Abstract
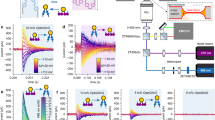
Sphingosine-1-phosphate (S1P) plays important roles as a signaling lipid in a variety of physiological and pathophysiological processes. S1P signals via a family of G-protein-coupled receptors (GPCRs) (S1P1–5) and intracellular targets. Here, we report on photoswitchable analogs of S1P and its precursor sphingosine, respectively termed PhotoS1P and PhotoSph. PhotoS1P enables optical control of S1P1–3, shown through electrophysiology and Ca2+ mobilization assays. We evaluated PhotoS1P in vivo, where it reversibly controlled S1P3-dependent pain hypersensitivity in mice. The hypersensitivity induced by PhotoS1P is comparable to that induced by S1P. PhotoS1P is uniquely suited for the study of S1P biology in cultured cells and in vivo because it exhibits prolonged metabolic stability compared to the rapidly metabolized S1P. Using lipid mass spectrometry analysis, we constructed a metabolic map of PhotoS1P and PhotoSph. The formation of these photoswitchable lipids was found to be light dependent, providing a novel approach to optically probe sphingolipid biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that all relevant data supporting the findings in this study are available within this paper and the Supplementary Information files.
References
Fyrst, H. & Saba, J. D. An update on sphingosine-1-phosphate and other sphingolipid mediators. Nat. Chem. Biol. 6, 489–497 (2010).
Hannun, Y. A. & Obeid, L. M. Principles of bioactive lipid signalling: lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 9, 139–150 (2008).
Maceyka, M., Harikumar, K. B., Milstien, S. & Spiegel, S. Sphingosine-1-phosphate signaling and its role in disease. Trends Cell Biol. 22, 50–60 (2012).
Proia, R. L. & Hla, T. Emerging biology of sphingosine-1-phosphate: its role in pathogenesis and therapy. J. Clin. Invest. 125, 1379–1387 (2015).
Hait, N. C. et al. Regulation of histone acetylation in the nucleus by sphingosine-1-phosphate. Science 325, 1254–1257 (2009).
Hill, R. Z. et al. The signaling lipid sphingosine-1-phosphate regulates mechanical pain. eLife 7, e33285 (2018).
Mair, N. et al. Genetic evidence for involvement of neuronally expressed s1p1 receptor in nociceptor sensitization and inflammatory pain. PLoS One 6, e17268 (2011).
Camprubí-Robles, M. et al. Sphingosine-1-phosphate-induced nociceptor excitation and ongoing pain behavior in mice and humans is largely mediated by S1P3 receptor. J. Neurosci. 33, 2582–2592 (2013).
Kim, R. H., Takabe, K., Milstien, S. & Spiegel, S. Export and functions of sphingosine-1-phosphate. Biochim. Biophys. Acta BBA 2009, 692–696 (1791).
Książek, M., Chacińska, M., Chabowski, A. & Baranowski, M. Sources, metabolism, and regulation of circulating sphingosine-1-phosphate. J. Lipid Res. 56, 1271–1281 (2015).
Brinkmann, V. et al. The immune modulator FTY720 targets sphingosine-1-phosphate receptors. J. Biol. Chem. 277, 21453–21457 (2002).
Chun, J. & Hartung, H.-P. Mechanism of action of oral fingolimod (FTY720) in multiple sclerosis. Clin. Neuropharmacol. 33, 91–101 (2010).
Wenk, M. R. Lipidomics: new tools and applications. Cell 143, 888–895 (2010).
Wenk, M. R. The emerging field of lipidomics. Nat. Rev. Drug Discov. 4, 594–610 (2005).
Schwarzmann, G., Arenz, C. & Sandhoff, K. Labeled chemical biology tools for investigating sphingolipid metabolism, trafficking and interaction with lipids and proteins. Biochim. Biophys. Acta BBA 2014, 1161–1173 (1841).
Haberkant, P. & Holthuis, J. C. M. Fat & fabulous: bifunctional lipids in the spotlight. Biochim. Biophys. Acta BBA 1841, 1022–1030 (2014).
Höglinger, D., Nadler, A. & Schultz, C. Caged lipids as tools for investigating cellular signaling. Biochim. Biophys. Acta BBA 1841, 1085–1096 (2014).
Feng, S. et al. Mitochondria-specific photoactivation to monitor local sphingosine metabolism and function. eLife 7, e34555 (2018).
Broichhagen, J., Frank, J. A. & Trauner, D. A roadmap to success in photopharmacology. Acc. Chem. Res. 48, 1947–1960 (2015).
Frank, J. A., Moroni, M., Moshourab, R., Sumser, M. & Lewin, G. R. Photoswitchable fatty acids enable optical control of TRPV1. Nat Commun. 6, 7118 (2015).
Leinders-Zufall, T. et al. PhoDAGs enable optical control of diacylglycerol-sensitive transient receptor potential channels. Cell Chem. Biol. 25, 215–223.e3 (2017).
Frank, J. A. et al. Optical control of GPR40 signalling in pancreatic β-cells. Chem. Sci. 8, 7604–7610 (2017).
Lichtenegger, M. et al. An optically controlled probe identifies lipid-gating fenestrations within the TRPC3 channel. Nat. Chem. Biol. 14, 396–404 (2018).
Frank, J. A., Franquelim, H. G., Schwille, P. & Trauner, D. Optical control of lipid rafts with photoswitchable ceramides. J. Am. Chem. Soc. 138, 12981–12986 (2016).
Pernpeintner, C. et al. Light-controlled membrane mechanics and shape transitions of photoswitchable lipid vesicles. Langmuir 33, 4083–4089 (2017).
Frank, J. A. et al. Photoswitchable diacylglycerols enable optical control of protein kinase C. Nat. Chem. Biol. 12, 755–762 (2016).
Hüll, K., Morstein, J. & Trauner, D. In vivo photopharmacology. Chem. Rev. 118, 10710–10747 (2018).
Bünemann, M. et al. A novel membrane receptor with high affinity for lysosphingomyelin and sphingosine-1-phosphate in atrial myocytes. EMBO J. 15, 5527–5534 (1996).
Troupiotis-Tsaïlaki, A. et al. Ligand chain length drives activation of lipid G protein-coupled receptors. Sci. Rep. 7, 2020 (2017).
Jeffery, T. Palladium-catalysed arylation of allylic alcohols: highly selective synthesis of β-aromatic carbonyl compounds or β-aromatic α,β-unsaturated alcohols. Tetrahedron Lett. 32, 2121–2124 (1991).
Yamamoto, T., Hasegawa, H., Hakogi, T. & Katsumura, S. Versatile synthetic method for sphingolipids and functionalized sphingosine derivatives via olefin cross metathesis. Org. Lett. 8, 5569–5572 (2006).
Lim, H.-S., Oh, Y.-S., Suh, P.-G. & Chung, S.-K. Syntheses of sphingosine-1-phosphate stereoisomers and analogues and their interaction with EDG receptors. Bioorg. Med. Chem. Lett. 13, 237–240 (2003).
Chan, K. W., Sui, J.-L., Vivaudou, M. & Logothetis, D. E. Control of channel activity through a unique amino acid residue of a G protein-gated inwardly rectifying K+ channel subunit. Proc. Natl Acad. Sci. USA 93, 14193–14198 (1996).
Atwood, B. K., Lopez, J., Wager-Miller, J., Mackie, K. & Straiker, A. Expression of G protein-coupled receptors and related proteins in HEK293, AtT20, BV2, and N18 cell lines as revealed by microarray analysis. BMC Genomics 12, 14 (2011).
Valentine, W. J. & Tigyi, G. in Sphingosine-1-Phosphate: Methods in Molecular Biology Vol. 874 (eds Pébay, A. & Turksen, K.) (Humana Press, 2012).
Rios Candelore, M. et al. Phytosphingosine-1-phosphate: a high affinity ligand for the S1P4/Edg-6 receptor. Biochem. Biophys. Res. Commun. 297, 600–606 (2002).
Basbaum, A. I., Bautista, D. M., Scherrer, G. & Julius, D. Cellular and molecular mechanisms of pain. Cell 139, 267–284 (2009).
Hill, R. Z., Morita, T., Brem, R. B. & Bautista, D. M. S1PR3 mediates itch and pain via distinct TRP channel-dependent pathways. J. Neurosci. 38, 7833–7843 (2018).
Harayama, T. & Riezman, H. Detection of genome-edited mutant clones by a simple competition-based PCR method. PLoS One 12, e0179165 (2017).
Harayama, T. & Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 19, 281–296 (2018).
Han, X., Yang, K. & Gross, R. W. Multi-dimensional mass spectrometry-based shotgun lipidomics and novel strategies for lipidomic analyses. Mass Spectrom. Rev. 31, 134–178 (2012).
Flock, T. et al. Selectivity determinants of GPCR–G-protein binding. Nature 545, 317–322 (2017).
Weth-Malsch, D. et al. Ablation of sphingosine 1-phosphate receptor subtype 3 impairs hippocampal neuron excitability in vitro and spatial working memory in vivo. Front. Cell. Neurosci. 10, 258 (2016).
Alvarez, S. E. et al. Sphingosine-1-phosphate is a missing cofactor for the E3 ubiquitin ligase TRAF2. Nature 465, 1084–1088 (2010).
Takasugi, N. et al. BACE1 activity is modulated by cell-associated sphingosine-1-phosphate. J. Neurosci. 31, 6850–6857 (2011).
Molecular Operating Environment (MOE) v.2013.08 (Chemical Computing Group ULC, 2018).
Berman, H. M. et al. The protein data bank. Acta Crystallogr. 58, 899–907 (2002).
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012).
Inagaki, Y. et al. Sphingosine-1-phosphate analogue recognition and selectivity at S1P4 within the endothelial differentiation gene family of receptors. Biochem. J. 389, 187–195 (2005).
Wang, D. A. et al. A single amino acid determines lysophospholipid specificity of the S1P1 (EDG1) and LPA1 (EDG2) phospholipid growth factor receptors. J. Biol. Chem. 276, 49213–49220 (2001).
Wilson, S. R. et al. TRPA1 Is required for histamine-independent, MAS-related G protein-coupled receptor-mediated itch. Nat. Neurosci. 14, 595–602 (2011).
Guan, X. L. et al. Functional interactions between sphingolipids and sterols in biological membranes regulating cell physiology. Mol. Biol. Cell 20, 2083–2095 (2009).
da Silveira dos Santos, A. X. et al. Systematic lipidomic analysis of yeast protein kinase and phosphatase mutants reveals novel insights into regulation of lipid homeostasis. Mol. Biol. Cell 25, 3234–3246 (2014).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Liao, S., Tammaro, M. & Yan, H. Enriching CRISPR-Cas9 targeted cells by co-targeting the HPRT gene. Nucleic Acids Res. 43, e134 (2015).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Acknowledgements
J.M. thanks the German Academic Scholarship Foundation (Studienstiftung) for a PhD Fellowship. J.M. and A.J.E.N. thank New York University for MacCracken PhD fellowships. T.H. was supported by the Japan Society for the Promotion of Science Postdoctoral Fellowships for Research Abroad. D.D.N. and G.Y.T. were supported by NCI grant No. CA092160. H.R. was supported by Sinergia, the Swiss National Science Foundation (No. CRSII3-154405) and the NCCR Chemical Biology funded by the Swiss National Science Foundation (No. 51NF40-160589). D.M.B was supported by NIH grants Nos. AR059385 and NS077224, and by an HHMI Faculty Scholar Award. E.Y.I. was supported by NIH grant 1U01MH109069-01. We thank B. Hetzler for critical discussion of the photophysical characterization and C. Lin for assistance with NMR experiments. S. Lee is acknowledged for performing TNA-α assays with PhotoS1P on S1P receptor subtypes (data not included).
Author information
Authors and Affiliations
Contributions
J.M. and D.T. designed and coordinated the study with critical input from all authors. A.J.E.N. performed chemical synthesis with input from B.M.W. P.C.D. and J.M. performed electrophysiology experiments under supervision by E.Y.I. and with input from J.A.F. D.D.N. and J.M. performed Ca2+ mobilization experiments under supervision by G.J.T. A.L.P. performed receptor homology modeling and docking experiments. R.Z.H. performed DRG neuronal Ca2+ imaging and in vivo pain physiology experiments under supervision by D.B. J.M. performed SphK assays. S.F. performed lipid mass spectrometry analysis. T.H. performed whole-lipidome analysis. T.H. generated CRISPR KO cell lines under supervision by H.R. J.M. and D.T. wrote the manuscript with critical input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Results
Supplementary Tables 1 and 2, Supplementary Figs. 1–12
Supplementary Note
Synthetic Procedures
Rights and permissions
About this article
Cite this article
Morstein, J., Hill, R.Z., Novak, A.J.E. et al. Optical control of sphingosine-1-phosphate formation and function. Nat Chem Biol 15, 623–631 (2019). https://doi.org/10.1038/s41589-019-0269-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0269-7
This article is cited by
-
A photoswitchable inhibitor of TREK channels controls pain in wild-type intact freely moving animals
Nature Communications (2023)
-
Optical control of neuronal activities with photoswitchable nanovesicles
Nano Research (2023)
-
Coenzyme Q10 in the eye isomerizes by sunlight irradiation
Scientific Reports (2022)
-
Advances and opportunities in the exciting world of azobenzenes
Nature Reviews Chemistry (2021)
-
Photoswitchable paclitaxel-based microtubule stabilisers allow optical control over the microtubule cytoskeleton
Nature Communications (2020)