Abstract
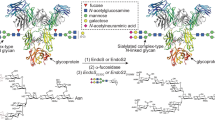
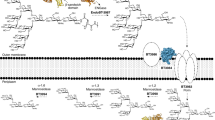
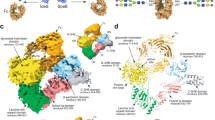
The identification of host protein substrates is key to understanding effector glycosyltransferases secreted by pathogenic bacteria and to using them for glycoprotein engineering. Here we report a chemical method for tagging, enrichment, and site-specific proteomic profiling of effector-modified proteins in host cells. Using this method, we discover that Legionella effector SetA α-O-glucosylates various eukaryotic proteins by recognizing a S/T-X-L-P/G sequence motif, which can be exploited to site-specifically introduce O-glucose on recombinant proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on reasonable request.
References
Schmaltz, R. M., Hanson, S. R. & Wong, C.-H. Chem. Rev. 111, 4259–4307 (2011).
Moremen, K. W. et al. Nat. Chem. Biol. 14, 156–162 (2017).
Nothaft, H. & Szymanski, C. M. Nat. Rev. Microbiol. 8, 765–778 (2010).
Keys, T. G. & Aebi, M. Curr. Opin. Syst. Biol. 5, 23–31 (2017).
Schwarz, F. et al. Nat. Chem. Biol. 6, 264–266 (2010).
Valderrama-Rincon, J. D. et al. Nat. Chem. Biol. 8, 434–436 (2012).
Naegeli, A. et al. J. Biol. Chem. 289, 24521–24532 (2014).
Xu, Y. et al. Chem. Commun. (Camb.) 53, 9075–9077 (2017).
Kightlinger, W. et al. Nat. Chem. Biol. 14, 627–635 (2018).
Lu, Q., Li, S. & Shao, F. Trends Microbiol. 23, 630–641 (2015).
Jank, T., Belyi, Y. & Aktories, K. Cell Microbiol. 17, 1752–1765 (2015).
Sun, Y., Willis, L. M., Batchelder, H. R. & Nitz, M. Chem. Commun. (Camb.) 52, 13024–13026 (2016).
Just, I. et al. Nature 375, 500–503 (1995).
Belyi, Y. et al. Proc. Natl. Acad. Sci. USA 103, 16953–16958 (2006).
López Aguilar, A. et al. ACS. Chem. Biol. 12, 611–621 (2017).
Jank, T. et al. Cell Microbiol. 14, 852–868 (2012).
O’Shea, J. P. et al. Nat. Methods 10, 1211–1212 (2013).
Rana, N. A. & Haltiwanger, R. S. Curr. Opin. Struct. Biol. 21, 583–589 (2011).
Yu, H. et al. Nat. Chem. Biol. 12, 735–740 (2016).
Steinemann, M., Schlosser, A., Jank, T. & Aktories, K. Proc. Natl. Acad. Sci. USA 115, 9580–9585 (2018).
Levanova, N. et al. Naunyn Schmiedebergs Arch. Pharmacol. 322, 390 (2018).
Wang, Z. et al. Cell Disc. 4, 53 (2018).
Shen, D. L. et al. ACS Chem. Biol. 12, 206–213 (2017).
Darabedian, N., Gao, J., Chuh, K. N., Woo, C. M. & Pratt, M. R. J. Am. Chem. Soc. 140, 7092–7100 (2018).
Li, S. et al. Nature 501, 242–246 (2013).
Berger, K. H. & Isberg, R. R. Mol. Microbiol. 7, 7–19 (1993).
Besanceney-Webler, C. et al. Angew. Chem. Int. Ed. Engl. 50, 8051–8056 (2011).
Qin, K. et al. ACS Chem. Biol. 13, 1983–1989 (2018).
Xu, L. et al. PLoS Pathog. 6, e1000822 (2010).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. Genome Res. 14, 1188–1190 (2004).
Eswar, N. et al. Curr. Protoc. Bioinformatics Chapter 5, Unit 5.6 (2006).
Larkin, M. A. et al. Bioinformatics 23, 2947–2948 (2007).
Maier, J. A. et al. J. Chem. Theory. Comput. 11, 3696–3713 (2015).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. J. Comput. Chem. 25, 1157–1174 (2004).
Petersen, H. G. J. Chem. Phys. 103, 3668–3679 (1995).
Acknowledgements
We thank Z. Luo for providing L. pneumophila strains, W. Zhou at the mass spectrometry facility of the National Center for Protein Sciences at Peking University for assistance with proteomics analysis, and the High Performace Computing Platform of the Center for Life Sciences for supporting protein structure modeling. This work is supported by the National Key R&D Program of China (numbers 2018YFA0507600 and 2016YFA0501500 to X.C. and 2016YFA0502300 to L.L.) and the National Natural Science Foundation of China (numbers 21425204, 21672013, 91753206, and 21521003 to X.C.).
Author information
Authors and Affiliations
Contributions
L.G. and X.C. conceived the project; L.G. performed experiments with the help of Q.S., Y.Z., T.W., N.D., J.X., and F.S.; H.L. and L.L. performed the structural modeling; L.G. and X.C. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Note 1
Rights and permissions
About this article
Cite this article
Gao, L., Song, Q., Liang, H. et al. Legionella effector SetA as a general O-glucosyltransferase for eukaryotic proteins. Nat Chem Biol 15, 213–216 (2019). https://doi.org/10.1038/s41589-018-0189-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0189-y
This article is cited by
-
Core-predominant gut fungus Kazachstania slooffiae promotes intestinal epithelial glycolysis via lysine desuccinylation in pigs
Microbiome (2023)
-
Lactobacillus reuteri-derived extracellular vesicles maintain intestinal immune homeostasis against lipopolysaccharide-induced inflammatory responses in broilers
Journal of Animal Science and Biotechnology (2021)