Abstract
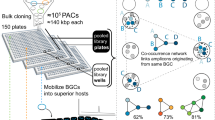
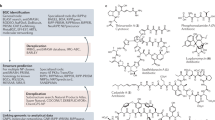
Bacteria contain an immense untapped trove of novel secondary metabolites in the form of ‘silent’ biosynthetic gene clusters (BGCs). These can be identified bioinformatically but are not expressed under normal laboratory growth conditions. Methods to access their products would dramatically expand the pool of bioactive compounds. We report a universal high-throughput method for activating silent BGCs in diverse microorganisms. Our approach relies on elicitor screening to induce the secondary metabolome of a given strain and imaging mass spectrometry to visualize the resulting metabolomes in response to ~500 conditions. Because it does not require challenging genetic, cloning, or culturing procedures, this method can be used with both sequenced and unsequenced bacteria. We demonstrate the power of the approach by applying it to diverse bacteria and report the discovery of nine cryptic metabolites with potentially therapeutic bioactivities, including a new glycopeptide chemotype with potent inhibitory activity against a pathogenic virus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and the supplementary material. NMR data used to characterize the cryptic metabolites are available from the corresponding author upon reasonable request. The DNA sequence of the ker gene cluster from A. keratiniphila has been deposited in GenBank (accession no. MH428036). The LAESI–IMS data for S. canus and A. keratiniphila, including the raw data for Figs. 3a and 4a as well as the source code used to generate the 3D plots, have been deposited in the Global Natural Products Social Molecular Networking (GNPS) database (MassIVE accession number MSV000082658).
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 79, 629–661 (2016).
Cragg, G. M. & Newman, D. J. Natural products: a continuing source of novel drug leads. Biochim. Biophys. Acta 1830, 3670–3695 (2013).
Cragg, G. M., Grothaus, P. G. & Newman, D. J. Impact of natural products on developing new anti-cancer agents. Chem. Rev. 109, 3012–3043 (2009).
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002).
Ikeda, H. et al. Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat. Biotechnol. 21, 526–531 (2003).
Oliynyk, M. et al. Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338. Nat. Biotechnol. 25, 447–453 (2007).
Nett, M., Ikeda, H. & Moore, B. S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 26, 1362–1384 (2009).
Liu, X. & Cheng, Y. Q. Genome-guided discovery of diverse natural products from Burkholderia sp. J. Ind. Microbiol. Biotechnol. 41, 275–284 (2014).
Baltz, R. H. Gifted microbes for genome mining and natural product discovery. J. Ind. Microbiol. Biotechnol. 44, 573–588 (2017).
Okada, B. K. & Seyedsayamdost, M. R. Antibiotic dialogues: induction of silent biosynthetic gene clusters by exogenous small molecules. FEMS Microbiol. Rev. 41, 19–33 (2017).
Ochi, K. & Hosaka, T. New strategies for drug discovery: activation of silent or weakly expressed microbial gene clusters. Appl. Microbiol. Biotechnol. 97, 87–98 (2013).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 13, 509–523 (2015).
Zhu, H., Sandiford, S. K. & van Wezel, G. P. Triggers and cues that activate antibiotic production by actinomycetes. J. Ind. Microbiol. Biotechnol. 41, 371–386 (2014).
Nah, H. J., Pyeon, H. R., Kang, S. H., Choi, S. S. & Kim, E. S. Cloning and heterologous expression of a large-sized natural product biosynthetic gene cluster in streptomyces species. Front. Microbiol. 8, 394 (2017).
Ren, H., Wang, B. & Zhao, H. Breaking the silence: new strategies for discovering novel natural products. Curr. Opin. Biotechnol. 48, 21–27 (2017).
Yoon, V. & Nodwell, J. R. Activating secondary metabolism with stress and chemicals. J. Ind. Microbiol. Biotechnol. 41, 415–424 (2014).
Guo, F. et al. Targeted activation of silent natural product biosynthesis pathways by reporter-guided mutant selection. Metab. Eng. 28, 134–142 (2015).
Seyedsayamdost, M. R. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc. Natl Acad. Sci. USA 111, 7266–7271 (2014).
Xu, F., Nazari, B., Moon, K., Bushin, L. B. & Seyedsayamdost, M. R. Discovery of a cryptic antifungal compound from Streptomyces albus J1074 using high-throughput elicitor screens. J. Am. Chem. Soc. 139, 9203–9212 (2017).
Rosen, P. C. & Seyedsayamdost, M. R. Though much is taken, much abides: finding new antibiotics using old ones. Biochemistry 56, 4925–4926 (2017).
Watrous, J. D. & Dorrestein, P. C. Imaging mass spectrometry in microbiology. Nat. Rev. Microbiol. 9, 683–694 (2011).
Gross, H. et al. The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem. Biol. 14, 53–63 (2007).
Nemes, P. & Vertes, A. Laser ablation electrospray ionization for atmospheric pressure, in vivo, and imaging mass spectrometry. Anal. Chem. 79, 8098–8106 (2007).
Li, H., Balan, P. & Vertes, A. Molecular imaging of growth, metabolism, and antibiotic inhibition in bacterial colonies by laser ablation electrospray ionization mass spectrometry. Angew. Chem. Int. Edn Engl. 55, 15035–15039 (2016).
Fincher, J. A. et al. Enhanced sensitivity and metabolite coverage with remote laser ablation electrospray ionization-mass spectrometry aided by coaxial plume and gas dynamics. Analyst 142, 3157–3164 (2017).
Li, H. & Vertes, A. Solvent gradient electrospray for laser ablation electrospray ionization mass spectrometry. Analyst 142, 2921–2927 (2017).
Heinemann, B., Kaplan, M. A., Muir, R. D. & Hooper, I. R. Amphomycin, a new antibiotic. Antibiot. Chemother. (Northfield) 3, 1239–1242 (1953).
Bodanszky, M., Sigler, G. F. & Bodanszky, A. Structure of the peptide antibiotic amphomycin. J. Am. Chem. Soc. 95, 2352–2357 (1973).
Maksimov, M. O., Pan, S. J. & Link, A.J. Lasso peptides: structure, function, biosynthesis, and engineering. Nat. Prod. Rep. 29, 996–1006 (2012).
Hegemann, J. D., Zimmermann, M., Xie, X. & Marahiel, M. A. Lasso peptides: an intriguing class of bacterial natural products. Acc. Chem. Res. 48, 1909–1919 (2015).
Güntert, P., Mumenthaler, C. & Wüthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Herrmann, T., Güntert, P. & Wüthrich, K. Protein NMR structure determination with automated NOE-identification in the NOESY spectra using the new software ATNOS. J. Biomol. NMR 24, 171–189 (2002).
Zhu, S. et al. Insights into the unique phosphorylation of the lasso peptide paeninodin. J. Biol. Chem. 291, 13662–13678 (2016).
Tietz, J. I. et al. A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat. Chem. Biol. 13, 470–478 (2017).
Price, J. C., Barr, E. W., Tirupati, B., Bollinger, J. M. Jr. & Krebs, C. The first direct characterization of a high-valent iron intermediate in the reaction of an α-ketoglutarate-dependent dioxygenase: a high-spin FeIV complex in taurine/α-ketoglutarate dioxygenase (TauD) from Escherichia coli. Biochemistry 42, 7497–7508 (2003).
Krebs, C., Galonić Fujimori, D., Walsh, C. T. & Bollinger, J. M. Jr. Non-heme Fe(IV)-oxo intermediates. Acc. Chem. Res. 40, 484–492 (2007).
Tiwari, K. & Gupta, R. K. Rare actinomycetes: a potential storehouse for novel antibiotics. Crit. Rev. Biotechnol. 32, 108–132 (2012).
Nicolaou, K. C., Boddy, C. N., Bräse, S. & Winssinger, N. Chemistry, biology, and medicine of the glycopeptide antibiotics. Angew. Chem. Int. Edn Engl. 38, 2096–2152 (1999).
Hubbard, B. K. & Walsh, C. T. Vancomycin assembly: nature’s way. Angew. Chem. Int. Edn Engl. 42, 730–765 (2003).
Shen, B., Liu, W. & Nonaka, K. Enediyne natural products: biosynthesis and prospect towards engineering novel antitumor agents. Curr. Med. Chem. 10, 2317–2325 (2003).
Everest, G. J. & Meyers, P. R. Evaluation of the antibiotic biosynthetic potential of the genus Amycolatopsis and description of Amycolatopsis circi sp. nov., Amycolatopsis equina sp. nov. and Amycolatopsis hippodromi sp. nov. J. Appl. Microbiol. 111, 300–311 (2011).
Heald, S. L., Mueller, L. & Jeffs, P. W. Actinoidins A and A2: structure determination using 2D NMR methods. J. Antibiot. (Tokyo) 40, 630–645 (1987).
Berdnikova, T. F., Lomakina, N. N. & Potapova, N. P. [Structure of actinoidins A and B]. Antibiotiki 27, 252–258 (1982).
Diekema, D. J. & Jones, R. N. Oxazolidinone antibiotics. Lancet 358, 1975–1982 (2001).
Jorquera, P. A. & Tripp, R. A. Respiratory syncytial virus: prospects for new and emerging therapeutics. Expert Rev. Respir. Med. 11, 609–615 (2017).
Yang, Y. L., Xu, Y., Straight, P. & Dorrestein, P. C. Translating metabolic exchange with imaging mass spectrometry. Nat. Chem. Biol. 5, 885–887 (2009).
Kersten, R. D. et al. A mass spectrometry-guided genome mining approach for natural product peptidogenomics. Nat. Chem. Biol. 7, 794–802 (2011).
Traxler, M. F., Watrous, J. D., Alexandrov, T., Dorrestein, P. C. & Kolter, R. Interspecies interactions stimulate diversification of the Streptomyces coelicolor secreted metabolome. mBio 4, e00459–13 (2013).
Du, L. et al. Unique amalgamation of primary and secondary structural elements transform peptaibols into potent bioactive cell-penetrating peptides. Proc. Natl Acad. Sci. USA 114, E8957–E8966 (2017).
Craney, A., Ozimok, C., Pimentel-Elardo, S. M., Capretta, A. & Nodwell, J. R. Chemical perturbation of secondary metabolism demonstrates important links to primary metabolism. Chem. Biol. 19, 1020–1027 (2012).
Goodwin, C. R. et al. Structuring microbial metabolic responses to multiplexed stimuli via self-organizing metabolomics maps. Chem. Biol. 22, 661–670 (2015).
Okada, B. K., Wu, Y., Mao, D., Bushin, L. B. & Seyedsayamdost, M. R. Mapping the trimethoprim-induced secondary metabolome of Burkholderia thailandensis. ACS Chem. Biol. 11, 2124–2130 (2016).
Davies, J., Spiegelman, G. B. & Yim, G. The world of subinhibitory antibiotic concentrations. Curr. Opin. Microbiol. 9, 445–453 (2006).
Yim, G., Wang, H. H. & Davies, J. Antibiotics as signalling molecules. Phil. Trans. R. Soc. Lond. B 362, 1195–1200 (2007).
Romero, D., Traxler, M. F., López, D. & Kolter, R. Antibiotics as signal molecules. Chem. Rev. 111, 5492–5505 (2011).
Hu, H. & Ochi, K. Novel approach for improving the productivity of antibiotic-producing strains by inducing combined resistant mutations. Appl. Environ. Microbiol. 67, 1885–1892 (2001).
Yilmaz, E. M. & Güntert, P. NMR structure calculation for all small molecule ligands and non-standard residues from the PDB Chemical Component Dictionary. J. Biomol. NMR 63, 21–37 (2015).
Acknowledgements
We thank the National Institutes of Health (DP2-AI-124786 to M.R.S.), the Burroughs Wellcome Fund, and the Princeton IP Accelerator Fund for support of this work. K.M.D. was supported by an Arnold O. Beckman Postdoctoral Fellowship. L.B.B. was supported by a National Science Foundation Graduate Research Fellowship.
Author information
Authors and Affiliations
Contributions
F.X., Y.W., and M.R.S. designed the research; F.X., Y.W., and C.Z. carried out high-throughput elicitor screens and validations; Y.W. conducted the LAESI–IMS experiments and analyzed the results; F.X. purified and solved the structures of all secondary metabolites, with assistance from K.M.D. and K.M. Y.W. conducted structure calculations, with assistance from L.B.B. F.X. carried out bioinformatic analyses. M.R.S. wrote the manuscript, to which all authors contributed. F.X. and Y.W. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–14 and Supplementary Figures 1–6
Rights and permissions
About this article
Cite this article
Xu, F., Wu, Y., Zhang, C. et al. A genetics-free method for high-throughput discovery of cryptic microbial metabolites. Nat Chem Biol 15, 161–168 (2019). https://doi.org/10.1038/s41589-018-0193-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0193-2
This article is cited by
-
Escherichia coli has an undiscovered ability to inhibit the growth of both Gram-negative and Gram-positive bacteria
Scientific Reports (2024)
-
Manipulation and epigenetic control of silent biosynthetic pathways in actinobacteria
World Journal of Microbiology and Biotechnology (2024)
-
Microbial antagonists: diversity, formulation and applications for management of pest–pathogens
Egyptian Journal of Biological Pest Control (2023)
-
Cirsiliol induces autophagy and mitochondrial apoptosis through the AKT/FOXO1 axis and influences methotrexate resistance in osteosarcoma
Journal of Translational Medicine (2023)
-
Newest perspectives of glycopeptide antibiotics: biosynthetic cascades, novel derivatives, and new appealing antimicrobial applications
World Journal of Microbiology and Biotechnology (2023)