Abstract
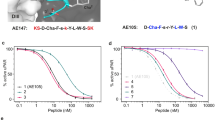
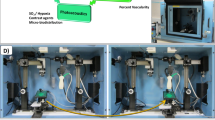
Activation of hepatocyte growth factor (HGF) by proteolytic processing is triggered in cancer microenvironments, and subsequent signaling through the MET receptor is involved in cancer progression. However, the structure of HGF remains elusive, and few small/medium-sized molecules can modulate HGF. Here, we identified HiP-8, a macrocyclic peptide consisting of 12 amino acids, which selectively recognizes active HGF. Biochemical analysis and real-time single-molecule imaging by high-speed atomic force microscopy demonstrated that HiP-8 restricted the dynamic domains of HGF into static closed conformations, resulting in allosteric inhibition. Positron emission tomography using HiP-8 as a radiotracer enabled noninvasive visualization and simultaneous inhibition of HGF–MET activation status in tumors in a mouse model. Our results illustrate the conformational change in proteolytic activation of HGF and its detection and inhibition by a macrocyclic peptide, which may be useful for diagnosis and treatment of cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary information or from the authors upon reasonable request.
Code availability
The authors declare that no custom code was used in this study. Software code or mathematical algorithm used in this study is available within the article and its Supplementary Information or from the authors upon reasonable request.
References
Naka, D. et al. Activation of hepatocyte growth factor by proteolytic conversion of a single chain form to a heterodimer. J. Biol. Chem. 141, 20114–20119 (1992).
Kataoka, H. et al. Activation of hepatocyte growth factor/scatter factor in colorectal carcinoma. Cancer Res. 60, 6148–6159 (2000).
Kawaguchi, M. & Kataoka, H. Mechanisms of hepatocyte growth factor activation in cancer tissues. Cancers 6, 1890–1904 (2014).
K Trusolino, L., Bertotti, A. & Comoglio, P. M. MET signalling: principles and functions in development, organ regeneration and cancer. Nat. Rev. Mol. Cell. Biol. 11, 834–848 (2010).
Gherardi, E., Birchmeier, W., Birchmeier, C. & Woude, G. V. Targeting MET in cancer: rationale and progress. Nat. Rev. Cancer 12, 89–103 (2012).
Sakai, K., Aoki, S. & Matsumoto, K. Hepatocyte growth factor and MET in drug discovery. J. Biochem. 157, 271–284 (2015).
Burggraaf, J. et al. Detection of colorectal polyps in humans using an intravenously administered fluorescent peptide targeted against c-MET. Nat. Med. 21, 955–961 (2015).
Han, Z. et al. Analysis of progress and challenges for various patterns of c-MET-targeted molecular imaging: a systematic review. EJNMMI Res. 7, 41 (2017).
Engelman, J. A. et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 16, 1039–1043 (2007).
Yano, S. et al. Hepatocyte growth factor induces gefitinib resistance of lung adenocarcinoma with epidermal growth factor receptor-activating mutations. Cancer Res. 68, 9479–9487 (2008).
Straussman, R. et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature 487, 500–504 (2012).
Corso, S. & Giordano, S. Cell-autonomous and non-cell-autonomous mechanisms of HGF/MET-driven resistance to targeted therapies: from basic research to a clinical perspective. Cancer Discov. 3, 978–992 (2013).
Cecchi, F., Rabe, D. C. & Bottaro, D. P. Targeting the HGF/MET signaling pathway in cancer therapy. Expert Opin. Ther. Targets 16, 553–572 (2012).
Furlan, A. et al. Thirty years of research on met receptor to move a biomarker from bench to bedside. Cancer Res. 74, 6737–6744 (2014).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Bendinelli, P., Maroni, P., Matteucci, E. & Desiderio, M. A. Epigenetic regulation of HGF/MET receptor axis is critical for the outgrowth of bone metastasis from breast carcinoma. Cell Death Dis. 8, e2578 (2017).
Matsumoto, K. et al. Hepatocyte growth factor/MET in cancer progression and biomarker discovery. Cancer Sci. 108, 296–307 (2017).
Grootjans, W. et al. PET in the management of locally advanced and metastatic NSCLC. Nat. Rev. Clin. Oncol. 12, 395–407 (2015).
Stamos, J. et al. Crystal structure of the HGF beta-chain in complex with the sema domain of the met receptor. EMBO J. 23, 2325–2335 (2004).
Kirchhofer, D. et al. Structural and functional basis of the serine protease-like hepatocyte growth factor beta-chain in met binding and signaling. J. Biol. Chem. 279, 39915–39924 (2004).
Kirchhofer, D. et al. Utilizing the activation mechanism of serine proteases to engineer hepatocyte growth factor into a met antagonist. Proc. Natl Acad. Sci. USA 104, 5306–5311 (2007).
Landgraf, K. E. et al. An allosteric switch for pro-HGF/MET signaling using zymogen activator peptides. Nat. Chem. Biol. 10, 567–573 (2014).
Gherardi, E. et al. Structural basis of hepatocyte growth factor/scatter factor and MET signalling. Proc. Natl Acad. Sci. USA 103, 4046–4051 (2006).
Winter, A. et al. Developing antagonists for the MET-HGF/SF protein–protein interaction using a fragment-based approach. Mol. Cancer Ther. 15, 3–14 (2016).
Tam, E. M. et al. Noncompetitive inhibition of hepatocyte growth factor-dependent MET signaling by a phage-derived peptide. J. Mol. Biol. 385, 79–90 (2009).
Valeur, E. et al. New modalities for challenging targets in drug discovery. Angew. Chem. Int. Ed. Engl. 56, 10294–10323 (2017).
Dougherty, P. G., Qian, Z. & Pei, D. Macrocycles as protein-protein interaction inhibitors. Biochem. J. 474, 1109–1125 (2017).
Villar, E. A. et al. How proteins bind macrocycles. Nat. Chem. Biol. 10, 723–731 (2014).
Millward, S. W., Fiacco, S., Austin, R. J. & Roberts, R. W. Design of cyclic peptides that bind protein surfaces with antibody-like affinity. ACS Chem. Biol. 2, 625–634 (2007).
Heinis, C., Rutherford, T., Freund, S. & Winter, G. Phage-encoded combinatorial chemical libraries based on bicyclic peptides. Nat. Chem. Biol. 5, 502–507 (2009).
Shi, Y., Yang, X., Garg, N. & Van Der Donk, W. A. Production of lantipeptides in Escherichia coli. J. Am. Chem. Soc. 133, 2338–2341 (2011).
Schlippe, Y. V., Hartman, M. C., Josephson, K. & Szostak, J. W. In vitro selection of highly modified cyclic peptides that act as tight binding inhibitors. J. Am. Chem. Soc. 134, 10469–10477 (2012).
Li, Y. et al. Versatile protein recognition by the encoded display of multiple chemical elements on a constant macrocyclic scaffold. Nat. Chem. 10, 441–448 (2018).
Kale, S. S. et al. Cyclization of peptides with two chemical bridges affords large scaffold diversities. Nat. Chem. 10, 715–723 (2018).
Passioura, T. & Suga, H. Flexizyme-mediated genetic reprogramming as a tool for noncanonical peptide synthesis and drug discovery. Chemistry. 19, 6530–6536 (2013).
Josephson, K., Ricardo, A. & Szostak, J. W. mRNA display: from basic principles to macrocycle drug discovery. Drug Discov. Today 19, 388–399 (2014).
Goto, Y., Katoh, T. & Suga, H. Flexizymes for genetic code reprogramming. Nat. Protoc. 6, 779–790 (2011).
Ito, K. et al. Artificial human met agonists based on macrocycle scaffolds. Nat. Commun. 6, 6372 (2015).
Veronese, F. M. & Pasut, G. PEGylation, successful approach to drug delivery. Drug Discov. Today 10, 1451–1458 (2005).
Holmes, O. et al. Insights into the structure/function of hepatocyte growth factor/scatter factor from studies with individual domains. J. Mol. Biol. 367, 395–408 (2007).
Lokker, N. A. et al. Structure-function analysis of hepatocyte growth-factor-identification of variants that lack mitogenic activity yet retain high-affinity receptor-binding. EMBO J. 11, 2503–2510 (1992).
Matsumoto, K., Kataoka, H., Date, K. & Nakamura, T. Cooperative interaction between α- and β-chains of hepatocyte growth factor on c-MET receptor confers ligand-induced receptor tyrosine phosphorylation and multiple biological responses. J. Biol. Chem. 273, 22913–22920 (1998).
Umitsu, M. et al. Probing conformational and functional states of human hepatocyte growth factor by a panel of monoclonal antibodies. Sci. Rep. 6, 33149 (2016).
Shibata, M. et al. High-speed atomic force microscopy shows dynamic molecular processes in photoactivated bacteriorhodopsin. Nat. Nanotech. 5, 208–212 (2010).
Uchihashi, T., Iino, R., Ando, T. & Noji, H. High-speed atomic force microscopy reveals rotary catalysis of rotorless F1-ATPase. Science 333, 755–758 (2011).
Ando, T., Uchihashi, T. & Scheuring, S. Filming biomolecular processes by high-speed atomic force microscopy. Chem. Rev. 114, 3120–3188 (2014).
Shibata, M. et al. Real-space and real-time dynamics of CRISPR-Cas9 visualized by high-speed atomic force microscopy. Nat. Commun. 8, 1430 (2017).
Chirgadze, D. Y. et al. Crystal structure of the NK1 fragment of HGF/SF suggests a novel mode for growth factor dimerization and receptor binding. Nat. Struct. Biol. 6, 72–79 (1999).
Yu, H. et al. Macrocycle peptides delineate locked-open inhibition mechanism for microorganism phosphoglycerate mutases. Nat. Commun. 8, 14932 (2017).
Wu, A. M. & Olafsen, T. Antibodies for molecular imaging of cancer. Cancer J. 14, 191–197 (2008).
Fukuta, K., Matsumoto, K. & Nakamura, T. Multiple biological responses are induced by glycosylation-deficient hepatocyte growth factor. Biochem. J. 388, 555–562 (2005).
Suzuki, Y. et al. Inhibition of MET/HGF receptor and angiogenesis by NK4 leads to suppression of tumor growth and migration in malignant pleural mesothelioma. Int. J. Cancer 127, 1948–1957 (2010).
Isozaki, H. et al. Non-small cell lung cancer cells acquire resistance to the ALK inhibitor alectinib by activating alternative receptor tyrosine kinases. Cancer Res. 76, 1506–1516 (2016).
Mukai, H., Wada, Y. & Watanabe, Y. The synthesis of 64Cu-chelated porphyrin photosensitizers and their tumor-targeting peptide conjugates for the evaluation of target cell uptake and PET image-based pharmacokinetics of targeted photodynamic therapy agents. Ann. Nucl. Med. 27, 625–639 (2013).
Zeng, D. et al. New cross-bridged cyclam derivative CB-TE1K1P, an improved bifunctional chelator for copper radionuclides. Chem. Commun. 50, 43–45 (2014).
Mukai, H. et al. Quantitative evaluation of the improvement in the pharmacokinetics of a nucleic acid drug delivery system by dynamic PET imaging with (18)F-incorporated oligodeoxynucleotides. J. Control. Release 180, 92–99 (2014).
Acknowledgements
This work was supported by World Premier International Research Center Initiative (WPI), MEXT, Japan. This work was supported in part by the A-STEP (Adaptable and Seamless Technology Transfer Program through Target-driven R&D) (grant no. AS262Z) from the Japan Science and Technology Agency (JST), the Medical Research Fund of Takeda Science Foundation, the Mitani Foundation for Research and Development, the Grant-in-Aid for JSPS Scientific Research (C) (no. 16K08544) to K.S., a Grant-in-Aid for JSPS Scientific Research (B) (no. 15K14473) to K.M., Project for Cancer Research and Therapeutic Evolution (P-CREATE) from the Japan Agency for Medical Research and development (AMED) to Y.W., H.M., K.M. and T.P., Basic Science and Platform Technology Program for Innovative Biological Medicine from AMED to H. Suga, a Grant-in-Aid for JSPS Research Activity Start-up (no. 16H06830) to H. Sato, a Grant-in-Aid for JSPS Fellows (no. 23-7727) to K.I., a Grant-in-Aid for JSPS Scientific Research (B) (no. 18K01836) to M.S. This work was performed under the Cooperative Research Program of the Institute for Protein Research, Osaka University (no. CR15-05) and an Extramural Collaborative Research Grant from the Cancer Research Institute (Kanazawa University). We thank T. Ando (Kanazawa University) for providing HS-AFM apparatus, T. Uchihashi (Nagoya University) for providing the analytical software of HS-AFM, Y. Kanayama and R. Zochi for their assistance in 64Cu production, Y. Wada and E. Hayashinaka for their assistance in reconstructing the PET images and Enago (www.enago.jp) for the English language review.
Author information
Authors and Affiliations
Contributions
K.S., H. Suga and K.M. conceived and designed the study. K.S. expressed and purified HGF and HGF fragment proteins. M.U. and J.T. expressed and purified Xa-modified scHGF, tcHGF, K2–4–SP and K4–SP proteins. K.I., K.S. and T.P. performed RaPID and peptide synthesis. K.S., H. Sato and K.I performed cell-based assays. K.S. and K.I. performed biochemical analysis. H. Sato performed immunohistochemistry and the in vivo efficacy studies. M.S. and H.F. performed HS-AFM observations. H.M., H. Sato, S.W., M.Z. and Y.W. designed and performed PET studies. Y.K. developed t5A11. S.Y. developed HGF-expressing PC-9. All authors analyzed the experimental data, discussed the results and were involved in preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–21
Supplementary Video 1 HS-AFM analysis of NK1.
NK1 consisting of two domains attached to the AP-mica surface. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 2 HS-AFM analysis of NK4.
NK4 attached to the AP-mica surface predominantly through NK1. NK4 had additional flexible domains corresponding to K2–K4, which did not interact with the AP-mica surface. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 3 HS-AFM analysis of SP.
SP sparingly bound to the AP-mica surface. Pixel sizes: 180 × 108 pixels. 200 × 120 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 4 HS-AFM analysis of scHGF.
scHGF attached to the AP-mica surface predominantly through NK1. scHGF had additional flexible domains corresponding to K2–SP, which did not interact with the AP-mica surface. The SP domain was bended toward N-terminus in scHGF. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 5 HS-AFM analysis of tcHGF.
tcHGF attached to the AP-mica surface predominantly through NK1. tcHGF had additional flexible domains corresponding to K2–SP, which did not interact with the AP-mica surface. The SP domain was more open toward N-terminus in tcHGF compared to scHGF. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 6 HS-AFM analysis of tcHGF/HiP-8 complex (Shape 1).
A representative tcHGF/HiP-8 complex on the AP-mica surface showing a static elongated shape (Shape 1). Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 7 HS-AFM analysis of tcHGF/HiP-8 complex (Shape 2).
A representative tcHGF/HiP-8 complexe on the AP-mica surface showing a static closed circular shape (Shape 2). Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated three times independently with similar results.
Supplementary Video 8 HS-AFM analysis of scHGF treated with HiP-8.
A representative scHGF treated with HiP-8 on the AP-mica surface showing an unchanged molecular shape and flexible conformation compared with free scHGF. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated twice independently with similar results.
Supplementary Video 9 HS-AFM analysis of tcHGF/t5A11 antibody complex.
Three representative tcHGF/t5A11 antibody complexes on the AP-mica surface. t5A11 bound to tcHGF between NK1 and SP domains. tcHGF molecules maintain their flexible conformation. Pixel sizes: 120 × 72 pixels. 80 × 47 nm2. This experiment was repeated twice independently with similar results.
Supplementary Video 10 Dynamic maximum intensity projection PET image of mice bearing PC-9 tumors intravenously administered with 64Cu-labeled HiP-8-PEG11.
This experiment was repeated three times independently with similar results.
Supplementary Video 11 Rotating maximum intensity projection image along with z axis of mice bearing PC-9 tumors intravenously administered with 64Cu-labeled HiP-8-PEG11 at 90 min.
This experiment was repeated three times independently with similar results.
Rights and permissions
About this article
Cite this article
Sakai, K., Passioura, T., Sato, H. et al. Macrocyclic peptide-based inhibition and imaging of hepatocyte growth factor. Nat Chem Biol 15, 598–606 (2019). https://doi.org/10.1038/s41589-019-0285-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0285-7
This article is cited by
-
Designing receptor agonists with enhanced pharmacokinetics by grafting macrocyclic peptides into fragment crystallizable regions
Nature Biomedical Engineering (2022)
-
Lasso-grafting of macrocyclic peptide pharmacophores yields multi-functional proteins
Nature Communications (2021)
-
Selection for constrained peptides that bind to a single target protein
Nature Communications (2021)
-
Quantitation of ligand is critical for ligand-dependent MET signalling activation and determines MET-targeted therapeutic response in gastric cancer
Gastric Cancer (2021)