Abstract
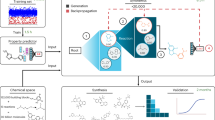
Specific RNA structures control numerous metabolic processes that impact human health, and yet efforts to target RNA structures de novo have been limited. In eukaryotes, the self-splicing group II intron is a mitochondrial RNA tertiary structure that is absent in vertebrates but essential for respiration in plants, fungi and yeast. Here we show that this RNA can be targeted through a process of high-throughput in vitro screening, SAR and lead optimization, resulting in high-affinity compounds that specifically inhibit group IIB intron splicing in vitro and in vivo and lack toxicity in human cells. The compounds are potent growth inhibitors of the pathogen Candida parapsilosis, displaying antifungal activity comparable to that of amphotericin B. These studies demonstrate that RNA tertiary structures can be successfully targeted de novo, resulting in pharmacologically valuable compounds.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Authors can confirm that all relevant data are included in the paper and/or its supplementary information files.
References
Blount, K. F. & Breaker, R. R. Riboswitches as antibacterial drug targets. Nat. Biotechnol. 24, 1558–1564 (2006).
Howe, J. A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015).
Howe, J. A. et al. Atomic resolution mechanistic studies of ribocil: a highly selective unnatural ligand mimic of the E. coli FMN riboswitch. RNA Biol. 13, 946–954 (2016).
Wang, H. et al. Dual-targeting small-molecule inhibitors of the Staphylococcus aureus FMN riboswitch disrupt riboflavin homeostasis in an infectious setting. Cell Chem. Biol. 24, 576–588 (2017).
Kim, J. N. et al. Design and antimicrobial action of purine analogues that bind guanine riboswitches. ACS Chem. Biol. 4, 915–927 (2009).
Blount, K. F., Wang, J. X., Lim, J., Sudarsan, N. & Breaker, R. R. Antibacterial lysine analogs that target lysine riboswitches. Nat. Chem. Biol. 3, 44–49 (2007).
Blount, K. F. et al. Novel riboswitch-binding flavin analog that protects mice against Clostridium difficile infection without inhibiting cecal flora. Antimicrob. Agents Chemother. 59, 5736–5746 (2015).
von Ahsen, U., Davies, J. & Schroeder, R. Antibiotic inhibition of group I ribozyme function. Nature 353, 368–370 (1991).
Disney, M. D., Childs, J. L. & Turner, D. H. Hoechst 33258 selectively inhibits group I intron self-splicing by affecting RNA folding. ChemBioChem 5, 1647–1652 (2004).
Mei, H. Y., Cui, M., Lemrow, S. M. & Czarnik, A. W. Discovery of selective, small-molecule inhibitors of RNA complexes. II. Self-splicing group I intron ribozyme. Bioorg. Med. Chem. 5, 1185–1195 (1997).
Patwardhan, N. N. et al. Amiloride as a new RNA-binding scaffold with activity against HIV-1 TAR. MedChemComm 8, 1022–1036 (2017).
Schroeder, R., Waldsich, C. & Wank, H. Modulation of RNA function by aminoglycoside antibiotics. EMBO J. 19, 1–9 (2000).
Colameco, S. & Elliot, M. A. Non-coding RNAs as antibiotic targets. Biochem. Pharmacol. 133, 29–42 (2017).
Wilson, D. N. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat. Rev. Microbiol. 12, 35–48 (2014).
Blaha, G. M., Polikanov, Y. S. & Steitz, T. A. Elements of ribosomal drug resistance and specificity. Curr. Opin. Struct. Biol. 22, 750–758 (2012).
Eubanks, C. S., Forte, J. E., Kapral, G. J. & Hargrove, A. E. Small molecule-based pattern recognition to classify RNA structure. J. Am. Chem. Soc. 139, 409–416 (2017).
Morgan, B. S., Forte, J. E., Culver, R. N., Zhang, Y. & Hargrove, A. E. Discovery of key physicochemical, structural, and spatial properties of RNA-targeted bioactive ligands. Angew. Chem. Int. Edn. Engl. 56, 13498–13502 (2017).
Bernat, V. & Disney, M. D. RNA structures as mediators of neurological diseases and as drug targets. Neuron 87, 28–46 (2015).
Costales, M. G. et al. Small molecule inhibition of microRNA-210 reprograms an oncogenic hypoxic circuit. J. Am. Chem. Soc. 139, 3446–3455 (2017).
Yang, W. Y., Gao, R., Southern, M., Sarkar, P. S. & Disney, M. D. Design of a bioactive small molecule that targets r(AUUCU) repeats in spinocerebellar ataxia 10. Nat. Commun. 7, 11647 (2016).
Dibrov, S. M. et al. Hepatitis C virus translation inhibitors targeting the internal ribosomal entry site. J. Med. Chem. 57, 1694–1707 (2014).
Barros, S. A., Yoon, I. & Chenoweth, D. M. Modulation of the E. coli rpoH temperature sensor with triptycene-cased small molecules. Angew. Chem. Int. Edn Engl. 55, 8258–8261 (2016).
Sztuba-Solinska, J. et al. Identification of biologically active, HIV TAR RNA-binding small molecules using small molecule microarrays. J. Am. Chem. Soc. 136, 8402–8410 (2014).
Lorenz, D. A., Song, J. M. & Garner, A. L. High-throughput platform assay technology for the discovery of pre-microRNA-selective small molecule probes. Bioconjug. Chem. 26, 19–23 (2015).
Kett, D. H., Azoulay, E., Echeverria, P. M. & Vincent, J. L. Candida bloodstream infections in intensive care units: analysis of the extended prevalence of infection in intensive care unit study. Crit. Care Med. 39, 665–670 (2011).
Guinea, J. Global trends in the distribution of Candida species causing candidemia. Clin. Microbiol. Infect. 20, 5–10 (2014).
Morales, D. K. et al. Control of Candida albicans metabolism and biofilm formation by Pseudomonas aeruginosa phenazines. MBio 4, e00526–12 (2013).
Marcia, M. & Pyle, A. M. Principles of ion recognition in RNA: insights from the group II intron structures. RNA 20, 516–527 (2014).
Zhao, C. & Pyle, A. M. Structural insights into the mechanism of group II intron splicing. Trends Biochem. Sci. 42, 470–482 (2017).
Perlman, P. S. Genetic analysis of RNA splicing in yeast mitochondria. Methods Enzymol. 181, 539–558 (1990).
Rossignol, T. et al. Correlation between biofilm formation and the hypoxic response in Candida parapsilosis. Eukaryot. Cell 8, 550–559 (2009).
Richard, M. L., Nobile, C. J., Bruno, V. M. & Mitchell, A. P. Candida albicans biofilm-defective mutants. Eukaryot. Cell 4, 1493–1502 (2005).
Su, L. J., Waldsich, C. & Pyle, A. M. An obligate intermediate along the slow folding pathway of a group II intron ribozyme. Nucleic Acids Res. 33, 6674–6687 (2005).
Perez-Martinez, X., Broadley, S. A. & Fox, T. D. Mss51p promotes mitochondrial Cox1p synthesis and interacts with newly synthesized Cox1p. EMBO J. 22, 5951–5961 (2003).
Dziembowski, A. et al. The yeast mitochondrial degradosome. Its composition, interplay between RNA helicase and RNase activities and the role in mitochondrial RNA metabolism. J. Biol. Chem. 278, 1603–1611 (2003).
Luedtke, N. W., Liu, Q. & Tor, Y. RNA-ligand interactions: affinity and specificity of aminoglycoside dimers and acridine conjugates to the HIV-1 Rev response element. Biochemistry 42, 11391–11403 (2003).
Tanner, M. & Cech, T. Activity and thermostability of the small self-splicing group I intron in the pre-tRNA(lle) of the purple bacterium Azoarcus. RNA 2, 74–83 (1996).
Li, C. F., Costa, M., Bassi, G., Lai, Y. K. & Michel, F. Recurrent insertion of 5′-terminal nucleotides and loss of the branchpoint motif in lineages of group II introns inserted in mitochondrial preribosomal RNAs. RNA 17, 1321–1335 (2011).
Moen, M. D., Lyseng-Williamson, K. A. & Scott, L. J. Liposomal amphotericin B: a review of its use as empirical therapy in febrile neutropenia and in the treatment of invasive fungal infections. Drugs 69, 361–392 (2009).
Velagapudi, S. P. et al. Design of a small molecule against an oncogenic noncodingRNA. Proc. Natl Acad. Sci. USA 113, 5898–5903 (2016).
Mulhbacher, J. et al. Novel riboswitch ligand analogs as selective inhibitors of guanine-related metabolic pathways. PLoS Pathog. 6, e1000865 (2010).
Baell, J. & Walters, M. A. Chemistry: chemical con artists foil drug discovery. Nature 513, 481–483 (2014).
Baell, J. B. & Nissink, J. W. M. Seven year itch: pan-assay interference compounds (PAINS) in 2017—utility and limitations. ACS Chem. Biol. 13, 36–44 (2018).
Jasial, S., Hu, Y. & Bajorath, J. How frequently are pan-assay interference compounds active? Large-scale analysis of screening data reveals diverse activity profiles, low global hit frequency, and many consistently inactive compounds. J. Med. Chem. 60, 3879–3886 (2017).
Capuzzi, S. J., Muratov, E. N. & Tropsha, A. Phantom PAINS: problems with the utility of alerts for pan-assay INterference CompoundS. J. Chem. Inf. Model. 57, 417–427 (2017).
Re, A., Joshi, T., Kulberkyte, E., Morris, Q. & Workman, C. T. RNA-protein interactions: an overview. Methods Mol. Biol. 1097, 491–521 (2014).
Wincott, F. et al. Synthesis, deprotection, analysis and purification of RNA and ribozymes. Nucleic Acids Res. 23, 2677–2684 (1995).
Dickey, T. H. & Pyle, A. M. The SMAD3 transcription factor binds complex RNA structures with high affinity. Nucleic Acids Res. 45, 11980–11988 (2017).
Chin, K. & Pyle, A. M. Branch-point attack in group II introns is a highly reversible transesterification, providing a potential proofreading mechanism for 5′-splice site selection. RNA 1, 391–406 (1995).
Fedorova, O., Mitros, T. & Pyle, A. M. Domains 2 and 3 interact to form critical elements of the group II intron active site. J. Mol. Biol. 330, 197–209 (2003).
Pyle, A. M. & Green, J. B. Building a kinetic framework for group II intron ribozyme activity: quantitation of interdomain binding and reaction rate. Biochemistry 33, 2716–2725 (1994).
Zingler, N., Solem, A. & Pyle, A. M. Dual roles for the Mss116 cofactor during splicing of the ai5γ group II intron. Nucleic Acids Res. 38, 6602–6609 (2010).
Daniels, D. L., Michels, W. J. Jr & Pyle, A. M. Two competing pathways for self-splicing by group II introns: a quantitative analysis of in vitro reaction rates and products. J. Mol. Biol. 256, 31–49 (1996).
Zhang, J. H., Chung, T. D. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Clinical and Laboratory Standards Institute. Reference Method For Broth Dilution Antifungal SusceptibilityTesting of Yeasts; Approved Standard—Third Edition (Clinical and Laboratory Standards Institute, Wayne, PA,2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We thank S. Umlauf, P. Gareiss and J. Merkel at Yale Center for Molecular Discovery for their help with high-throughput screening. We are grateful to K. Blount and R. Breaker for help with setting up MIC experiments. We thank J. Sinclair and A. Schepartz for sharing their expertise on cytotoxicity experiments. We gratefully acknowledge T. Fox (Cornell University) for sharing wild-type and mtDNA intronless S. cerevisiae strains. We thank S. Woodson (Johns Hopkins Unversity) for sharing the Azo-pre-tRNA plasmid. We thank D. Chenoweth and A. DeBerardinis for helpful discussions. We are grateful to S. Herzon and R. Breaker for comments on the manuscript. We are grateful to C. Zhao for help in making Supplementary Fig. 1. A.M.P. is an Investigator, and O.F is a Research Specialist in the Howard Hughes Medical Institute. This work was supported by NIH grants RO1GM50313 to A.M.P. and R43 AI115951 to M.V.Z.
Author information
Authors and Affiliations
Contributions
A.M.P., O.F. and G.E.J. designed the study. All authors contributed to this work as follows: O.F. performed in vitro biochemical studies of splicing inhibition, carried out cytotoxicity and MIC experiments; G.E.J. designed small-molecule inhibitors and carried out organic synthesis of small molecules; R.L.A. performed in vivo splicing inhibition experiments in S. cerevisiae and C. parapsilosis; L.Y. performed organic synthesis of small molecules; M.C.V.Z. designed small-molecule inhibitors; A.M.P., O.F., G.E.J. and R.L.A. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
Yale University has filed a provisional patent application on the work developed in this manuscript.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–3
Supplementary Note
Synthetic procedures
Rights and permissions
About this article
Cite this article
Fedorova, O., Jagdmann, G.E., Adams, R.L. et al. Small molecules that target group II introns are potent antifungal agents. Nat Chem Biol 14, 1073–1078 (2018). https://doi.org/10.1038/s41589-018-0142-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0142-0
This article is cited by
-
Bioengineered materials with selective antimicrobial toxicity in biomedicine
Military Medical Research (2023)
-
Targeting RNA structures with small molecules
Nature Reviews Drug Discovery (2022)
-
Alternative Treatment Strategies for Secondary Bacterial and Fungal Infections Associated with COVID-19
Infectious Diseases and Therapy (2022)
-
Splicing modulators: on the way from nature to clinic
The Journal of Antibiotics (2021)
-
The splice is right
Nature Chemical Biology (2018)