Abstract
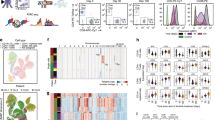
The Bruton tyrosine kinase (BTK) inhibitor ibrutinib has substantially improved therapeutic options for chronic lymphocytic leukemia (CLL). Although ibrutinib is not curative, it has a profound effect on CLL cells and may create new pharmacologically exploitable vulnerabilities. To identify such vulnerabilities, we developed a systematic approach that combines epigenome profiling (charting the gene-regulatory basis of cell state) with single-cell chemosensitivity profiling (quantifying cell-type-specific drug response) and bioinformatic data integration. By applying our method to a cohort of matched patient samples collected before and during ibrutinib therapy, we identified characteristic ibrutinib-induced changes that provide a starting point for the rational design of ibrutinib combination therapies. Specifically, we observed and validated preferential sensitivity to proteasome, PLK1, and mTOR inhibitors during ibrutinib treatment. More generally, our study establishes a broadly applicable method for investigating treatment-specific vulnerabilities by integrating the complementary perspectives of epigenetic cell states and phenotypic drug responses in primary patient samples.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The ATAC-seq and pharmacoscopy data are available from http://cll-combinations.computational-epigenetics.org. The ATAC-seq data are also available from NCBI GEO under accession number GSE100672. The source code for ATAC-seq data processing is available from a Github repository linked on the above website.
References
Woyach, J. A., Johnson, A. J. & Byrd, J. C. The B-cell receptor signaling pathway as a therapeutic target in CLL. Blood 120, 1175–1184 (2012).
Stevenson, F. K., Krysov, S., Davies, A. J., Steele, A. J. & Packham, G. B-cell receptor signaling in chronic lymphocytic leukemia. Blood 118, 4313–4320 (2011).
Burger, J. A. & Chiorazzi, N. B cell receptor signaling in chronic lymphocytic leukemia. Trends Immunol. 34, 592–601 (2013).
Byrd, J. C. et al. Targeting BTK with ibrutinib in relapsed chronic lymphocytic leukemia. N. Engl. J. Med. 369, 32–42 (2013).
O’Brien, S. et al. Ibrutinib for patients with relapsed or refractory chronic lymphocytic leukaemia with 17p deletion (RESONATE-17): a phase 2, open-label, multicentre study. Lancet. Oncol. 17, 1409–1418 (2016).
Burger, J. A. et al. Ibrutinib as initial therapy for patients with chronic lymphocytic leukemia. N. Engl. J. Med. 373, 2425–2437 (2015).
Herman, S. E. et al. Bruton tyrosine kinase represents a promising therapeutic target for treatment of chronic lymphocytic leukemia and is effectively targeted by PCI-32765. Blood 117, 6287–6296 (2011).
Ponader, S. et al. The Bruton tyrosine kinase inhibitor PCI-32765 thwarts chronic lymphocytic leukemia cell survival and tissue homing in vitro and in vivo. Blood 119, 1182–1189 (2012).
de Rooij, M. F. et al. The clinically active BTK inhibitor PCI-32765 targets B-cell receptor- and chemokine-controlled adhesion and migration in chronic lymphocytic leukemia. Blood 119, 2590–2594 (2012).
Chen, S. S. et al. BTK inhibition results in impaired CXCR4 chemokine receptor surface expression, signaling and function in chronic lymphocytic leukemia. Leukemia 30, 833–843 (2016).
Burger, J. A. et al. Leukemia cell proliferation and death in chronic lymphocytic leukemia patients on therapy with the BTK inhibitor ibrutinib. JCI Insight 2, e89904 (2017).
Burger, J. A. et al. Clonal evolution in patients with chronic lymphocytic leukaemia developing resistance to BTK inhibition. Nat. Commun. 7, 11589 (2016).
Herman, S. E. et al. Ibrutinib inhibits BCR and NF-κB signaling and reduces tumor proliferation in tissue-resident cells of patients with CLL. Blood 123, 3286–3295 (2014).
Bottoni, A. et al. Targeting BTK through microRNA in chronic lymphocytic leukemia. Blood 128, 3101–3112 (2016).
Landau, D. A. et al. The evolutionary landscape of chronic lymphocytic leukemia treated with ibrutinib targeted therapy. Nat. Commun. 8, 2185 (2017).
Maddocks, K. & Jones, J. A. Bruton tyrosine kinase inhibition in chronic lymphocytic leukemia. Semin. Oncol. 43, 251–259 (2016).
Maddocks, K. J. et al. Etiology of ibrutinib therapy discontinuation and outcomes in patients with chronic lymphocytic leukemia.JAMA Oncol. 1, 80–87 (2015).
Woyach, J. A. et al. Resistance mechanisms for the Bruton’s tyrosine kinase inhibitor ibrutinib. N. Engl. J. Med. 370, 2286–2294 (2014).
Lamothe, B. et al. Proteasome inhibitor carfilzomib complements ibrutinib’s action in chronic lymphocytic leukemia. Blood 125, 407–410 (2015).
Deng, J. et al. Bruton’s tyrosine kinase inhibition increases BCL-2 dependence and enhances sensitivity to venetoclax in chronic lymphocytic leukemia. Leukemia 31, 2075–2084 (2017).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Snijder, B. et al. Image-based ex-vivo drug screening for patients with aggressive haematological malignancies: interim results from a single-arm, open-label, pilot study. Lancet Haematol. 4, e595–e606 (2017).
Rendeiro, A. F. et al. Chromatin accessibility maps of chronic lymphocytic leukaemia identify subtype-specific epigenome signatures and transcription regulatory networks. Nat. Commun. 7, 11938 (2016).
Nückel, H. et al. Lipoprotein lipase expression is a novel prognostic factor in B-cell chronic lymphocytic leukemia. Leuk. Lymphoma 47, 1053–1061 (2006).
O’Brien, S. et al. Single-agent ibrutinib in treatment-naïve and relapsed/refractory chronic lymphocytic leukemia: a 5-year experience. Blood 131, 1910–1919 (2018).
Sheffield, N. C. & Bock, C. LOLA: enrichment analysis for genomic region sets and regulatory elements in R and Bioconductor. Bioinformatics 32, 587–589 (2016).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Bucur, O. et al. PLK1 is a binding partner and a negative regulator of FOXO3 tumor suppressor. Discoveries (Craiova) 2, e16 (2014).
Shehata, M. et al. Reconstitution of PTEN activity by CK2 inhibitors and interference with the PI3-K/Akt cascade counteract the antiapoptotic effect of human stromal cells in chronic lymphocytic leukemia. Blood 116, 2513–2521 (2010).
Bock, C. & Lengauer, T. Managing drug resistance in cancer: lessons from HIV therapy. Nat. Rev. Cancer 12, 494–501 (2012).
Chari, A. et al. Phase 1 trial of ibrutinib and carfilzomib combination therapy for relapsed or relapsed and refractory multiple myeloma. Leuk. Lymphoma. 59:11, 2588–2594.
Dietrich, S. et al. Drug-perturbation-based stratification of blood cancer. J. Clin. Invest. 128, 427–445 (2018).
Tao, Y. F. et al. Inhibiting PLK1 induces autophagy of acute myeloid leukemia cells via mammalian target of rapamycin pathway dephosphorylation. Oncol. Rep. 37, 1419–1429 (2017).
Giulino-Roth, L. et al. Inhibition of Hsp90 suppresses PI3K/AKT/mTOR signaling and has antitumor activity in Burkitt lymphoma. Mol. Cancer Ther. 16, 1779–1790 (2017).
Murray, M.Y. et al. Ibrutinib inhibits BTK-driven NF-kappaB p65 activity to overcome bortezomib-resistance in multiple myeloma. Cell Cycle 14, 2367–75 (2015).
Mathews Griner, L. A. et al. High-throughput combinatorial screening identifies drugs that cooperate with ibrutinib to kill activated B-cell-like diffuse large B-cell lymphoma cells. Proc. Natl Acad. Sci. USA 111, 2349–2354 (2014).
Ezell, S. A. et al. Synergistic induction of apoptosis by combination of BTK and dual mTORC1/2 inhibitors in diffuse large B cell lymphoma. Oncotarget 5, 4990–5001 (2014).
Tenenbaum, D. KEGGREST: Client-side REST access to KEGG. R package version 1.14.1. (2017).
Jiang, H., Lei, R., Ding, S. W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinformatics 15, 182 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome. Biol. 9, R137 (2008).
Hoffman, M. M. et al. Integrative annotation of chromatin elements from ENCODE data. Nucleic Acids Res. 41, 827–841 (2013).
Ernst, J. & Kellis, M. Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues. Nat. Biotechnol. 33, 364–376 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome. Biol. 15, 550 (2014).
Vladimer, G. I. et al. Global survey of the immunomodulatory potential of common drugs. Nat. Chem. Biol. 13, 681–690 (2017).
Griffith, M. et al. DGIdb: mining the druggable genome. Nat. Methods 10, 1209–1210 (2013).
Acknowledgements
We thank all patients who have donated their samples for this study. We also thank the Biomedical Sequencing Facility at CeMM for assistance with next generation sequencing and J. Bigenzahn, M. Rebsamen as well as the G.S.-F. and C.B. labs for help and advice. This work was performed in the context of the following grants and fellowships: WWTF LS16-034 to G.S.-F. and U.J.; FWF SFB F 4711-B20 to G.S.-F.; EMBO Long-Term Fellowship 1543-2012 to G.I.V. and 733-2016 to T.P.; Swiss National Science Foundation Fellowship P300P3_147897 and PP00P3_163961 to B.S.; Marie-Sklodowska Curie Action Fellowship 703668 to N.K.; Feodor Lynen Fellowship of the Alexander von Humboldt Foundation to C. Schmidl; Marie Curie Action International Outgoing Fellowship (PIOF-2013-624924) to M.G.; Initiative Krebsforschung (UE71104017, UE71104005, UE71504001, and UE711043037), Austrian Society of Hematology and Oncology (ÖGHO AP00359OFF), and Anniversary Fund of the Austrian National Bank (OeNB AP130120ONB) to M.S.; Austrian Academy of Sciences New Frontiers Group Award and ERC Starting Grant (European Union’s Horizon 2020 research and innovation programme) 679146 to C.B.
Author information
Authors and Affiliations
Contributions
C. Schmidl and T.K. performed ATAC-seq experiments; G.I.V., C.T., A.R., and K.R. performed image-based chemosensitivity experiments; C. Schmidl, G.I.V., A.F.R., N.K., B.S., O.L.d.l.F., and S.K. analyzed the ATAC-seq and image-based chemosensitivity data; S.S., C.T., T.P., M.A., R.H., D.D., M.H., and M.S. handled patient samples and performed validation experiments; M.S. and U.J. were responsible for study ethics; C. Skrabs, E.P., M.G., G.H., P.B.S., M.S., and U.J. provided and analyzed clinical data or oversaw patient care and ethics; C. Schmidl, G.I.V., A.F.R., T.P., M.S., G.S.-F., U.J., and C.B. wrote the manuscript; M.S., G.S.-F., U.J., and C.B. oversaw the project.
Corresponding author
Ethics declarations
Competing interests
G.I.V., N.K., B.S., G.S.-F. are co-founders of Allcyte GmbH, which has licensed the pharmacoscopy technology, and they are listed as inventors on patent applications for the pharmacoscopy / single-cell imaging methodology. G.I.V. and N.K. have become employees of Allcyte GmbH during the course of this study. U.J. received research grants and honoraria from Janssen Cilag, Abbvie, Novartis, and Roche Austria.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7
Source Data for figures
Source data for figures
Supplementary Table 1
Overview and clinical annotation of patient samples included in this study
Supplementary Table 2
Sample-specific sequencing statistics for the ATAC-seq experiments
Supplementary Table 3
List of all chromatin accessible regions detected in the ATAC-seq dataset
Supplementary Table 4
List of genomic regions with differential chromatin accessibility upon ibrutinib treatment
Supplementary Table 5
List of drugs and small molecules for the pharmacoscopy experiments
Supplementary Table 6
Selectivity scores before and during ibrutinib treatment for 131 drugs
Rights and permissions
About this article
Cite this article
Schmidl, C., Vladimer, G.I., Rendeiro, A.F. et al. Combined chemosensitivity and chromatin profiling prioritizes drug combinations in CLL. Nat Chem Biol 15, 232–240 (2019). https://doi.org/10.1038/s41589-018-0205-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0205-2
This article is cited by
-
Standardized assays to monitor drug sensitivity in hematologic cancers
Cell Death Discovery (2023)
-
MiR-145 inhibits the differentiation and proliferation of bone marrow stromal mesenchymal stem cells by GABARAPL1 in steroid-induced femoral head necrosis
BMC Musculoskeletal Disorders (2022)
-
Chromatin accessibility profiling by ATAC-seq
Nature Protocols (2022)
-
Chromatin accessibility profiling methods
Nature Reviews Methods Primers (2021)
-
Survey of ex vivo drug combination effects in chronic lymphocytic leukemia reveals synergistic drug effects and genetic dependencies
Leukemia (2020)