Abstract
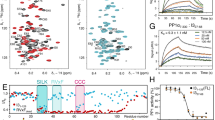
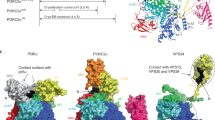
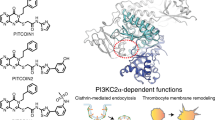
We have discovered a class of PI3Kγ inhibitors exhibiting over 1,000-fold selectivity over PI3Kα and PI3Kβ. On the basis of X-ray crystallography, hydrogen-deuterium exchange–mass spectrometry and surface plasmon resonance experiments we propose that the cyclopropylethyl moiety displaces the DFG motif of the enzyme away from the adenosine tri-phosphate binding site, inducing a large conformational change in both the kinase- and helical domains of PI3Kγ. Site directed mutagenesis explained how the conformational changes occur. Our results suggest that these cyclopropylethyl substituted compounds selectively inhibit the active state of PI3Kγ, which is unique to these compounds and to the PI3Kγ isoform, explaining their excellent potency and unmatched isoform selectivity that were confirmed in cellular systems. This is the first example of a Class I PI3K inhibitor achieving its selectivity by affecting the DFG motif in a manner that bears similarity to DFG in/out for type II protein kinase inhibitors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Crystal structures for mPI3Kδ in complex with AZ3 have been deposited in the Protein Data Bank under accession codes PDB 6GY0. All other data generated or analyzed during the study in this published article (and its supplementary information files) are available upon request.
References
Hirsch, E. et al. Central role for G protein-coupled phosphoinositide 3-kinase γ in inflammation. Science 287, 1049–1053 (2000).
Wymann, M. P., Zvelebil, M. & Laffargue, M. Phosphoinositide 3-kinase signalling—which way to target? Trends Pharmacol. Sci. 24, 366–376 (2003).
Rommel, C., Camps, M. & Ji, H.PI3Kδ and PI3Kγ: partners in crime in inflammation in rheumatoid arthritis and beyond? Nat. Rev. Immunol. 7, 191–201 (2007).
Camps, M. et al.Blockade of PI3Kγ suppresses joint inflammation and damage in mouse models of rheumatoid arthritis. Nat. Med. 11, 936–943 (2005).
Fougerat, A. et al. Genetic and pharmacological targeting of phosphoinositide 3-kinase-γ reduces atherosclerosis and favors plaque stability by modulating inflammatory processes. Circulation 117, 1310–1317 (2008).
Smirnova, N. F. et al. Targeting PI3Kγ activity decreases vascular trauma-induced intimal hyperplasia through modulation of the Th1 response. J. Exp. Med. 211, 1779–1792 (2014).
Gayral, S. et al. Elastin-derived peptides potentiate atherosclerosis through the immune Neu1-PI3Kγ pathway. Cardiovasc. Res. 102, 118–127 (2014).
Laffargue, M. et al. Phosphoinositide 3-kinase γ is an essential amplifier of mast cell function. Immunity 16, 441–451 (2002).
Bohnacker, T. et al. PI3Kγ adaptor subunits define coupling to degranulation and cell motility by distinct PtdIns(3,4,5)P3 pools in mast cells. Sci. Signal. 2, ra27 (2009).
Collmann, E. et al. Transient targeting of phosphoinositide 3-kinase acts as a roadblock in mast cells’ route to allergy. J. Allergy Clin. Immunol. 132, 959–968 (2013).
Patrucco, E. et al. PI3Kγ modulates the cardiac response to chronic pressure overload by distinct kinase-dependent and -independent effects. Cell 118, 375–387 (2004).
Vecchione, C. et al. Protection from angiotensin II-mediated vasculotoxic and hypertensive response in mice lacking PI3Kγ. J. Exp. Med. 201, 1217–1228 (2005).
Becattini, B. et al. PI3Kγ within a nonhematopoietic cell type negatively regulates diet-induced thermogenesis and promotes obesity and insulin resistance. Proc. Natl Acad. Sci. USA 108, E854–E863 (2011).
Wymann, M. P. & Solinas, G. Inhibition of phosphoinositide 3-kinase γ attenuates inflammation, obesity, and cardiovascular risk factors. Ann. NY Acad. Sci. 1280, 44–47 (2013).
Breasson, L. et al. PI3Kγ activity in leukocytes promotes adipose tissue inflammation and early-onset insulin resistance during obesity. Sci. Signal. 10, eaaf2969 (2017).
Kaneda, M. M. et al. PI3Kγ is a molecular switch that controls immune suppression. Nature 539, 437–442 (2016).
Lamb, D., Lunn, G., O’Reilly, M., Butler, C. & Kilty, I. In vitro and in vivo assessment of Pi3Kγ inhibitors for anti-inflammatory indications: challenges of selectivity over Pi3Kα. J. Pulm. Respir. Med. 3, 157 (2013).
Collier, P. N. et al. Structural basis for isoform selectivity in a class of benzothiazole inhibitors of phosphoinositide 3-kinase γ. J. Med. Chem. 58, 517–521 (2015).
Collier, P. N. et al. Discovery of highly isoform selective thiazolopiperidine inhibitors of phosphoinositide 3-kinase γ. J. Med. Chem. 58, 5684–5688 (2015).
Evans, C. A. et al. Discovery of a selective phosphoinositide-3-kinase (PI3K)-γ inhibitor (IPI-549) as an immuno-oncology clinical candidate. ACS Med. Chem. Lett. 7, 862–867 (2016).
Pemberton, N. et al. Discovery of highly isoform selective orally bioavailable phosphoinositide 3-kinase (PI3K)-γ inhibitors. J. Med. Chem. 61, 5435–5441 (2018).
Walker, E. H., Perisic, O., Ried, C., Stephens, L. & Williams, R. L. Structural insights into phosphoinositide 3-kinase catalysis and signalling. Nature 402, 313–320 (1999).
Carles, F., Bourg, S., Meyer, C. & Bonnet, P. PKIDB: a curated, annotated and updated database of protein kinase inhibitors in clinical trials. Molecules 23, 908 (2018).
Kufareva, I. & Abagyan, R. Type-II kinase inhibitor docking, screening, and profiling using modified structures of active kinase states. J. Med. Chem. 51, 7921–7932 (2008).
Fabbro, D. 25 years of small molecular weight kinase inhibitors: potentials and limitations. Mol. Pharmacol. 87, 766–775 (2015).
Geitmann, M. et al. Interaction kinetic and structural dynamic analysis of ligand binding to acetylcholine-binding protein. Biochemistry 49, 8143–8154 (2010).
Geitmann, M. & Danielson, U. H. Studies of substrate-induced conformational changes in human cytomegalovirus protease using optical biosensor technology. Anal. Biochem. 332, 203–214 (2004).
Flatmark, T., Stokka, A. J. & Berge, S. V. Use of surface plasmon resonance for real-time measurements of the global conformational transition in human phenylalanine hydroxylase in response to substrate binding and catalytic activation. Anal. Biochem. 294, 95–101 (2001).
Gestwicki, J. E., Hsieh, H. V. & Pitner, J. B. Using receptor conformational change to detect low molecular weight analytes by surface plasmon resonance. Anal. Chem. 73, 5732–5737 (2001).
Weis, D. D., Wales, T. E., Engen, J. R., Hotchko, M. & Ten Eyck, L. F. Identification and characterization of EX1 kinetics in H/D exchange mass spectrometry by peak width analysis. J. Am. Soc. Mass Spectrom. 17, 1498–1509 (2006).
Zhang, X. et al. Structure of lipid kinase p110β/p85β elucidates an unusual SH2-domain-mediated inhibitory mechanism. Mol. Cell 41, 567–578 (2011).
Vadas, O. et al. Molecular determinants of PI3Kγ-mediated activation downstream of G-protein-coupled receptors (GPCRs). Proc. Natl Acad. Sci. USA 110, 18862–18867 (2013).
Edgar, K. et al. Abstract 156: Preclinical characterization of GDC-0077, a specific PI3K alpha inhibitor in early clinical development. Cancer Res. 77, 156 (2017).
Walser, R. et al. PKCβ phosphorylates PI3Kγ to activate it and release it from GPCR control. PLoS Biol. 11, e1001587 (2013).
Suire, S. et al. p84, a new Gβγ-activated regulatory subunit of the type IB phosphoinositide 3-kinase p110γ. Curr. Biol. 15, 566–570 (2005).
Ali, K. et al. Essential role for the p110δ phosphoinositide 3-kinase in the allergic response. Nature 431, 1007–1011 (2004).
Bohnacker, T. et al. Deconvolution of Buparlisib’s mechanism of action defines specific PI3K and tubulin inhibitors for therapeutic intervention. Nat. Commun. 8, 14683 (2017).
Berndt, A. et al. The p110δ structure: mechanisms for selectivity and potency of new PI(3)K inhibitors. Nat. Chem. Biol. 6, 117–124 (2010).
Giordanetto, F. et al. Discovery of 9-(1-phenoxyethyl)-2-morpholino-4-oxo-pyrido[1,2-a]pyrimidine-7-carboxamides as oral PI3Kβ inhibitors, useful as antiplatelet agents. Bioorg. Med. Chem. Lett. 24, 3936–3943 (2014).
Beaufils, F. et al. 5-(4,6-Dimorpholino-1,3,5-triazin-2-yl)-4-(trifluoromethyl)pyridin-2-amine (PQR309), a potent, brain-penetrant, orally bioavailable, pan-class I PI3K/mTOR inhibitor as clinical candidate in oncology. J. Med. Chem. 60, 7524–7538 (2017).
Bauer, T. M., Patel, M. R. & Infante, J. R. Targeting PI3 kinase in cancer. Pharmacol. Ther. 146, 53–60 (2015).
Walker, E. H. et al. Structural determinants of phosphoinositide 3-kinase inhibition by wortmannin, LY294002, quercetin, myricetin, and staurosporine. Mol. Cell 6, 909–919 (2000).
Burke, J. E. et al. Dynamics of the phosphoinositide 3-kinase p110δ interaction with p85α and membranes reveals aspects of regulation distinct from p110α. Structure 19, 1127–1137 (2011).
Terstiege, I. et al. Discovery of triazole aminopyrazines as a highly potent and selective series of PI3Kδ inhibitors. Bioorg. Med. Chem. Lett. 27, 679–687 (2017).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D Biol. Crystallogr. 66, 133–144 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Klein, T. et al. Structural and dynamic insights into the energetics of activation loop rearrangement in FGFR1 kinase. Nat. Commun. 6, 7877 (2015).
Houde, D., Berkowitz, S. A. & Engen, J. R. The utility of hydrogen/deuterium exchange mass spectrometry in biopharmaceutical comparability studies. J. Pharm. Sci. 100, 2071–2086 (2011).
Acknowledgements
The authors would like to thank A. Gunnarsson, AstraZeneca, for input on membrane interactions, M. McAllister; AstraZeneca for facilitating the project and proofreading the manuscript and N. Bond, MedImmune, for initial HDX–MS experiments. G.G. is a fellow of the AstraZeneca post doc program. R.L.W. acknowledges support by the Medical Research Council (MC_U105184308) and an MRC—AstraZeneca/MedImmune Blue Sky Collaborative Research Grant (MC_A024-5PF9G). M.P.W. was supported by the Swiss Commission for Technology and Innovation; the Stiftung für Krebsbekämpfung Grant 341 and Swiss National Science Foundation Grants 310030_153211 and 316030_133860.
Author information
Authors and Affiliations
Contributions
G.G. and R.R. carried out HDX–MS experiments. G.G. and G.D. carried out SPR characterization of compounds. G.G. S.B. and J.G. carried out expression and purification of all protein reagents used in this study. K.K., C.T., N.P., M.M. and M.W.D.P. designed and synthesized compounds used in this study. J.P. and L.O. crystallized and determined the X-ray structure of mPI3Kδ-AZ3. G.G. and H.L. carried out kinetic characterization. T.B. and M.P.W. carried out studies in BMMCs. G.G., G.D., T.B., T.P., M.P.W., R.L.W. and J.P. contributed to data analysis and manuscript preparation. G.G., G.D. and J.P. designed and supervised the study, analyzed data and wrote the manuscript. R.L.W. contributed to writing the manuscript. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors, except for T.B., R.L.W. and M.P.W., are employees (and stockholders) of AstraZeneca UK Ltd or were at the time that this study was conducted.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5, Supplementary Figures 1–21
Supplementary Video 1
The movie based on our crystal structure and HDX-MS observations illustrates our model of how AZ2 binds to PI3Kγ and induces large-scale conformational changes that bring the PI3Kγ into its active conformation, but with the inhibitor bound. The movie was prepared with PyMOL (Schrödinger). In the first 1/3 of the move, AZ2 binds in the ATP-binding pocket. This initiates two conformational transitions that were created by the PyMOL Morph command. In the first transition forming the second 1/3 of the movie, the active site of the basal-state PI3Kγ (represented by the 4FUL PDB structure) 5 with the “Lock” intact morphs into a conformation with the lock broken and the helix kα12 lifted off the catalytic and activation loops, a conformation represented by our structure of p110δ bound to AZ2. We propose that this transition gives rise to the initial catalytically competent state of PI3Kγ. In the second transition, forming the last 1/3 of the movie, helix kα12 undergoes a large conformational change accompanied by widespread changes in conformation of the kinase domain, as the enzyme assumes the fully active conformation capable of carrying out catalysis on the membrane surface. We believe that the binding of this highly specific PI3Kγ inhibitor provides some insight into the conformational rearrangements that accompany the natural process of activation of the enzyme on membranes.
Supplementary Note 1
Synthetic Procedures
Rights and permissions
About this article
Cite this article
Gangadhara, G., Dahl, G., Bohnacker, T. et al. A class of highly selective inhibitors bind to an active state of PI3Kγ. Nat Chem Biol 15, 348–357 (2019). https://doi.org/10.1038/s41589-018-0215-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0215-0
This article is cited by
-
Development of PI3Kγ selective inhibitors: the strategies and application
Acta Pharmacologica Sinica (2024)
-
Molecular basis for Gβγ-mediated activation of phosphoinositide 3-kinase γ
Nature Structural & Molecular Biology (2024)
-
The role of PI3Kγ in the immune system: new insights and translational implications
Nature Reviews Immunology (2022)
-
QSAR analysis on a large and diverse set of potent phosphoinositide 3-kinase gamma (PI3Kγ) inhibitors using MLR and ANN methods
Scientific Reports (2022)
-
PI3K inhibitors are finally coming of age
Nature Reviews Drug Discovery (2021)