Abstract
Interferon-induced transmembrane proteins (IFITMs 1, 2 and 3) have emerged as important innate immune effectors that prevent diverse virus infections in vertebrates. However, the cellular mechanisms and live-cell imaging of these small membrane proteins have been challenging to evaluate during viral entry of mammalian cells. Using CRISPR–Cas9-mediated IFITM-mutant cell lines, we demonstrate that human IFITM1, IFITM2 and IFITM3 act cooperatively and function in a dose-dependent fashion in interferon-stimulated cells. Through site-specific fluorophore tagging and live-cell imaging studies, we show that IFITM3 is on endocytic vesicles that fuse with incoming virus particles and enhances the trafficking of this pathogenic cargo to lysosomes. IFITM3 trafficking is specific to restricted viruses, requires S-palmitoylation and is abrogated with loss-of-function mutants. The site-specific protein labeling and live-cell imaging approaches described here should facilitate the functional analysis of host factors involved in pathogen restriction as well as their mechanisms of regulation.
Similar content being viewed by others
Main
Interferon (IFN)-induced transmembrane proteins (IFITM1, IFITM2 and IFITM3) have emerged as important gatekeepers of pathogen infection in vertebrates1. Initial siRNA knockdown studies of IFITM3 increased the susceptibility of mammalian cell lines to influenza A virus (IAV), and overexpression showed that IFITM1, IFITM2 and IFITM3 could also inhibit infection by other strains of IAV as well as flaviviruses such as West Nile virus (WNV) and dengue virus (DENV), with IFITM3 being the most active against IAV2. In addition to IFITM1, IFITM2 and IFITM3, humans have two other IFITM isoforms (5 and 10) and mice have four others (5, 6, 7 and 10) that are not induced by IFNs1,2,3. Subsequent studies have demonstrated that IFITM1, IFITM2 and IFITM3 are active against many other viruses1,3, including Ebola virus (EBOV), vesicular stomatitis virus (VSV), SARS coronavirus (SARS-CoV), human immunodeficiency virus (HIV), respiratory syncytial virus (RSV), non-enveloped mammalian reovirus, hepatitis C virus (HCV) and Zika virus (ZIKV)4,5. IFITMs are, however, not active against DNA viruses such as papillomavirus, cytomegalovirus and adenovirus6 and even promote human coronavirus OC43 infection7,8. SiRNA knockdown of IFITM1, IFITM2 and IFITM3 together has also been reported to increase Mycobacterium tuberculosis (Mtb) infection in macrophages9. Moreover, IFITM3 was shown to associate with Mtb in late endosomal compartments and found to restrict Mtb by imaging flow cytometry9. IFITMs are active in multiple mammalian cell types, including primary CD8+ T cells and respiratory dendritic cells directly involved in adaptive immunity to IAV10. Notably, ifitm3−/− mice are more susceptible to infection by IAV11,12. Moreover, IFITM3 appears to be important for the intrinsic and inducible antiviral activity of human embryonic stem cells13. IFITMs have also been suggested to limit virus infection in a variety of other vertebrates1,3, highlighting their possible evolutionarily conserved function in host immunity.
IFITMs encode relatively small membrane proteins (107–144 amino acids)1 that were originally predicted to be dual-pass transmembrane proteins based on two hydrophobic domains2 (Fig. 1a). However, subsequent epitope mapping studies in cells14,15,16 and preliminary structural studies in vitro17 have suggested that IFITMs are type IV membrane proteins in which only the second hydrophobic domain is a transmembrane domain, with the C terminus facing the outside of cells. In contrast, the first hydrophobic domain of IFITM3 is suggested to function as an amphipathic helix and regulate interactions with the inner leaflet of host membranes18. This IFITM membrane topology model is also supported by the discovery of post-translational modifications on IFITM3 (S-fatty-acylation of Cys71, Cys72 and Cys105, phosphorylation of Tyr20, ubiquitination of Lys24, Lys83, Lys88 and Lys104 and methylation of Lys88)19. In mammalian cells, IFITM2 and IFITM3 are mostly distributed through early and late endocytic vesicles and on lysosomes, as shown by immunofluorescence1,3,19 and live-cell imaging studies20. IFITM1 has a truncated N terminus, lacks the AP2 sorting motif (YXXΦ) found in IFITM2 and IFITM3, and shows more plasma membrane localization1,3,19. The cellular localization of the IFITMs may be a crucial determinant for their specificities toward viruses, as IFITM1 is more active against viruses that enter through the plasma membrane or early endocytic vesicles, whereas IFITM2 and IFITM3 are more active against viruses that enter within late endocytic compartments1,3,19. In humans, individuals with the IFITM3 SNP rs12252-C allele, which encodes a predicted 21-amino acid truncation of the N terminus (IFITM3-N21Δ), show increased morbidity and mortality from H1N1 strains of IAV12. The IFITM3 rs12252-C allele is more prevalent in Asian populations21 and is associated with fatal IAV H7N9 infection22,23, as well as acute HIV-1 infection24. However, recent meta-analyses25,26,27, overexpression studies28 and direct transcript analyses29 have challenged the significance of this allele in humans. Further studies of IFITM3 SNPs have identified rs34481144 as another loss-of-function allele in the IFITM3 5′ UTR that results in lower mRNA expression, which correlates with reduced numbers of CD8+ T cells in patient airways and increased severity of influenza-mediated illness30. These studies highlight that IFITMs are highly regulated in mammalian cells and are important for host susceptibility to diverse pathogens.
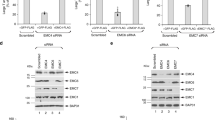
a, Alignment and topology of human IFITM1, IFITM2 and IFITM3. Key domains: amphipathic (AP) helix, hydrophobic (HP), and transmembrane (TM); mutations: N21Δ SNP rs12252-C allele and post-translational modifications are highlighted. b,c, Expression levels of human IFITM1, IFITM2 and IFITM3 in IFN-treated HeLa and A549 cell lines. Full gels for b are shown in Supplementary Fig. 17a. d, IAV infection of naïve and IFN-α stimulated HeLa WT and IFITM2/3-KO cell lines. Cells were stimulated with IFN-α (100 μg/mL) for 16 h and then infected with IAV (multiplicity of infection (MOI) = 2.5). At 12 h post-infection, cells were harvested, stained for IAV and analyzed by flow cytometry. The percentage of IAV-positive cells was normalized to WT cells and plotted as relative infection. Data represent the mean and s.d. of three independent experiments. e, Stable reconstitution with IFITM2 and IFITM3 in IFITM2/3-KO HeLa cells (C5-2). Low, medium and high IFITM2/3 protein expression levels are shown. f, IAV infection of IFITM2- and IFITM3-reconstituted HeLa and A549 cell clones stimulated with IFN-α (100 μg/mL). Experiments were performed and data were analyzed as described in d.
Though expression levels, subcellular localization and mutation of IFITM1, IFITM2 and IFITM3 clearly modulate host susceptibility toward different viruses, the precise cellular and biochemical mechanisms by which IFITMs inhibit virus entry in IFN-stimulated cells are still unclear1. Overexpression studies showed that IFITMs do not affect the expression levels of host ligands and receptors for viruses (for example, sialic acid levels)2 or uptake of IAV particles into mammalian cells31, suggesting that these small membrane proteins may interfere with intermediate or late stages of virus entry rather than initial binding or internalization into host cells. Notably, overexpression of IFITM3 appears to arrest internalized virus particles within endocytic vesicles and prevent the translocation of viral contents into the cytosol31. Overexpression studies have also suggested that IFITM3 expands Rab7- and LAMP1-positive late endocytic compartments31 and potentially alters the localization and activity of v-ATPase32 without affecting lysosomal cathepsin activity33. Alternatively, overexpression studies have also suggested that IFITMs can influence cell–cell fusion34 or indirectly alter intracellular cholesterol homeostasis35 to block viral hemifusion with host membranes. However, live-cell imaging of IAV entry and lipid mixing has shown that IFITMs do not affect virus hemifusion in endosomes36 or localization to endosomes37, indicating a possible role in blocking fusion pore formation. Interestingly, IFITMs may also be incorporated in HIV-1 virions and other viruses to inhibit virus spreading or be transferred through exosomes to propagate antiviral activity to neighboring cells38. The broad activity of IFITMs against different classes of viruses and perhaps even intracellular bacterial pathogens suggests a general mechanism of action that is functional in many different cell types. However, the lack of IFITM1, IFITM2 and IFITM3 mutant cell lines and tools for live imaging of IFITMs has limited their direct mechanistic investigation in host cells during infection, as well as dissection of their regulatory mechanisms.
To address the antiviral mechanism of IFITMs and their interactions with viruses in mammalian cells, we generated human IFITM1, IFITM2, and IFITM3 mutant cell lines by CRISPR–Cas9 gene editing and established site-specific fluorophore labeling of IFITM3 (ref. 20) for live-cell imaging during virus infection. We discovered that IFITM1, IFITM2 and IFITM3 act cooperatively in IFN-stimulated cells against diverse viruses (IAV, VSV, DENV, WNV and ZIKV) and enhance the trafficking of both endogenous (activated EGFR) and exogenous (virus particles) cargo in late endocytic vesicles to lysosomes for degradation. Using live-cell IFITM3 imaging experiments, we observed that internalized IAV particles primarily undergo lipid mixing or hemifusion with host membranes after fusion of virus-bearing vesicles with IFITM3-positive vesicles. The prefusion colocalization of virus particles with IFITM3-positive vesicles was specific to restricted viruses (IAV and EBOV) and was markedly decreased with loss-of-function IFITM3 mutants such as the N21Δ SNP rs12252-C allele. We also demonstrated that site-specific S-fatty-acylation of Cys72 is important for IFITM3 trafficking to incoming IAV particles. Collectively, our results showed that IFITM3 can engage vesicles bearing incoming virus particles and promote shuttling of late endocytic cargo to lysosomes for degradation, which highlights a general mechanism by which IFITMs may intercept pathogens and prevent their entry into host cells. These studies were uniquely enabled by site-specific protein labeling and imaging methods, which should also facilitate the functional analysis of other host factors involved in pathogen restriction as well as their mechanisms of regulation.
Results
IFITM1, IFITM2 and IFITM3 act cooperatively in IFN-stimulated cells
To evaluate antiviral activity of endogenously expressed human IFITMs, we employed CRISPR–Cas9 gene editing to generate human IFITM2 and IFITM3 knockout (KO) cell lines for HeLa (cervical epithelial) cells and A549 (lung epithelial) cells, which normally express high levels of human IFITM1, IFITM2 and IFITM3 following IFN stimulation (Fig. 1a). Among the gRNAs we evaluated targeting exon 1 of human IFITM3 (Supplementary Fig. 1b), gRNA3 afforded the most efficient cleavage of genomic IFITM3 (Supplementary Fig. 1c). Western blot analysis of CRISPR–Cas9-gRNA3-transfected stable cell clones after puromycin selection yielded several HeLa and A549 clones with decreased and/or deleted IFITM2 and IFITM3 using an N-terminally-directed antibody that cross-reacts with both proteins (Supplementary Fig. 1d). Because human IFITM2 and IFITM3 have high sequence identity in their first exons and also flank IFITM1 on chromosome 11, we evaluated the expression of IFITM1, IFITM2 and IFITM3 using isoform-specific antibodies (Fig. 1b). From this analysis, we identified HeLa cell clones with reduced expression in IFITM2 (C4–4), IFITM3 (C345–5), or both (C5-2, IFITM2/3-KO), as well as one clone that showed loss of IFITM1, IFITM2 and IFITM3 expression (C5-9) compared to Cas9 empty-vector-treated HeLa cells (WT HeLa) (Fig. 1b). For A549 cells, we only isolated IFITM2/IFITM3 double KO clones with partially reduced expression of IFITM1 (C1–3) (Fig. 1c). A summary of the IFITM2 and IFITM3 mutations in these HeLa and A549 mutant cell lines is included in Supplmentary Table 1.
We next evaluated the virus susceptibility of these HeLa and A549 IFITM (1, 2 and 3) mutant cell lines with and without IFN-α treatment. Infection of the HeLa cell clones with IAV (A/PR/8/34 (H1N1) strain) showed that reduced expression of IFITM2 or IFITM3 increased virus susceptibility approximately two- or four-fold, respectively, compared to controls (Fig. 1d; Supplementary Fig. 1e). However, HeLa cell clones with complete loss of IFITM2 and IFITM3 (C5-2, IFITM2/3-KO) or all three isoforms (C5-9) exhibited greater than 20-fold increased infection, as judged by anti-influenza A virus nucleoprotein (NP) staining (Fig. 1d; Supplementary Fig. 1e). The cooperative antiviral activity among IFITMs was even more pronounced in IFN-α treated cells, in which reduced expression of IFITM2 or IFITM3 alone did not have a major effect on IAV infection, whereas deletion of both or all three isoforms again resulted in greater than 20-fold increased susceptibility (Fig. 1d; Supplementary Fig. 1e). No significant difference in IAV infection of clones with KO of IFITM2/3 (C5-2,) or all three isoforms (C5-9) was observed, even with IFN-α treatment (Fig. 1d; Supplementary Fig. 1e), suggesting that IFITM1 does not play a significant role in preventing IAV infection. In contrast, the analysis of IFITM mutant cell lines with other viruses showed that VSV and Venezuelan equine encephalitis virus (VEEV) infected naïve HeLa cell clones at similar levels, but with IFN-α treatment, reduced expression or deletion of IFITM1, IFITM2 or IFITM3 increased infection, particularly when all three isoforms were deleted (Supplementary Fig. 1f). DENV also showed increased infectivity in the naïve IFITM-mutant HeLa cell lines (Supplementary Fig. 1f). However, loss of IFITM2 or IFITM3 expression alone in IFN-α treated cells only partially attenuated DENV infection compared to HeLa cells without IFITM1, IFITM2 and IFITM3 (Supplementary Fig. 1f). YFV and o’nyong’nyong virus (ONNV) showed enhanced infection of naïve IFITM-mutant cells but were attenuated with IFN-α treatment (Supplementary Fig. 1f), suggesting that other IFN-effectors also contribute to host restriction of these viruses. We observed similar trends with virus infections of naïve and IFN-α treated IFITM2/3-mutant A549 cells compared to wild-type (WT) controls (Supplementary Fig. 1g). To confirm that the virus susceptibility was indeed due to reduced expression or deletion of IFITM2 or IFITM3, we generated stable clones reconstituted with IFITM2 or IFITM3. In the HeLa IFITM2/3-KO clone (C5-2), we generated cells expressing low, medium or near-WT levels of IFITM3 and a clone expressing only IFITM2 (Fig. 1e). IAV infection of these HeLa cell lines showed dose-dependent antiviral activity by IFITM3 and indicated that IFITM3 or IFITM2 expression alone was sufficient to render these IFITM2/3-KO HeLa and A549 cells resistant to virus infection (Fig. 1f). These results show that human IFITM3 functions in a dose-dependent manner and may be redundant with IFITM2 toward some viruses, but it may act synergistically with IFITM1 to restrict a broad spectrum of viruses in IFN-stimulated cells. At endogenous levels in IFN-stimulated cells, IFITM2 and IFITM3 are most important for IAV and VEEV, whereas all three IFITMs are involved in preventing VSV and DENV infections.
IFITMs enhance trafficking of late endocytic cargo to lysosomes
To elucidate cellular mechanisms by which IFITMs inhibit virus infection, we evaluated the localization of endogenous IFITM3, endocytic vesicle distribution and cargo trafficking in IFN-α-treated cells. Immunofluorescence analysis showed that endogenous as well as reconstituted IFITM3 is present on intracellular vesicular structures that partially colocalize with CD63-positive late endocytic compartments in IFN-α treated A549 cells (Fig. 2a). Fluorescence imaging of endocytic markers (GFP-Rab5a, GFP-Rab7a and LAMP1-mCherry) did not show significant differences in the distribution of early endosomes, late endosomes and lysosomes between A549 WT and IFITM2/3-KO cells (Supplementary Fig. 2a,b). We also did not observe significant enlargement or expansion of LAMP1-mCherry compartments in IFN-α-treated control cells compared to IFITM2/3-KO cells (Supplementary Fig. 2a,b), in contrast to results of previous IFITM3 overexpression studies31. Moreover, the uptake of exogenous cargo (dextran particles) was similar between IFN-α-treated WT and IFITM2/3-KO cells (Supplementary Fig. 2c,d). IFN-α-treated WT and IFITM2/3-KO cells also exhibited similar labeling with fluorogenic cell-penetrating peptides that enter through the endocytic pathway39 (Supplementary Fig. 3). These results collectively suggest that IFITM2 and IFITM3 do not significantly regulate early endocytic events or generally alter endocytic vesicle membrane permeability.
a, Localization of IFITM3 and CD63 in IFN-α stimulated A549 cells. Pearson correlation coefficient (PCC) analyses are included below. Top row, endogenous IFITM3 and CD63 in IFN-α stimulated WT A549 cells (PCC = 0.58); middle row, endogenous IFITM3 and CD63 in IFN-α stimulated IFITM2/3-KO cells (PCC = n/a); bottom row, IFITM2/3-KO cells + stable re-expression of IFITM3 (PCC = 0.46). b, EGFR levels in IFN-α stimulated A549 WT, IFITM2/3-KO and IFITM2/3-KO cells + stable re-expression of IFITM3. Cells were treated with IFN-α (100 μg/ml) for 16 h, serum-starved for 2 h, treated with 25 μg/ml cycloheximide for 1 h and then treated with 100 ng/ml EGF for 30 min at 4 °C in the continued presence of cycloheximide. Cells were subsequently shifted to 37 °C and analyzed at various time points. Full gels are shown in Supplementary Fig. 17b. c, Quantitation of EGFR levels in b normalized to tubulin levels. Data represent the mean and s.d. of three independent experiments.
As early endocytic events are not affected and endogenous IFITM3 is mainly localized to late endosomes or lysosomes, we evaluated whether IFITM2/3 may affect late endocytic vesicle cargo trafficking and degradation. Our initial pulse-chase labeling studies with fluorescent dextran particles suggested that exogenous cargo trafficking to lysosomes was modestly delayed in IFN-α treated IFITM2/3-KO cells (Supplementary Fig. 2c,d). To evaluate the trafficking and degradation of endogenous endocytic cargo, we focused on the turnover of activated epidermal growth factor receptor (EGFR), which is one of the major sialylated cell surface receptors for IAV40 and is readily detectable in A549 cells. While the cell surface levels of EGFR were similar in WT and IFITM2/3-KO A549 cells (Supplementary Fig. 4a), following EGF-stimulation of IFN-α-treated A549 cells, EGFR was largely degraded after 90 min (Fig. 2b,c). Notably, EGFR turnover was markedly delayed in IFN-α-treated IFITM2/3-KO cells, with significant levels still present after 90 min (Fig. 2b,c). Stable reconstitution of IFITM3 in IFN-α-treated IFITM2/3-KO cells rescued and even accelerated the degradation of EGFR (Fig. 2b,c). By contrast, treatment of these cells with transferrin (Tf) in the same time frame (Supplementary Fig. 4b,c) suggests IFITM2 and IFITM3 do not significantly affect the turnover of activated transferrin receptor (TfR, CD71), a GPI-anchor-modified membrane protein that is recycled to the plasma membrane. Analysis of lysosomal protease activity using a fluorogenic cathepsin B substrate and cysteine protease activity-based profiling showed similar levels of endolysosomal proteolytic activity in IFN-α-treated A549 and HeLa cells regardless of IFITM2/3 expression levels (Supplementary Fig. 4d,e). These data, together with pulse-chase analysis of fluorescent-dextran particles, suggest that IFITM2 and IFITM3 do not significantly alter endolysosomal protease activity but may enhance the trafficking of late endocytic cargo to lysosomes for degradation, even in the absence of virus infection.
IFITM2/3 increase the rate of virus trafficking to lysosomes
To investigate the effects of IFITMs on viral entry and trafficking, we employed live-cell fluorescence microscopy to monitor individual IAV particles during synchronized infection. Using a membrane-incorporated dye, 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindodicarbocyanine (DiD), which readily labels IAV particles in vitro (Supplementary Fig. 5) and exhibits fluorescence dequenching upon hemifusion or full fusion between viral and endosomal membranes41, we were able to track viral membrane fusion, as indicated by glycoprotein-mediated lipid mixing, and monitor the trafficking of virus-fused vesicles. Because IAV is known to fuse with late endosomes, we first evaluated IAV lipid mixing and trafficking in WT and IFITM2/3-KO cells transduced to express LAMP1-GFP. These experiments were performed in HeLa cells, which exhibit less contour than A549 cells and enabled more comprehensive quantitative imaging of DiD-IAV particles by wide-field epifluorescence microscopy. IAV lipid mixing occurred almost exclusively in LAMP1-positive compartments (Fig. 3a,b; Supplementary Video 1). The rate at which IAV particles underwent lipid mixing was similar in WT and IFITM2/3-KO cells (Fig. 3c, black and white circles), with a t1/2 of approximately 41 min for both. In IFN-α-treated cells, the DiD-IAV dequenching rate was delayed in IFITM2/3-KO HeLa cells compared to WT cells (Fig. 3c, black and white squares). The t1/2 for lipid mixing in IFN-treated IFITM2/3-KO HeLa cells occurred 11 min later than that in WT cells. Moreover, the trafficking of DiD-IAV particles to LAMP1-positive compartments in IFN-α-treated WT HeLa cells was also ~10 min faster, on average, even though virus trafficking times may vary widely even in synchronized infections (Fig. 3d). No significant difference in trafficking of DiD-IAV particles to LAMP1-positive compartments was observed in unstimulated cells (Fig. 3d). Although IFN-α treatment reduced the relative amount of dequenched DiD-IAV particles, the percentage of dequenched DiD-IAV particles (Fig. 3e) and the degree of colocalization with LAMP1-positive compartments were similar between HeLa WT and IFITM2/3-KO cells at 2 h post-infection (Fig. 3f). These results show that IFN-α treatment impacts the kinetics, but not total extent, of viral lipid mixing in IFITM2/3-KO cells compared to WT cells, which suggests that IFITM2 and IFITM3 primarily influence the rate of virus trafficking to late endosomal compartments rather than the gross composition of the endosomal milieu, consistent with our data on dextran particle trafficking (Supplementary Fig. 2c,d) EGFR (Fig. 2b,c) and TfR turnover (Supplementary Fig. 4b,c).
a, Representative images of lipid mixing by an IAV particle in a LAMP1 + compartment in a HeLa cell as depicted by DID dequenching. The arrowheads indicate the virion of interest. The number show the time post-infection (minutes). Scale bar, 5 μm. b, Fluorescence intensity trace for the indicated particle in a. a.u., arbitrary units. c, IAV lipid mixing events following synchronized infection in HeLa WT or IFITM2/3-KO cells in the presence or absence of IFN-α (n > 100 dequenching particles tracked in three independent experiments). d, Times of IAV delivery to LAMP1 + compartments following synchronized infection in unstimulated and IFN-α stimulated HeLa WT or IFITM2/3-KO cells. For each sample, n > 100 particles tracked in three independent experiments. Mean and s.d. values are indicated. P < 0.0001 by and unpaired, two-sided t-test. e, Total percentage of cell-associated IAV particles in HeLa WT or IFITM2/3-KO cells that undergo lipid mixing by 2 h post-infection. Data represent the mean and s.d. of three independent experiments (n = 800–1,000 cell-associated particles total for each sample). P values were determined by one-way ANOVA with a post-hoc Tukey test. f, Total percentage of cell-associated particles colocalized with LAMP1 at 2 h post-infection. Data represent the mean and s.d. of three independent experiments (n = 800–1,000 cell-associated particles total for each sample). P value is nonsignificant (ns) by unpaired, two-sided t-test.
IFITM3 vesicles directly fuse with restricted virus particles
The trafficking of IFITMs in live cells during virus infection has not been previously described. Immunofluorescence and electron microscopy studies have suggested that IFITM3 is localized around IAV and VSV16,33,35,42 as well as Mtb-containing vesicles9. Indeed, our immunofluorescence studies also showed that endogenous IFITM3 in IFN-α-treated A549 cells partially colocalized with internalized IAV particles visualized by anti-IAV NP antibody staining (Supplementary Fig. 6a) and perhaps even surrounded virus particles when imaged with higher resolution structured illumination microscopy (Supplementary Fig. 6b). However, these studies do not reveal whether IFITMs interact with incoming virus particles or affect their trafficking within endocytic compartments.
To address this key aspect of IFITM function during virus entry, we recently optimized site-specific fluorophore labeling of proteins using amber-codon-suppression-mediated unnatural amino acid mutagenesis to evaluate IFITM3 trafficking in live mammalian cells20. Briefly, the co-transfection of mammalian cells with plasmids encoding with HA-IFITM3-F8-TAG and Mm-PylRS-AF/Pyl-tRNACUA enables site-specific incorporation of trans-cyclooct-2-ene-lysine (TCOK) and subsequent bioorthogonal labeling with BODIPY-tetrazine (BODIPY-Tz) (Supplementary Fig. 7). Importantly, HA-IFITM3-F8-TCOK labeled with BODIPY-Tz retains antiviral activity toward IAV (Supplementary Fig. 8a). The analysis of IFITM3 expression levels by α-IFITM3 antibody staining showed that HA-IFITM3 site-specifically labeled with TCOK in IFITM2/3-KO cells yielded slightly lower levels of total protein compared to WT HeLa cells ± IFN-α, as shown by western blot and flow cytometry analysis (Supplementary Fig. 8b,c), and a similar pattern of vesicle staining as indicated by immunofluorescence (Supplementary Fig. 8d). These results demonstrate that TCOK-labeled IFITM3 expression levels from transient transfection are within the physiological range of endogenous IFITM3 protein levels and exhibit comparable subcellular localization and antiviral activity.
We then performed these imaging studies in IFITM2/3-KO cells, which enabled us to monitor all IFITM3-positive compartments and avoid potential heterotypic interactions with endogenous IFITMs. Live imaging of IFITM2/3-KO cells revealed that relatively little BODIPY-IFITM3 colocalized with Rab5 at steady state (Manders correlation coefficient (MCC) 0.45 ± 0.13; Supplementary Fig. 9). Instead, we observed a significantly greater overlap of BODIPY-IFITM3 with Rab7 (MCC 0.98 ± 0.06) and LAMP1 (MCC 0.97 ± 0.03) (Supplementary Fig. 9), which shows that IFITM3 largely resides in late endocytic vesicles and is consistent with previous immunofluorescence9,16,33,35 and live-cell imaging studies of IFITM3 in WT HeLa cells20. Using DiD-labeled IAV and epifluorescence imaging, we observed BODIPY-IFITM3 colocalization with the majority of internalized virus particles before lipid mixing (Fig. 4a,b; Supplementary Video 2), with another 20 percent of DiD-dequenching events occurring in compartments without detectable BODIPY-IFITM3 (Fig. 4c). In these latter events, BODIPY-IFITM3 vesicles were observed to colocalize with DiD-dequenched puncta later in time, where secondary DiD dequenching events could occasionally be observed (Fig. 4d; Supplementary Video 3). This secondary DiD dequenching likely represents further dye diffusion due to endosome fusion with an IFITM3-positive vesicle and expansion of the limiting membrane. Reintroduction of IFITM3 into HeLa IFITM2/3-KO cells did not significantly alter the total percentage of IAV particles undergoing lipid mixing by 2 h post-infection (Supplementary Fig. 10a). Consistent with our previous experiments (Fig. 3), these results show that although virus particles may fuse with IFITM3-positive vesicles, IFITM3 does not significantly inhibit IAV lipid mixing or hemifusion with endocytic membranes. More importantly, IFITM3 colocalized with nearly 80 percent of DiD-IAV particles at the time of lipid mixing (Fig. 4e), but this was markedly decreased with VSV particles pseudotyped with the glycoprotein complex (GPC) of Lassa virus (LASV), a late-penetrating arenavirus that is not restricted by IFITMs2 (Fig. 4e; Supplementary Video 4). In these experiments, both IAV and LASV GPC-pseudotyped VSV still trafficked to late endocytic compartments, as indicated by colocalization with LAMP1 (Fig. 4e), and the extent of colocalization of both viruses with IFITM3 at 1 h post-infection was similar (Supplementary Fig. 10b). However, most LASV GPC-bearing particles dequenched before their endosomes acquired IFITM3 (Fig. 4f), whereas IAV dequenching generally occurred shortly after colocalization with IFITM3 (Fig. 4f). To further investigate the specificity of IFITM3 trafficking toward restricted viruses, we analyzed VSV particles pseudotyped with the glycoprotein of Ebola virus (EBOV), a late endosome/lysosome-entering virus that is restricted by IFITM3 (refs. 33,42). Like IAV, the majority of DiD-EBOV particles colocalized with IFITM3 prior to lipid mixing (Fig. 4e,f). These results demonstrate for the first time that incoming virus particles interact with IFITM3-positive vesicles before fusion and that this phenomenon is specific to restricted viruses such as IAV and EBOV.
a, IAV lipid mixing in an IFITM3 + compartment over time. Bar, 5 μm. b, The fluorescence intensity trace for the indicated particle in a. c, Example of IAV lipid mixing before colocalization with IFITM3 in HeLa cells. Scale bar, 5 μm. For a and c, the arrows indicate the virion of interest in each panel. The numbers indicate the time post-infection (minutes). d, The fluorescence intensity trace for the indicated particle in c. e, Percentage of particles of IAV, and VSV pseudotyped with LASV or EBOV glycoproteins undergoing lipid mixing in LAMP1 + or IFITM3 + compartments in HeLa cells. Data represent the mean and s.d. of three independent experiments (n = 800–1,000 cell-associated particles total for each sample). P values were determined by one-way ANOVA with a post-hoc Tukey test. *P < 0.05, ***P < 0.001, ****P < 0.0001. f, Time of lipid mixing relative to IAV, LASV- or EBOV-pseudotyped particle colocalization with IFITM3 in HeLa cells (n > 50 dequenching particles tracked in three independent experiments for each sample).
Functional IFITM3 is required for virus particle fusion
To evaluate the features of IFITM3 that are required for trafficking to incoming IAV particles, we analyzed loss-of-function and key post-translational modification mutants. In particular, we evaluated the rs12252-C allele of IFITM3 that yields an N21Δ truncation mutant and is suggested to affect host cell susceptibility to IAV infection12,43. Stable expression of IFITM3-N21Δ in A549 IFITM2/3-KO cells revealed more peripheral plasma membrane distribution (Supplementary Fig. 11a), lower protein levels and reduced antiviral activity against IAV compared to WT IFITM3 (Supplementary Fig. 11b,c), consistent with previous studies12,43. Moreover, IFITM3-N21Δ also exhibited decreased antiviral activity toward IAV and ZIKV, in contrast to the similar levels of full-length IFITM2 or IFITM3 in HeLa IFITM2/3-KO (C5-2) cells (Fig. 5a–c). To analyze IFITM3-N21Δ trafficking during IAV infection, we generated a HA-IFITM3-N21Δ-V93-TAG construct for site-specific fluorophore labeling with BODIPY-Tz. Whereas wide-field epifluorescence microscopy showed that the BODIPY-labeled IFITM3 constructs were primarily in intracellular vesicles (Supplementary Fig. 12), confocal fluorescence microscopy showed that HA-IFITM3-N21Δ-V93-BODIPY exhibited slightly more co-staining with a plasma membrane marker compared to the HA-IFITM3-F8-BODIPY and HA-IFITM3-V93-BODIPY constructs (Supplementary Fig. 13a,b). For DiD-IAV imaging studies, we observed a 55% reduction in the number of IAV particles colocalized with IFITM3-N21Δ at the time of lipid mixing relative to WT IFITM3 (Fig. 5d–f), indicating that the truncated protein is less likely to be incorporated into, but not completely absent from, endosomal compartments supporting viral fusion (Fig. 5d–f). Reintroduction of WT IFITM3 and IFITM3-N21Δ into HeLa IFITM2/3-KO cells also did not significantly alter the total percentage of IAV particles undergoing lipid mixing by 2 h post-infection (Supplementary Fig. 10c). These results demonstrate that a quantitative difference in IFITM3 trafficking to incoming virus particles is correlated with altered steady state localization and antiviral activity, further suggesting that direct engagement or fusion of IFITM3-positive vesicles with pathogens is important for its cellular mechanism of action.
a, Western blot analysis of IFITM2 and IFITM2/3 in IFN-α stimulated HeLa WT and IFITM2/3-KO cells. High and low exposures of blots are included. Full gels are shown in Supplementary Fig. 17c. b, IAV infection of IFN-α stimulated HeLa WT and IFITM2/3-KO cells. Data represent the mean and s.d. of three independent experiments. c, ZIKV infection of IFN-α stimulated HeLa WT and IFITM2/3-KO cells. Data represent the mean and s.d. of three independent experiments. d, IAV-DID dequenching and trafficking with IFITM3-N21Δ. The arrows indicate the virion of interest. The numbers indicate the time post-infection (in minutes). Scale bar, 5 μm. e, Fluorescence intensity trace for the indicated particle in d. f, Relative percentage of DiD-IAV particles colocalized with IFITM3 and IFITM3-N21Δ at the time of dequenching. Data represent the mean and s.d. of three independent experiments (n > 100 dequenching particles total for each sample. P < 0.001 by an unpaired, two-sided t-test).
To further investigate mechanisms that control IFITM3 trafficking to virus particles, we analyzed site-specific mutants of S-palmitoylation14,44,45. Previous studies from our group demonstrated that Cys72 in human IFITM3 is the most prominent site of S-palmitoylation45,46 and is crucial for antiviral activity toward IAV45,47 as well as DENV47. However, the mechanism by which site-specific S-palmitoylation regulates IFITM3 antiviral activity is unclear, as similar levels of steady state protein expression and cellular localization patterns were observed with WT IFITM3 and individual Ala mutants at Cys71, Cys72 and Cys105 (ref. 45). We therefore generated F8-TAG constructs of these IFITM3 Cys-to-Ala mutants and analyzed their trafficking properties during IAV entry. These IFITM3 Cys-to-Ala mutants were expressed at similar levels to WT IFITM3-F8-TAG and showed decreased levels of S-fatty-acylation in HeLa IFITM2/3-KO (C5-2) cells, particularly with the C72A mutant (Supplementary Fig. 14a,b). Although the relative levels of site-specific S-fatty-acylation on these TAG-modified constructs of human IFITM3 are less dramatic than those of our previous studies with murine IFITM3 (ref. 45), Cys72 is still the most prominent site of modification46. Importantly, mutation of Cys72 to Ala most significantly decreased the antiviral activity of IFITM3 toward IAV, whereas Cys71 and Cys105 Ala mutants retained more activity (Supplementary Fig. 14c), consistent with previous studies from our group45 and others47. The steady state localization of these BODIPY-labeled constructs was similar to that in WT IFITM3 (Supplementary Figs. 9 and 15). However, the trafficking and colocalization of the IFITM3-C72A mutant with dequenched DiD-IAV particles were significantly decreased to ~20% compared to WT IFITM3 or Ala mutants at Cys71 and Cys105 (Fig. 6; Supplementary Fig. 16). In comparison to WT, IFITM3-C72A mutant trafficking to DiD-IAV particles was also delayed (Supplementary Fig. 14d). These results show that although S-fatty-acylation does not significantly affect IFITM3 steady state cellular distribution, site-specific S-palmitoylation of IFITM3 at Cys72 is important for its rate of trafficking to IAV particles during infection, which was only revealed by site-specific fluorophore labeling and live-cell imaging.
a, Intensity trace for DiD-IAV dequenching in the presence of IFITM3-F8-BODIPY-C71A. b, Intensity trace for DiD-IAV dequenching in the presence of IFITM3-F8-BODIPY-C72A. c, Intensity trace for DiD-IAV dequenching in the presence of IFITM3-F8-BODIPY-C105A. d, Total percentage of DiD-IAV particles colocalized with IFITM3 Cys mutants at the time of lipid mixing. Data represent the mean and s.d. of three independent experiments (n > 100 dequenching particles total for each sample). P values determined by one-way ANOVA with a post-hoc Tukey test. *P < 0.05, ***P < 0.001.
Discussion
IFITM1, IFITM2 and IFITM3 are important IFN-stimulated effectors that prevent diverse virus infections in a variety of vertebrate animals and are emerging as key virus susceptibility factors in humans. Nonetheless, the precise cellular and biochemical mechanisms of action for these small membrane proteins have been challenging to elucidate. By generating single and multiple IFITM-deficient mammalian cell lines, we demonstrated that IFITM2 and IFITM3 may be redundant and act synergistically with IFITM1 toward different restricted viruses in naïve and IFN-α-stimulated cells. Furthermore, we demonstrated that these IFITMs function in a dose-dependent fashion and do not affect early stages of exogenous cargo or pathogen uptake, but rather enhance the trafficking and turnover of late endocytic cargo to lysosomes. In the absence of infection, IFITM2/3-deficient cells stimulated with IFN-α showed slower rates of EGFR turnover. During IAV infection, we observed that fluorescently labeled IAV particles enter host cells and undergo hemifusion at similar rates in WT and IFITM2/3-deficient cells, consistent with previous siRNA knockdown studies of IFITM3 (ref. 36). However, in IFITM2/3-KO cells we observed that fluorescently labeled IAV particles traffic to lysosomes approximately 10 min more slowly on average. Our findings suggest that beyond potentially inhibiting virus pore formation36, IFITM2/3 may prevent virus entry by altering rates of virus-endosome fusion and/or accelerating the trafficking of endosomal cargo to lysosomes for destruction.
Although previous studies have correlated the cellular distribution of IFITM1, IFITM2 or IFITM3 with their antiviral activity, their trafficking and interactions with viruses during infection were unknown. Using site-specific fluorophore labeling of IFITM3 in mammalian cells, we were able to directly visualize IFITM3 trafficking with fluorescently labeled virus particles and observed for the first time that IFITM3-positive vesicles fuse with incoming virus-bearing vesicles before hemifusion. Importantly, the IFITM3-positive vesicle fusion and colocalization before hemifusion were specific to restricted viruses such as IAV and pseudotyped EBOV, but not LASV-pseudotyped virus particles, and colocalization was significantly reduced with loss-of-function mutants such as IFITM3-N21Δ. The ability of IFITMs to engage endosomes bearing incoming virus particles and enhance their trafficking to late endocytic compartments provides an important mechanism to explain the specificity of IFITMs against different viruses and potentially predicts points of viral entry into host cells. Different viruses may therefore initiate fusion in different subsets of late endo/lysosomal compartments and exhibit differential sensitivity to IFN effector expression levels and trafficking properties. These interactions are not readily apparent from bulk viral entry studies, which highlight the utility of our site-specific protein labeling and live single-particle imaging approaches that should facilitate future studies on virus-host factor dynamics.
Our ability to image IFITM3 trafficking in live cells has not only revealed interactions with pathogens during infection, but also uncovered how post-translational modifications regulate the function of these small membrane proteins. We initially discovered IFITM3 S-fatty-acylation14,44 and determined endogenous levels45 and specific sites of modification45,46, as well as confirmed its modification with palmitic acid46. Although these results have also been confirmed by others47,48,49 and have been shown for IFITM5 (ref. 50), how S-palmitoylation at Cys72 regulated IFITM3 antiviral activity was unclear from steady state localization studies. Using site-specific fluorophore labeling and live-cell imaging during infection, we now observe that inhibition of S-palmitoylation at Cys72, but not Cys71 or Cys105, by site-directed mutagenesis impairs IFITM3 trafficking to incoming IAV particles and thus reveals how site-specific S-palmitoylation regulates IFITM3 antiviral activity. The chemical imaging approach described here has uncovered important aspects of how IFITMs engage pathogens during infection, as well as key regulatory mechanisms, and should facilitate the analysis of other host factors involved in pathogen restriction. Nonetheless, key questions still remain regarding the cellular pathway and biochemical mechanism by which these IFN-effectors prevent pathogen entry and accelerate trafficking of incoming viral particles to lysosomes for destruction. Our ability to conduct site-specific labeling IFITM3 with unique chemical reactivity should enable additional imaging studies, discovery of specific protein-protein interaction partners and reconstitution of lipidated isoforms for biochemical and structural studies in the future.
Methods
Cell lines and reagents
Both HeLa and A549 cells were ordered from ATCC and cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco 12800–017) containing 10% FBS (Hyclone SN30087.02). The IFITM1, IFITM2 and IFITM3 antibodies were ordered from Cell Signaling Technology (13126 S), Proteintech group (66137–1-Ig) and Abgent (AP1153a). Anti-IFITM3 (polyclonal) 10088-604 was from VWR International. EGFR antibody [E235] was from Abcam (ab32077). TfR antibody was from R&D systems (AF2474). Alexa-Fluor 488-CD63 (sc-5275) was from Santa Cruz Biotechnology. Influenza A virus nucleoprotein antibody [AA5H] (ab20343) was from Abcam. Alexa-Fluor 647 Antibody Labeling Kit (A20186) was ordered from Life Technologies. The secondary antibodies HRP-conjugated goat anti-mouse IgG (HCL) (115-035-003) and HRP-conjugated goat anti-rabbit IgG (HCL) (111-035-003) were purchased from Jackson ImmunoResearch. Sheep IgG horseradish peroxidase-conjugated antibody (#HAF016) was ordered from Fisher Scientific. Anti-HA antibody conjugated to Alexa-Fluor 594 was purchased from Life Technologies. Anti-HA HRP-conjugated antibody was ordered from Roche. Anti-calnexin (ab22595) was from Abcam. Dextran-Alexa-Fluor 488 (D22910), dextran-pHrodo-red (P10361), transferrin-Alexa-Fluor 647 (T23366) and recombinant human EGF (10605HNAE25) were from Life Technologies. Human holo-Transferrin (T0665) was from Sigma-Aldrich. Human interferon-α (IFN-α, 8927SC) was from Cell Signaling Technology. Influenza A/PR/8/34 (H1N1) (10100374) was from Charles River Laboratories. Magic Red Cathepsin B Assay Kit (937) was from Immunochemistry Technologies. Bafilomycin A1 (ab120497) was from Abcam. 6 - TAMRA, Special Formulation (6-carboxytetramethylrhodamine, AS-81122) was from ANASPEC. QuikChange II Site-Directed Mutagenesis Kit (#200523) was from Agilent Technologies. T7 Endonuclease I (M0302S) was from New England BioLabs.
Plasmids
pEF-IFITM1, IFITM2, IFITM3 were cloned by PCR amplification and inserted into a pEF6-BSD vector. All mutants of IFITM3 were generated by using the QuikChange Multi Site-Directed Mutagenesis Kit (Stratagene). The pCAS9 plasmid and gRNA basic vector (pGL3-U6) were gifts from W. Wei (Peking University)51.
T7E1 assay
Cells were lysed for genome extraction after transfection with gRNA and cas9 plasmids for 60–72 h. The gRNA target site was amplified by using genome PCR primers (hIFITM3-GF1: AGGAAACT GTTGAGAAACCGAA; hIFITM3-GR1: GCTAGTGGATAGCCGGGGAC). The PCR products were gel purified, followed by denaturation and annealing. The annealing mixture was treated with T7 endonuclease I for 15 min at 37 °C, and the cleavage efficiency was then monitored by gel electrophoresis.
Generation and characterization of IFITM mutant cell lines by CRISPR–Cas9
The gRNA plasmid was constructed by inserting gRNA oligos into a pGL3-U6 vector through Golden Gate ligation. Cells were transfected with gRNA, pCas9 and pcDNA6-Puro with a ratio of 1:1:0.1 (μg). After 48 h, cells were detached and split into two parts. One part was lysed for T7E1 assay to check the cleavage efficiency of specific gRNA; another part was reseeded into six-well plates at different dilutions. Puromycin was added to select the positive clones. After selection for one week, the separated cell colonies were picked out into a new 24-well plate. New cell colonies were expanded and analyzed by western blot and genome sequencing to evaluate the knockout efficiency.
For western blot analysis, cells were lysed with sodium dodecyl sulfate (SDS) lysis buffer (2% SDS, 50 mM Tris, 2 mM EDTA) and boiled for 10 min at 98 °C. For immunoprecipitation, cells were lysed with Brij buffer (0.1 mM triethanolamine (TEA), 150 mM NaCl, 1% BrijO10 (pH7.4)) containing EDTA-free protease inhibitor mixture, then incubated with anti-HA affinity beads (Sigma). All primary antibodies were used at a 1:1,000 dilution, and secondary antibody was used at 1:5,000.
For genomic sequencing of IFITM clones, total genome was extracted using a genome extraction kit from QIAGEN. The target genomic site was amplified by gene-specific primers. The amplified fragments were added at the 3′ end by taq polymerase and then ligated with T vectors. After transformation, ten bacteria colonies were picked randomly and sent for sequencing.
Virus infections
A549, HeLa, Vero (WHO) and Huh-7.5 cells were maintained in DMEM supplemented with 10% FBS. A549 and HeLa cells were used in functional studies (viral replication assays), whereas Vero and Huh-7.5 cells were used for virus production and virus titration. All cell lines tested negative for contamination with mycoplasma and were obtained from ATCC (with the exception of Huh-7.5, which was from the Chisari lab (Scripps Research Institute, La Jolla, CA)).
HeLa or A549 cells were infected with influenza A/PR/8/34 virus (H1N1) at a MOI of 2.5 for 12 h. Infected cells were washed with PBS and harvested using 0.25% trypsin-EDTA. Cells were fixed in 3.7% paraformaldehyde (PFA) for 10 min and permeated with 0.1% Triton X-100 for 10 min. Cells were stained with anti-influenza nucleoprotein (NP) antibody (1:333; Abcam, ab 20343) and directly conjugated to Alexa-Fluor 647 using a 100 μg antibody labeling kit (Invitrogen). All antibodies were diluted in 0.1% Triton X-100 in PBS, and cells were stained for 20 min. Cells were washed twice with 0.1% Triton X-100 in PBS after each antibody treatment. PBS was used for final resuspension of cells for cytometric analysis using a FACS Canto II flow cytometer (BD Biosciences).
HeLa or A549 cells were seeded into 24-well plates (5 × 104 cells/well) and incubated overnight (16 h) in the presence or absence of IFN-α1 at a concentration of 100 μg/mL. Next, cells were washed with Opti-MEM (Gibco) before infection. Viruses were diluted in Opti-MEM at the indicated MOIs, and cells were inoculated for 2 h at 37 °C. Following incubation, viral inoculum was removed, and cultures were washed three times with Opti-MEM before DMEM was re-added. Treatment with IFN-α1 was continued throughout experiments. Cells infected with GFP-tagged viruses were harvested at indicated time points to measure infection by flow cytometry. The generation of viral stocks for the following viruses has been previously described: YFV-Venus52 (derived from YF17D-5′C25Venus2AUbi), WNV-GFP53 (derived from pBELO-WNV-GFP-RZ ic), DENV-GFP54 (derived from IC30P-A, a full-length infectious clone of strain 16681), VEEV-GFP55 (derived from pTC83-GFP infectious clone), ONNV-GFP56 (derived from infectious clone pONNV.GFP) and VSV-GFP (generously provided by J. Rose, Yale). ZIKV (PRVABC59 obtained from the CDC, Ft. Collins) was amplified in Huh-7.5 cells and titered by plaque assay on Huh-7.5 cells. Experiments with WNV were carried out in biosafety level 3 (BSL3) containment in compliance with institutional and federal guidelines. Infected cells were harvested into 300 μl Accumax cell dissociation medium (eBioscience) and transferred to a 96-well block containing 300 μl 4% PFA. Cells were pelleted at 930 r.c.f. for 5 min at 4 °C, resuspended in cold PBS containing 3% FBS, and stored at 4 °C until FACS analysis. Cells infected with ZIKV were harvested as described above and subsequently stained using a monoclonal antibody for flavivirus group antigen 4G2 (Millipore) at 1:500 as a primary antibody and anti-rabbit Alexa-Fluor 594 at 1:1,000 as a secondary antibody. In addition to cells, supernatants from ZIKV infections were harvested to determine viral titers by standard plaque assay performed on Huh-7.5 cells. All samples were analyzed using the LSRII flow cytometer (BD Biosciences) equipped with a 488-nm and a 561-nm laser for detection of GFP, YFP (Venus) and RFP. Data were analyzed using FlowJo software (Treestar) with a 0.1% compensation matrix.
Fluorescence microscopy
Cells were grown on glass coverslips in six-well plates. After treatments, cells were washed with PBS twice, fixed with 4% PFA for 10 min, permeabilized with 0.2% Triton X-100 for 10 min, blocked with 1% bovine serum albumin (BSA, Roche, 735094) for 1 h, and incubated with primary antibody rabbit anti-IFITM3 (1:100 diluted in 1% BSA; Proteintech group, 11714-1-AP) for 1.5 h, followed by secondary antibody anti-rabbit-Alexa Fluor 647 conjugate (1:1,000; Abcam, ab150083) for 1 h at room temperature. Finally, the coverslips were mounted onto slides using mounting medium containing DAPI, and the cells were examined by inverted LSM 780 laser scanning confocal microscope (Zeiss).
Exogenous cargo uptake assays
Cells were serum-starved for 45 min at 37 °C in DMEM and then incubated on ice for 5 min, which was followed by addition of 50 μg/ml Alexa-Fluor 647-conjugated transferrin (Invitrogen) in DMEM. After 30 min on ice, cells were transferred to 37 °C for 7 min or 20 min, pelleted, washed in ice-cold PBS, acid-washed (0.1 M glycine and 150 mM NaCl at pH 3) twice, washed with PBS once, and fixed (wt/vol 3% PFA and 4% sucrose in PBS) before flow cytometric analysis, as described above.
Analysis of dfTAT uptake and reduction
Synthesis of dfTAT (dimeric fluorescent TAT) by Y.-C. Wang was as previously described39. After pretreatment with or without 100 μg/ml IFN-α for 16 h, cells were incubated with 2 μM dfTAT in DMEM for the indicated time, placed on ice and washed by ice-cold PBS three times, and fixed (wt/vol 3% PFA and 4% sucrose in PBS) before flow cytometric analysis or fluorescence microscopy.
Analysis of EGFR and TfR turnover
Cells were cultured to 80% confluency, washed twice with PBS and starved in serum-free DMEM for 2 h at 37 °C to maximize surface EGFR and then pretreated with 10–25 μg/ml cycloheximide for 1 h in serum-free DMEM, preventing synthesis of new EGFR during stimulation by EGF. Immediately after, cells were continuously stimulated with 100 ng/ml EGF for 15, 30, 60, or 120 min in the presence of cycloheximide. The levels of EGFR and tubulin were determined by western blotting. For cell-surface staining experiments, cells were incubated with primary anti-EGFR antibody (Abcam, ab32077) on ice for 40 min and washed three times with PBS. Alexa-Fluor 647-conjugated secondary antibodies (Abcam, ab150083) were then incubated with the cells for 40 min on ice and washed three times with PBS. Cells were suspended in 2% FBS in PBS and immediately analyzed via flow cytometry.
For TfR turnover experiments, cells were cultured to 80% confluency, washed twice with PBS and starved in serum-free DMEM for 2 h at 37 °C, then pretreated with 10–25 μg/ml cycloheximide for 1 h in serum-free DMEM, preventing synthesis of new TfR. Immediately after, cells were continuously stimulated with 20 μg/ml Tf for 0, 30, 60, 120 or 180 min in the presence of cycloheximide. The levels of TfR, IFITM3 and tubulin were determined by western blotting.
Late endosome and lysosome fusion assay
Cells were incubated for 4 h in culture medium containing 25 μg/ml pHrodo-red dextran, followed by a 20 h chase in fluorophore-free medium to promote lysosomal accumulation of dextran. The cells were then incubated in medium containing 50 μg/ml dextran-Alexa Fluor 488 for 10 min at 37 °C before replacement with fluorophore-free medium. At the time points noted, cells were fixed with ice-cold 4% PFA for immunofluorescence staining.
Lysosomal protease activity assays
Cathepsin B activity was assayed using the cathepsin B activity kit following the protocol provided by the manufacturer. After treatment with IFN-α for 16 h with or without bafilomycin A1, cells were incubated with Magic Red (1:26) for 30–60 min at 37 °C before being washed twice with PBS. Cells were then fixed using 4% PFA and analyzed by flow cytometry.
Activity-based labeling of cathepsins (Vergent Bioscience, Pan Cathepsin probe) was performed as previously reported with minor modifications57. Briefly, cells were seeded 24 h before labeling, and cells were activated with IFN-α 16 h before labeling. Media were replaced, and cells were treated with 1 μM IABP Pan Cathepsin Probe for 2 h. Cells were washed with PBS and lysed on ice with a hypotonic lysis buffer (50 mM PIPES pH 7.4, 10 mM KCl, 5 mM MgCl2, 2 mM EDTA, 4 mM DTT, 1% NP-40). Cell debris was removed by spinning down lysates at 500 × g for 5 min at 4 °C. Supernatants were then mixed with 4X sample loading buffer, and 70 μg of total protein was analyzed by SDS–PAGE. The gel was washed twice with de-ionized water before in-gel fluorescence analysis on a Bio-Rad ChemiDoc MP Imaging System and Coomassie staining or transfer for western blot analysis.
DiD-labeling of viruses
Purified influenza virus A/PR/8/34 (H1N1) and recombinant VSV expressing LASV GPC (generated as previously described41) were labeled with self-quenching concentrations of the lipophilic dye 1,1′-dioctactadecyl-3,3,3′,3′-tetramethylindodicarbocyanine (DiD, Life Technologies). Purified virus (1 mg/ml) in PBS was incubated with 50 μM DiD while being agitated for 1 h at 4 °C. Virus was separated from excess dye by ultracentrifugation through a 10% sucrose cushion for 2 h at 107,000 × g and 4 °C using a SW41 rotor (Beckman Coulter). Labeled virus pellets were resuspended in PBS at a viral protein concentration of 1 mg/ml, aliquoted, and stored at −80 °C until use.
Site-specific labeling of IFITM3 in mammalian cells
For live-cell imaging studies, HeLa cells were seeded onto 35-mm glass-bottom dishes to be at approximately 70% confluency for imaging the next day. Following adherence, cells were transfected with the plasmid of interest containing a HA-IFITM3-TAG variant (1 μg per dish) and the Mm-PylRS-AF/Pyl-tRNACUA plasmid (1 μg per dish) using 3 μL Viafect (Promega) in complete cell growth media containing unnatural amino acids (TCOK). After overnight incubation, cell media were changed into fresh cell growth media without TCOK. After another 2 h culture at 37 °C/5% CO2, cells were labeled with tetrazine-fluorophores (250 nM) in FluoroBrite DMEM (Life Technologies) for 30 min at 37 °C and washed with complete cell growth media three times over 1 h.
Live cell imaging
For wide-field epifluorescence microscopy, cells were transduced with CellLight vectors (Life Technologies) for early endosomes (RFP-Rab5), late endosomes (RFP-Rab7), or lysosomes (RFP-LAMP1) approximately 18 h before imaging according to the manufacturer’s instructions. Live-cell microscopy was performed as previously described with an AxioObserver.Z1 wide-field epifluorescence microscope (Zeiss) equipped with a 40 ×/ 1.3 N.A. objective, DAPI/GFP/Texas Red/Cy5 filter set, and heated environmental enclosure maintained at 37 °C. HeLa cell monolayers were seeded onto fibronectin-coated 35-mm glass coverslip dishes (MatTek) 24 h before experiments. Cells were chilled on ice for several minutes before spinoculation of DiD-labeled virus onto monolayers at 1,500 × g and 6 °C for 20 min. Unbound particles were removed by five washes with cold PBS, and 500 μl cold imaging buffer (140 mM NaCl, 2.5 mM KCl, 1.8 mM MgCl2, 20 mM HEPES, 5 mM sucrose, 2 μM Hoechst 33342, and 2% FBS) was added to cover the cells. The dish was immediately mounted on the microscope objective and focused. The coverslip dish was then flooded with 1.5 ml warm imaging buffer to mark the start of experiments (t = 0). Images were acquired every 10 s over the duration of the experiments using a single Z-section, which encompassed nearly all cell-associated particles.
For confocal microscopy, HeLa cells were seeded in 35-mm glass-bottom dishes (ibidi) and cultured overnight. The next day cells were transfected with the plasmid of interest containing a HA-IFITM3-TAG variant (1 µg per dish) and the Mm-PylRS-AF/Pyl-tRNACUA plasmid (1 µg per dish) using 6 µl Lipofectamine 3000 in complete cell growth media containing TCOK. After 16 h incubation, cell media were changed into fresh cell growth media without TCOK. After another 6 h culture at 37 °C/5% CO2, cells were labeled with tetrazine-BODIPY (500 nM) in FluoroBrite DMEM (Life Technologies)/10% FBS for 0.5 h at 37 °C and washed with FluoroBrite DMEM (Life Technologies)/10% FBS four times over 2 h. Finally, cells were stained with NucBlue Live Cell Stain (Life Technologies) for 20 min and imaged in FluoroBrite DMEM/10% FBS with confocal microscopy. For plasma membrane staining, cells were stained with CellMask Red (1:1,000, Life Technologies) in Live Cell Imaging Solution (LCIS) for 5 min at 37 °C and washed with LCIS three times before staining with NucBlue Live Cell Stain. Cells were imaged on an inverted LSM 780 laser scanning confocal microscope (Zeiss) with a Zeiss Plan-Apochromatic 63×/1.4 N.A. oil immersion objective. The microscope was equipped with a stage-top incubator set at 37 °C/5% CO2 for live-cell imaging. Hoechst was excited with a 405-nm laser with emission spectra collected between 410 nm and 480 nm. Tz-BODIPY was excited with a 488-nm laser with emission spectra collected between 490 nm and 550 nm. CellMask Red was excited with a 561-nm laser with emission spectra collected between 570 nm and 620 nm. Images were acquired with the ZEN blue 2012 software (Zeiss) and analyzed by ImageJ (NIH). Pearson’s correlation coefficients were calculated in ImageJ using the Coloc 2 plugin.
Data analysis
Image analysis and single-particle tracking were performed using Volocity software (PerkinElmer) as previously described41. Image files were not manipulated, apart from minor adjustments in brightness and contrast. Viral puncta were thresholded by initial intensity and size. Puncta falling outside the range of 0.25 to 1 μm2 expected of individual DiD-labeled virions were excluded from single-particle analysis. Virions were considered colocalized with LAMP1 or IFITM3 only if the cellular marker punctum exceeded the background signal by 30% or more and if the GFP or BODIPY and DiD puncta co-trafficked with greater than 70% overlap of signals. Mean measurements ( ± s.d.) were derived from three separate experiments, unless otherwise indicated.
Analysis of HA-IFITM3-TCOK expression levels
For western blot analysis, HeLa cells were seeded in six-well plates and cultured overnight. IFITM2/3-KO cells were transfected with HA-IFITM3-F8TAG (2.5 µg) and Mm-PylRS-AF/Pyl-tRNACUA (2.5 µg) in the presence of TCOK (50 µM) for 16 h. Hela WT cells were left untreated or treated with IFN-α (100 ng/ml) for 16 h. The levels of IFITM3 were determined by western blotting using rabbit anti-IFITM3 (1:1,000; Proteintech group, 11714-1-AP) and secondary antibody HRP-conjugated goat anti-rabbit IgG (HCL; 111-035-003).
For immunofluorescence analysis, IFITM2/3-KO cells were transfected with HA-IFITM3-F8TAG (1 µg) and Mm-PylRS-AF/Pyl-tRNACUA (1 µg) in the presence of TCOK (50 µM) for 16 h. Hela WT cells were untreated or treated with IFN-α (100 ng/ml) for 16 h. Cells were then fixed with formaldehyde, permeabilized with saponin (0.5%), and immunostained with rabbit anti-IFITM3 (1:100) (Proteintech group, 11714-1-AP) and anti-rabbit-Alexa Fluor 647 conjugate (1:1,000) (Abcam, ab150083) for confocal fluorescence imaging. Hoescht (blue) is used to stain nuclei. Cells were imaged on an inverted LSM 780 laser scanning confocal microscope (Zeiss) with a Zeiss Plan-Apochromatic 63×/1.4 oil immersion objective. Images were taken under identical confocal microscope settings.
For flow cytometry analysis, IFITM2/3 KO cells were transfected with HA-IFITM3-F8TAG (1 µg), Mm-PylRS-AF//Pyl-tRNACUA (1 µg), in the presence of TCOK (50 µM) for 16 h. Hela WT cells were treated with or without IFN-α (100 ng/ml) for 16 h. Cells were washed twice with PBS and then fixed with 400 μl of PBS with 4% PFA for 10 min. The fixed cells were permeabilized with 200 μl of 0.2% saponin in PBS for 10 min and then blocked with 200 μl 0.2% BSA and 0.2% saponin in PBS for 10 min. Cells were treated with rabbit anti-IFITM3 (1:300) (Proteintech group, 11714-1-AP) and anti-rabbit-Alexa Fluor 647 (1:1,000) (Abcam, ab150083) in PBS with 0.02% saponin. After three washes with PBS, cells were resuspended in 150 μL PBS with 0.2% BSA and 0.02% saponin. The samples were analyzed by flow cytometry (BD LSRII). Data analysis was performed using FLowJo software.
IAV infection with IFITM3 Cys mutants
IFITM2/3 KO HeLa cells were seeded in 12 well plates (Corning) and cultured overnight. Cells were co-transfected with HA-hIFITM3 constructs (1 μg/well) and the Mm-PylRS-AF/Pyl-tRNA plasmid (1 μg/well) using 3 μl Lipofectamine 3000 in 1 ml of complete cell growth media containing 10 μl of 100 mM BocK (final concentration 1 mM). After 16 h, cells were infected with influenza virus A/PR/8/34 virus (H1N1) with MOI of 5. After 6 h of infection, cells were trypsinized and collected in cluster tubes (Corning). Cells were washed twice with PBS and then fixed with 400 μl of PBS with 4% PFA for 10 min. The fixed cells were permeabilized with 200 μl of 0.2% saponin in PBS for 10 min and then blocked with 200 μL 0.2% BSA and 0.2% saponin in PBS for 10 min. Cells were treated with anti-influenza NP antibody conjugated to AlexaFluor-647 and anti-HA antibody conjugated to Alexa-fluor-594 (1:250) in PBS with 0.02% saponin. After three washes with PBS, cells were resuspended in 150 μl PBS with 0.2% BSA and 0.02% saponin. The samples were analyzed by flow cytometry (BD LSRII). All samples were first gated by HA-positive staining, indicating successful transfection, and then NP-positive percentage of NP-positive staining, indicating successful infection. The HA-tag epitope is derived from an H3 influenza virus strain and is not present in the PR8 strain of H1N1 influenza virus. Data analysis was performed using FLowJo software.
S-palmitoylation assays
Metabolic labeling of cells with the alkyne-palmitic acid reporter alk-16 was performed as previously described58 with some modifications. Alk-16 and az-rho were synthesized as previously reported58. IFITM2/3 KO HeLa cells were transfected with HA-hIFITM3 constructs as described above. After 16 h, cells were incubated with 50 μM alk-16 in DMEM containing 10% FBS for 2 h. Cells were harvested, washed once with PBS and lysed in 1% Brij 97 (Sigma) in 50 mM TEA, 150 mM NaCl pH 7.4, 1X Roche protease inhibitor, and 1,500 units/ml benzonase (EMD). Protein concentrations were determined by the BCA assay. For immunoprecipitation, 150 μg of total protein was added to 20 μl of anti-HA antibody-conjugated agarose (Sigma) in a total volume of 150 μl and rocked at 4 °C for 1 h. Agarose beads were washed by resuspension in 500 μl of 50 mM HEPES buffer (containing 150 mM NaCl, pH 7.4) by centrifugation at 3,500 × g for 30 s. The beads were then resuspended in 45 μl of the above buffer and 5 μl of CuAAC reactant solution (0.5 μl of 10 mM azido-rhodamine (final concentration 100 μM), 1 μl of 50 mM freshly prepared CuSO4·5H2O in H2O (final concentration 1 mM, Sigma), 1 μl of 50 mM freshly prepared TCEP (final concentration 1 mM) and 2.5 μl of 10 mM Tris[(1-benzyl-1H-1,2,3-triazol-4-yl)methyl]amine (TBTA) (final concentration 500 μM)) was added. The samples were rocked at room temperature for 1 h and washed twice with RIPA buffer. Laemmli sample buffer (20 μl) was added to the samples (1.0:1.3 ratio of buffer to sample), which were heated for 10 min at 95 °C and separated by gel electrophoresis. In-gel fluorescence scanning was performed using a Bio-Rad ChemiDoc MP Imaging System. Western blots for HA-tagged proteins were performed using an anti-HA HRP conjugated antibody (1:1,000; Roche). Quantification of band intensities in fluorescence gels and western blots were performed with Image Lab (Bio-Rad). Data from three biological replicates were quantified and averaged for plotting.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Bailey, C. C., Zhong, G., Huang, I. C. & Farzan, M. IFITM-family proteins: the cell’s first line of antiviral defense. Annu. Rev. Virol. 1, 261–283 (2014).
Brass, A. L. et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 139, 1243–1254 (2009).
Perreira, J. M., Chin, C. R., Feeley, E. M. & Brass, A. L. IFITMs restrict the replication of multiple pathogenic viruses. J. Mol. Biol. 425, 4937–4955 (2013).
Savidis, G. et al. The IFITMs inhibit Zika virus replication. Cell Rep. 15, 2323–2330 (2016).
Monel, B. et al. Zika virus induces massive cytoplasmic vacuolization and paraptosis-like death in infected cells. EMBO J. 36, 1653–1668 (2017).
Warren, C. J. et al. The antiviral restriction factors IFITM1, 2 and 3 do not inhibit infection of human papillomavirus, cytomegalovirus and adenovirus. PLoS One 9, e96579 (2014).
Zhao, X. et al. Interferon induction of IFITM proteins promotes infection by human coronavirus OC43. Proc. Natl. Acad. Sci. USA 111, 6756–6761 (2014).
Zhao, X. et al. Identification of residues controlling restriction versus enhancing activities of IFITM proteins on the entry of human coronaviruses. J. Virol. 92, e01535-17 (2018).
Ranjbar, S., Haridas, V., Jasenosky, L. D., Falvo, J. V. & Goldfeld, A. E. A role for IFITM proteins in restriction of Mycobacterium tuberculosis infection. Cell Rep. 13, 874–883 (2015).
Wakim, L. M., Gupta, N., Mintern, J. D. & Villadangos, J. A. Enhanced survival of lung tissue-resident memory CD8+ T cells during infection with influenza virus due to selective expression of IFITM3. Nat. Immunol. 14, 238–245 (2013).
Bailey, C. C., Huang, I. C., Kam, C. & Farzan, M. Ifitm3 limits the severity of acute influenza in mice. PLoS Pathog. 8, e1002909 (2012).
Everitt, A. R. et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 (2012).
Wu, X. et al. Intrinsic immunity shapes viral resistance of stem cells. Cell 172, 423–438.e425 (2018).
Yount, J. S., Karssemeijer, R. A. & Hang, H. C. S-palmitoylation and ubiquitination differentially regulate interferon-induced transmembrane protein 3 (IFITM3)-mediated resistance to influenza virus. J. Biol. Chem. 287, 19631–19641 (2012).
Bailey, C. C., Kondur, H. R., Huang, I. C. & Farzan, M. Interferon-induced transmembrane protein 3 is a type II transmembrane protein. J. Biol. Chem. 288, 32184–32193 (2013).
Weston, S. et al. A membrane topology model for human interferon inducible transmembrane protein 1. PLoS One 9, e104341 (2014).
Ling, S. et al. Combined approaches of EPR and NMR illustrate only one transmembrane helix in the human IFITM3. Sci. Rep. 6, 24029 (2016).
Chesarino, N. M. et al. IFITM3 requires an amphipathic helix for antiviral activity. EMBO Rep. 18, 1740–1751 (2017).
Chesarino, N. M., McMichael, T. M. & Yount, J. S. Regulation of the trafficking and antiviral activity of IFITM3 by post-translational modifications. Future Microbiol. 9, 1151–1163 (2014).
Peng, T. & Hang, H. C. Site-specific bioorthogonal labeling for fluorescence imaging of intracellular proteins in living cells. J. Am. Chem. Soc. 138, 14423–14433 (2016).
Zhang, Y. H. et al. Interferon-induced transmembrane protein-3 genetic variant rs12252-C is associated with severe influenza in Chinese individuals. Nat. Commun. 4, 1418 (2013).
Wang, Z. et al. Early hypercytokinemia is associated with interferon-induced transmembrane protein-3 dysfunction and predictive of fatal H7N9infection. Proc. Natl. Acad. Sci. USA 111, 769–774 (2014).
Yang, X. et al. Interferon-inducible transmembrane protein 3 genetic variant rs12252 and influenza susceptibility and severity: a meta-analysis. PLoS One 10, e0124985 (2015).
Zhang, Y. et al. Interferon-induced transmembrane protein-3 rs12252-C is associated with rapid progression of acute HIV-1 infection in Chinese MSM cohort. AIDS 29, 889–894 (2015).
López-Rodríguez, M. et al. IFITM3 and severe influenza virus infection. No evidence of genetic association. Eur. J. Clin. Microbiol. Infect. Dis. 35, 1811–1817 (2016).
Naderi, M. et al. Evaluation of interferon-induced transmembrane protein-3 (IFITM3) rs7478728 and rs3888188 polymorphisms and the risk of pulmonary tuberculosis. Biomed. Rep. 5, 634–638 (2016).
Randolph, A. G. et al. Evaluation of IFITM3 rs12252 association with severe pediatric influenza infection. J. Infect. Dis. 216, 14–21 (2017).
Williams, D. E. et al. IFITM3 polymorphism rs12252-C restricts influenza A viruses. PLoS One 9, e110096 (2014).
Makvandi-Nejad, S. et al. Lack of truncated IFITM3 transcripts in cells homozygous for the rs12252-C variant that is associated with severe influenza infection. J. Infect. Dis. 217, 257–262 (2018).
Allen, E. K. et al. SNP-mediated disruption of CTCF binding at the IFITM3 promoter is associated with risk of severe influenza in humans. Nat. Med. 23, 975–983 (2017).
Feeley, E. M. et al. IFITM3 inhibits influenza A virus infection by preventing cytosolic entry. PLoS Pathog. 7, e1002337 (2011).
Wee, Y. S., Roundy, K. M., Weis, J. J. & Weis, J. H. Interferon-inducible transmembrane proteins of the innate immune response act as membrane organizers by influencing clathrin and v-ATPase localization and function. Innate Immun. 18, 834–845 (2012).
Huang, I. C. et al. Distinct patterns of IFITM-mediated restriction of filoviruses, SARS coronavirus, and influenza A virus. PLoS Pathog. 7, e1001258 (2011).
Li, K. et al. IFITM proteins restrict viral membrane hemifusion. PLoS Pathog. 9, e1003124 (2013).
Amini-Bavil-Olyaee, S. et al. The antiviral effector IFITM3 disrupts intracellular cholesterol homeostasis to block viral entry. Cell Host Microbe 13, 452–464 (2013).
Desai, T. M. et al. IFITM3 restricts influenza A virus entry by blocking the formation of fusion pores following virus-endosome hemifusion. PLoS Pathog. 10, e1004048 (2014).
Desai, T. M., Marin, M., Mason, C. & Melikyan, G. B. pH regulation in early endosomes and interferon-inducible transmembrane proteins control avian retrovirus fusion. J. Biol. Chem. 292, 7817–7827 (2017).
Shi, G., Schwartz, O. & Compton, A. A. More than meets the I: the diverse antiviral and cellular functions of interferon-induced transmembrane proteins. Retrovirology 14, 53 (2017).
Erazo-Oliveras, A. et al. Protein delivery into live cells by incubation with an endosomolytic agent. Nat. Methods 11, 861–867 (2014).
Eierhoff, T., Hrincius, E. R., Rescher, U., Ludwig, S. & Ehrhardt, C. The epidermal growth factor receptor (EGFR) promotes uptake of influenza A viruses (IAV) into host cells. PLoS Pathog. 6, e1001099 (2010).
Spence, J. S., Krause, T. B., Mittler, E., Jangra, R. K. & Chandran, K. Direct visualization of ebola virus fusion triggering in the endocytic pathway. mBio 7, e01857–e15 (2016).
Wrensch, F. et al. Interferon-induced transmembrane protein-mediated inhibition of host cell entry of ebolaviruses. J. Infect. Dis. 212, S210–S218 (2015).
Jia, R. et al. The N-terminal region of IFITM3 modulates its antiviral activity by regulating IFITM3 cellular localization. J. Virol. 86, 13697–13707 (2012).
Yount, J. S. et al. Palmitoylome profiling reveals S-palmitoylation-dependent antiviral activity of IFITM3. Nat. Chem. Biol. 6, 610–614 (2010).
Percher, A. et al. Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation. Proc. Natl. Acad. Sci. USA 113, 4302–4307 (2016).
Thinon, E., Fernandez, J. P., Molina, H. & Hang, H. C. Selective enrichment and direct analysis of protein S-palmitoylation sites. J. Proteome. Res. 17, 1907–1922 (2018).
John, S. P. et al. The CD225 domain of IFITM3 is required for both IFITM protein association and inhibition of influenza A virus and dengue virus replication. J. Virol. 87, 7837–7852 (2013).
McMichael, T. M. et al. The palmitoyltransferase ZDHHC20 enhances interferon-induced transmembrane protein 3 (IFITM3) palmitoylation and antiviral activity. J. Biol. Chem. 292, 21517–21526 (2017).
Narayana, S. K. et al. The Interferon-induced transmembrane proteins, IFITM1, IFITM2, and IFITM3 inhibit Hepatitis C virus entry. J. Biol. Chem. 290, 25946–25959 (2015).
Tsukamoto, T. et al. Role of S-palmitoylation on IFITM5 for the interaction with FKBP11 in osteoblast cells. PLoS One 8, e75831 (2013).
Zhou, Y. et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Jones, C. T. et al. Real-time imaging of hepatitis C virus infection using a fluorescent cell-based reporter system. Nat. Biotechnol. 28, 167–171 (2010).
McGee, C. E. et al. Infection, dissemination, and transmission of a West Nile virus green fluorescent protein infectious clone by Culex pipiens quinquefasciatus mosquitoes. Vector Borne Zoonotic Dis. 10, 267–274 (2010).
Schoggins, J. W. et al. Dengue reporter viruses reveal viral dynamics in interferon receptor-deficient mice and sensitivity to interferon effectors in vitro. Proc. Natl. Acad. Sci. USA 109, 14610–14615 (2012).
Petrakova, O. et al. Noncytopathic replication of Venezuelan equine encephalitis virus and eastern equine encephalitis virus replicons in Mammalian cells. J. Virol. 79, 7597–7608 (2005).
Brault, A. C. et al. Infection patterns of o’nyong nyong virus in the malaria-transmitting mosquito, Anopheles gambiae. Insect Mol. Biol. 13, 625–635 (2004).
Verdoes, M. et al. Improved quenched fluorescent probe for imaging of cysteine cathepsin activity. J. Am. Chem. Soc. 135, 14726–14730 (2013).
Charron, G. et al. Robust fluorescent detection of protein fatty-acylation with chemical reporters. J. Am. Chem. Soc. 131, 4967–4975 (2009).
Acknowledgements
E.T. acknowledges support a Marie Skłodowska-Curie postdoctoral fellowship. T.D. is supported by the Tri-Institutional Program in Chemical Biology at The Rockefeller University. H.-H.H. and C.M.R. acknowledge support from NIH R01AI091707. We thank W. Wei (Peking University) for sharing pCAS9 plasmid and gRNA vector (pGL3-U6). We thank Y.-C. Wang (The Rockefeller University) for the synthesis of dfTAT. We thank J. Yount and members of the Hang laboratory for helpful comments and discussion of the paper. T.P. acknowledges support from the National Natural Science Foundation of China (No. 21778010), the Shenzhen Science and Technology Innovation Committee (JCYJ20170412150832022), and Shenzhen Peacock Plan (KQTD2015032709315529). K.C. acknowledges grant support from NIH-NIAID R56AI088027. H.C.H. acknowledges grant support from NIH-NIGMS R01GM087544.
Author information
Authors and Affiliations
Contributions
J.S.S., R.H., T.P., K.C. and H.C.H. conceived the study. J.S.S., R.H., H.-H.H., E.T., T.D., C.M.R., T.P., K.C. and H.C.H. planned the experiments. R.H. generated IFITM1, IFITM2 and IFITM3 knockout mammalian cell lines and performed cell biology studies as well as IAV infection experiments. J.S.S. performed live-cell imaging studies of viruses and IFITM3. H.-H.H. performed other virus infection experiments. T.D. performed S-fatty-acylation experiments, TfR turnover and additional IFITM3 imaging experiments. E.T. performed protease activity studies. T.P. generated reagents for site-specific labeling and live-cell imaging of IFITM3. J.S.S., R.H., H.-H.H., E.T., T.D., C.M.R., T.P., K.C. and H.C.H. interpreted the data. C.M.R., K.C. and H.C.H. supervised the study. J.S.S. and H.C.H. wrote the manuscript with input from other co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–2, Supplementary Figures 1–18
Supplementary Video 1
Example of DiD-IAV dequenching in a LAMP-GFP-expressing HeLa IFITM2/3-KO cell. The white arrow marks the particle of interest, and the red arrow indicates the onset of dequenching.
Supplementary Video 2
Example of DiD-IAV dequenching following colocalization with IFITM3-F8-BODIPY in a HeLa IFITM2/3-KO cell. The white arrow marks the particle of interest, and the red arrow indicates the onset of dequenching.
Supplementary Video 3
Example of DiD-IAV dequenching prior to colocalization with IFITM3-F8-BODIPY in a HeLa IFITM2/3-KO cell. The white arrow marks the particle of interest, and the red arrow indicates the onset of dequenching.
Supplementary Video 4
Example of LASV GPC-pseudotyped DiD-VSV dequenching following colocalization with IFITM3-F8-BODIPY in a HeLa IFITM2/3-KO cell. The white arrow marks the particle of interest, and the red arrow indicates the onset of dequenching.
Rights and permissions
About this article
Cite this article
Spence, J.S., He, R., Hoffmann, HH. et al. IFITM3 directly engages and shuttles incoming virus particles to lysosomes. Nat Chem Biol 15, 259–268 (2019). https://doi.org/10.1038/s41589-018-0213-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0213-2
This article is cited by
-
Bioorthogonal Reactions in Bioimaging
Topics in Current Chemistry (2024)
-
The relationship between extreme inter-individual variation in macrophage gene expression and genetic susceptibility to inflammatory bowel disease
Human Genetics (2024)
-
Chaperone-mediated autophagy plays an important role in regulating retinal progenitor cell homeostasis
Stem Cell Research & Therapy (2022)
-
Transcriptome profile of spleen tissues from locally-adapted Kenyan pigs (Sus scrofa) experimentally infected with three varying doses of a highly virulent African swine fever virus genotype IX isolate: Ken12/busia.1 (ken-1033)
BMC Genomics (2022)
-
Ultrasensitive Ribo-seq reveals translational landscapes during mammalian oocyte-to-embryo transition and pre-implantation development
Nature Cell Biology (2022)