).
Zwart, P. H. et al. Automated structure solution with the PHENIX suite. Methods Mol. Biol. 426, 419–435 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Joosten, R. P., Joosten, K., Cohen, S. X., Vriend, G. & Perrakis, A. Automatic rebuilding and optimization of crystallographic structures in the Protein Data Bank. Bioinformatics 27, 3392–3398 (2011).
Horsburgh, M. J. et al. σB modulates virulence determinant expression and stress resistance: characterization of a functional rsbU strain derived from Staphylococcus aureus 8325-4. J. Bacteriol. 184, 5457–5467 (2002).
Acknowledgements
L.S.G.-D. is a PhD student funded by the Mexican government through a CONACYT scholarship. H.W.-M. is supported by a BBSRC MIBTP studentship. We thank the BBSRC and EPSRC for funding to upgrade the 600- and 800-MHz spectrometers, respectively (grant numbers BB/R000727/1 and EP/S01358X/1). The work in the laboratory of I.S. is supported by the Foundation for Polish Science (FNP) program, co-financed by the European Union under the European Regional Development Fund (grant TEAMTECH/2016-3/19).
Author information
Authors and Affiliations
Contributions
S.M. conceived the project and designed experiments with M.P.W. and A.L.L. A.M.H. and A.J.R. assigned the SH3b spectrum. L.S.G.-D. carried out all NMR experiments and analyzed them with the help of A.M.H. and M.P.W. B.S. and L.S.G.-D. built all SH3b recombinant proteins to carry out functional assays and crystallographic analyses. A.W.-M. crystallized the protein and solved the structure with the help of A.L.L. E.J. and I.S. provided reagents. L.S.G.-D., H.W.-M., B.S., A.M.H., M.P.W., A.L.L. and S.M. analyzed the data. L.S.G.-D., S.M., A.L.L. and M.P.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–5 and Supplementary Figures 1–12
Supplementary Note
Synthetic Procedures
Rights and permissions
About this article
Cite this article
Gonzalez-Delgado, L.S., Walters-Morgan, H., Salamaga, B. et al. Two-site recognition of Staphylococcus aureus peptidoglycan by lysostaphin SH3b. Nat Chem Biol 16, 24–30 (2020). https://doi.org/10.1038/s41589-019-0393-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0393-4
This article is cited by
-
LYZ2-SH3b as a novel and efficient enzybiotic against methicillin-resistant Staphylococcus aureus
BMC Microbiology (2023)
-
Staphylococcus aureus sacculus mediates activities of M23 hydrolases
Nature Communications (2023)
-
The measurement of binding affinities by NMR chemical shift perturbation
Journal of Biomolecular NMR (2022)
-
Influence of NaCl and pH on lysostaphin catalytic activity, cell binding, and bacteriolytic activity
Applied Microbiology and Biotechnology (2022)
-
Plate-associated localized osteitis in mini-pig by biofilm-forming Methicillin-resistant Staphylococcus aureus (MRSA): establishment of a novel experimental model
European Journal of Trauma and Emergency Surgery (2022)