Abstract
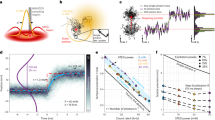
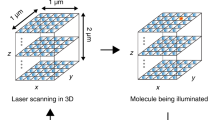
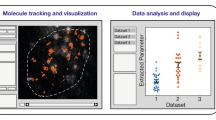
We describe three optical tags, ArrayG, ArrayD and ArrayG/N, for intracellular tracking of single molecules over milliseconds to hours. ArrayG is a fluorogenic tag composed of a green fluorescent protein–nanobody array and monomeric wild-type green fluorescent protein binders that are initially dim but brighten ~26-fold on binding with the array. By balancing the rates of binder production, photobleaching and stochastic binder exchange, we achieve temporally unlimited tracking of single molecules. High-speed tracking of ArrayG-tagged kinesins and integrins for thousands of frames reveals novel dynamical features. Tracking of single histones at 0.5 Hz for >1 hour with the import competent ArrayG/N tag shows that chromosomal loci behave as Rouse polymers with visco-elastic memory and exhibit a non-Gaussian displacement distribution. ArrayD, based on a dihydrofolate reductase nanobody array and dihydrofolate reductase–fluorophore binder, enables dual-color imaging. The arrays combine brightness, fluorogenicity, fluorescence replenishment and extended fluorophore choice, opening new avenues for tracking single molecules in living cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All datasets and analysis software are available upon request. All plasmid sequences have been submitted to NCBI GenBank and have been assigned the following accession numbers: MK317910, MK317911, MK317912, MK317913, MK317914, MK317915, MK317916, MK317917, MK317918, MK317919, MK317920. Plasmids are available upon request.
References
Peterman, E. J. G., Sosa, H. & Moerner, W. E. Single-molecule fluorescence spectroscopy and microscopy of biomolecular motors. Annu. Rev. Phys. Chem. 55, 79–96 (2004).
Xia, T., Li, N. & Fang, X. Single-molecule fluorescence imaging in living cells. Annu. Rev. Phys. Chem. 64, 459–480 (2013).
Cai, D., McEwen, D. P., Martens, J. R., Meyhofer, E. & Verhey, K. J. Single molecule imaging reveals differences in microtubule track selection between kinesin motors. PLoS Biol. 7, e1000216 (2009).
Buxbaum, A. R., Haimovich, G. & Singer, R. H. In the right place at the right time: visualizing and understanding mRNA localization. Nat. Rev. Mol. Cell Biol. 16, 95–109 (2015).
Bertrand, E. et al. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 2, 437–445 (1998).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Roukos, V., Burgess, R. C. & Misteli, T. Generation of cell-based systems to visualize chromosome damage and translocations in living cells. Nat. Protoc. 9, 2476–2492 (2014).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Liu, H. et al. Visualizing long-term single-molecule dynamics in vivo by stochastic protein labeling. Proc. Natl Acad. Sci. USA 115, 343–348 (2018).
Kamiyama, D. et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 7, 11046 (2016).
Cabantous, S., Terwilliger, T. C. & Waldo, G. S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat. Biotechnol. 23, 102–107 (2005).
Cabantous, S. & Waldo, G. S. In vivo and in vitro protein solubility assays using split GFP. Nat. Methods 3, 845–854 (2006).
Pinaud, F. & Dahan, M. Targeting and imaging single biomolecules in living cells by complementation-activated light microscopy with split-fluorescent proteins. Proc. Natl Acad. Sci. USA 108, E201–E210 (2011).
Kerppola, T. K. Bimolecular fluorescence complementation (BiFC) analysis as a probe of protein interactions in living cells. Annu. Rev. Biophys. 37, 465–487 (2008).
Li, C., Tebo, A. G. & Gautier, A. Fluorogenic labeling strategies for biological imaging. Int. J. Mol. Sci. 18, E1473 (2017).
Szent-Gyorgyi, C. et al. Fluorogen-activating single-chain antibodies for imaging cell surface proteins. Nat. Biotechnol. 26, 235–240 (2008).
De Meyer, T., Muyldermans, S. & Depicker, A. Nanobody-based products as research and diagnostic tools. Trends Biotechnol. 32, 263–270 (2014).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Kirchhofer, A. et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 17, 133–138 (2010).
Oyen, D., Wechselberger, R., Srinivasan, V., Steyaert, J. & Barlow, J. N. Mechanistic analysis of allosteric and non-allosteric effects arising from nanobody binding to two epitopes of the dihydrofolate reductase of Escherichia coli. Biochim. Biophys. Acta. 1834, 2147–2157 (2013).
Hocine, S., Raymond, P., Zenklusen, D., Chao, J. A. & Singer, R. H. Single-molecule analysis of gene expression using two-color RNA labeling in live yeast. Nat. Method 10, 119–121 (2013).
Viswanathan, S. et al. High-performance probes for light and electron microscopy. Nat. Method 12, 568–576 (2015).
Vale, R. D. The molecular motor toolbox for intracellular transport. Cell 112, 467–480 (2003).
Woehlke, G. et al. Microtubule interaction site of the kinesin motor. Cell 90, 207–216 (1997).
Zacharias, D. A., Violin, J. D., Newton, A. C. & Tsien, R. Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Phair, R. D. & Misteli, T. High mobility of proteins in the mammalian cell nucleus. Nature 404, 604–609 (2000).
Lowe, A. R. et al. Selectivity mechanism of the nuclear pore complex characterized by single cargo tracking. Nature 467, 600–603 (2010).
Shamir, M., Bar-On, Y., Phillips, R. & Milo, R. SnapShot: timescales in cell biology. Cell 164, 1302–1302.e1 (2016).
Shinkai, S., Nozaki, T., Maeshima, K. & Togashi, Y. Dynamic nucleosome movement provides structural information of topological chromatin domains in living human cells. PLoS Comput. Biol. 12, e1005136 (2016).
Hajjoul, H. et al. High-throughput chromatin motion tracking in living yeast reveals the flexibility of the fiber throughout the genome. Genome Res. 23, 1829–1838 (2013).
Weber, S. C., Spakowitz, A. J. & Theriot, J. A. Bacterial chromosomal loci move subdiffusively through a viscoelastic cytoplasm. Phys. Rev. Lett. 104, 238102 (2010).
Spichal, M. et al. Evidence for a dual role of actin in regulating chromosome organization and dynamics in yeast. J. Cell Sci. 129, 681–692 (2016).
Churchman, L. S., Flyvbjerg, H. & Spudich, J. A. A non-Gaussian distribution quantifies distances measured with fluorescence localization techniques. Biophys. J. 90, 668–671 (2006).
He, W. et al. Dynamic heterogeneity and non-Gaussian statistics for acetylcholine receptors on live cell membrane. Nat. Commun. 7, 11701 (2016).
Lampo, T. J., Stylianidou, S., Backlund, M. P., Wiggins, P. A. & Spakowitz, A. J. Cytoplasmic RNA-protein particles exhibit non-Gaussian subdiffusive behavior. Biophys. J. 112, 532–542 (2017).
Wang, B., Anthony, S. M., Bae, S. C. & Granick, S. Anomalous yet Brownian. Proc. Natl Acad. Sci. USA 106, 15160–15164 (2009).
Katrukha, E. A. et al. Probing cytoskeletal modulation of passive and active intracellular dynamics using nanobody-functionalized quantum dots. Nat. Commun. 8, 14772 (2017).
Perillo, E. P. et al. Deep and high-resolution three-dimensional tracking of single particles using nonlinear and multiplexed illumination. Nat. Commun. 6, 7874 (2015).
Ibach, J. et al. Single particle tracking reveals that EGFR signaling activity is amplified in clathrin-coated pits. PLoS One 10, e0143162 (2015).
Rossier, O. et al. Integrins β1 and β3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Cell Biol. 14, 1057–1067 (2012).
Schiller, H. B. et al. β1- and αv-class integrins cooperate to regulate myosin II during rigidity sensing of fibronectin-based microenvironments. Nat. Cell Biol. 15, 625–636 (2013).
Persson, F., Lindén, M., Unoson, C. & Elf, J. Extracting intracellular diffusive states and transition rates from single-molecule tracking data. Nat. Methods 10, 265–269 (2013).
Roca-Cusachs, P., Gauthier, N. C., del Rio, A. & Sheetz, M. P. Clustering of α5β1 integrins determines adhesion strength whereas αvβ3 and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Schvartzman, M. et al. Nanolithographic control of the spatial organization of cellular adhesion receptors at the single-molecule level. Nano Lett. 11, 1306–1312 (2011).
Li, J. et al. Conformational equilibria and intrinsic affinities define integrin activation. EMBO J. 36, 629–645 (2017).
Kong, F., García, A. J., Mould, A. P., Humphries, M. J. & Zhu, C. Demonstration of catch bonds between an integrin and its ligand. J. Cell Biol. 185, 1275–1284 (2009).
Grimm, J. B. et al. A general method to fine-tune fluorophores for live-cell and in vivo imaging. Nat. Methods 14, 987–994 (2017).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Liu, Z., Lavis, L. D. & Betzig, E. Imaging live-cell dynamics and structure at the single-molecule level. Mol. Cell 58, 644–659 (2015).
Cranfill, P. J. et al. Quantitative assessment of fluorescent proteins. Nat. Methods 13, 557–562 (2016).
McRae, S. R., Brown, C. L. & Bushell, G. R. Rapid purification of EGFP, EYFP, and ECFP with high yield and purity. Protein Expr. Purif. 41, 121–127 (2005).
Graham, J. S., Johnson, R. C. & Marko, J. F. Counting proteins bound to a single DNA molecule. Biochem. Biophys. Res. Commun. 415, 131–134 (2011).
Gosselin, P., Mohrbach, H., Kulić, I. M. & Ziebert, F. On complex, curved trajectories in microtubule gliding. Physica D. 318, 105–111 (2016).
Tarantino, N. et al. TNF and IL-1 exhibit distinct ubiquitin requirements for inducing NEMO-IKK supramolecular structures. J. Cell Biol. 204, 231–245 (2014).
Weber, S. C., Spakowitz, A. J. & Theriot, J. A. Nonthermal ATP-dependent fluctuations contribute to the in vivo motion of chromosomal loci. Proc. Natl Acad. Sci. USA 109, 7338–7343 (2012).
Acknowledgements
This work was partially supported by the National Institutes of Health, National Institute Of General Medical Sciences/National Cancer Institute (NCI) grant no. GM77856, NCI Physical Sciences Oncology Center grant no. U54CA143836, National Science Foundation Graduate Fellowship Program no. DGE-114747, National Institute Of Biomedical Imaging and Bioengineering/4D Nucleome Roadmap Initiative no. 1U01EB021237, National Institutes of Health Training Grant T32GM008294, National Science Foundation Graduate Fellowship Program no DGE-1656518 and National Science Foundation (NSF), Physics of Living Systems Program (PHY-1707751).
Author information
Authors and Affiliations
Contributions
R.P.G., J.M.F., W.D. and J.T.L. designed the research. R.P.G. and W.D. did most of the cloning. R.P.G generated and optimized most of the cell lines. R.P.G. and J.M.F carried out most of the imaging. W.D. set up the confocal calibration and did the HiLoTIRFM GFP counting. R.P.G and Q.S. carried out the nuclear sequestration experiments. Q.S performed the flow sorting experiments. J.M.F., W.D., Q.S. and R.P.G. analyzed most of the data. A.J.S. and B.B. analyzed the multiscale chromatin dynamics data. R.P.G., J.M.F. and J.T.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–13, Supplementary Note, Supplementary Tables 1–3
Supplementary Video 1
Laser-scanning confocal movie of dynamic droplet-like behavior of KIF560-ArrayG aggregates in presence of eGFP.
Supplementary Video 2
TIRF movie of integrin β1-ArrayG16x stably expressed in pKOαVβ1 cell line co-expressing mwtGFP.
Supplementary Video 3
20 Hz prolonged tracking of integrin β1-ArrayG16x + mwtGFP.
Supplementary Video 4
20 Hz HiLo-TIRF movie of H2B-ArrayGN + mwtGFP (left) and corresponding single-molecule trajectories (right).
Supplementary Video 5
HiLo-TIRF movie of H2B-ArrayGN + mwtGFP imaged under 20 Hz (left) and 0.5 Hz (right) showing gradual signal decay for 20 Hz case and constant signal strength for 0.5 Hz case (total illumination times in both cases are the same).
Supplementary Video 6
20 Hz HiLo-TIRFM movie of KIF560-ArrayG + mwtGFP (left) and corresponding single-molecule trajectories (right).
Supplementary Video 7
80 Hz HiLo-TIRFM movie of KIF560-ArrayG + mwtGFP on labeled tubulin.
Supplementary Video 8
20 Hz HiLo-TIRF movie of KIF560-ArrayD + DHFR-mGFP (left) and corresponding single-molecule trajectories (right).
Supplementary Video 9
20 Hz HiLo-TIRF movie of KIF560-ArrayD + DHFR-mCherry (left) and corresponding single-molecule trajectories (right).
Supplementary Video 10
20 Hz Dual-color TIRF imaging of cells co-expressing KIF560-ArrayG + mwtGFP (green) and KIF560-ArrayD + DHFR-mCherry (red) (left) with corresponding single molecule trajectories (right).
Supplementary Video 11
High temporal resolution (180 Hz) tracking of single KIF560-ArrayG molecules, using HiLo-TIRFM.
Supplementary Video 12
20 Hz TIRF imaging of vinculin-mCherry and β1-ArrayG16x + mwtGFP (left) with corresponding single-molecule trajectories (right).
Rights and permissions
About this article
Cite this article
Ghosh, R.P., Franklin, J.M., Draper, W.E. et al. A fluorogenic array for temporally unlimited single-molecule tracking. Nat Chem Biol 15, 401–409 (2019). https://doi.org/10.1038/s41589-019-0241-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0241-6
This article is cited by
-
Live-cell imaging of chromatin contacts opens a new window into chromatin dynamics
Epigenetics & Chromatin (2023)
-
Optical control of fast and processive engineered myosins in vitro and in living cells
Nature Chemical Biology (2021)
-
Nanobody-mediated control of gene expression and epigenetic memory
Nature Communications (2021)
-
Tracking single particles for hours via continuous DNA-mediated fluorophore exchange
Nature Communications (2021)
-
Advanced imaging and labelling methods to decipher brain cell organization and function
Nature Reviews Neuroscience (2021)