Abstract
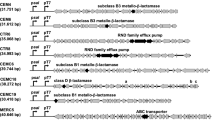
The soil microbiome can produce, resist, or degrade antibiotics and even catabolize them. While resistance genes are widely distributed in the soil, there is a dearth of knowledge concerning antibiotic catabolism. Here we describe a pathway for penicillin catabolism in four isolates. Genomic and transcriptomic sequencing revealed β-lactamase, amidase, and phenylacetic acid catabolon upregulation. Knocking out part of the phenylacetic acid catabolon or an apparent penicillin utilization operon (put) resulted in loss of penicillin catabolism in one isolate. A hydrolase from the put operon was found to degrade in vitro benzylpenicilloic acid, the β-lactamase penicillin product. To test the generality of this strategy, an Escherichia coli strain was engineered to co-express a β-lactamase and a penicillin amidase or the put operon, enabling it to grow using penicillin or benzylpenicilloic acid, respectively. Elucidation of additional pathways may allow bioremediation of antibiotic-contaminated soils and discovery of antibiotic-remodeling enzymes with industrial utility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
27 December 2018
In the version of the article originally published, the x axis of the graph in Fig. 4d was incorrectly labeled as “Retention time (min)”. It should read “Reaction time (min)”. The ‘deceased’ footnote was also formatted incorrectly when published. The footnote text itself should include the name of co-author Tara A. Gianoulis in addition to the previous link to her name in the author list through footnote number 10. The errors have been corrected in the HTML and PDF versions of the article.
References
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74, 417–433 (2010).
D’Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461 (2011).
Crofts, T. S., Gasparrini, A. J. & Dantas, G. Next-generation approaches to understand and combat the antibiotic resistome. Nat. Rev. Microbiol. 15, 422–434 (2017).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Aust, M.-O. et al. Distribution of sulfamethazine, chlortetracycline and tylosin in manure and soil of Canadian feedlots after subtherapeutic use in cattle. Environ. Pollut. 156, 1243–1251 (2008).
Wright, P. M., Seiple, I. B. & Myers, A. G. The evolving role of chemical synthesis in antibacterial drug discovery. Angew. Chem. Int. Ed. Engl. 53, 8840–8869 (2014).
Pramer, D. & Starkey, R. L. Decomposition of streptomycin. Science 113, 127 (1951).
Kameda, Y., Kimura, Y., Toyoura, E. & Omori, T. A method for isolating bacteria capable of producing 6-aminopenicillanic acid from benzylpenillin. Nature 191, 1122–1123 (1961).
Abd-El-Malek, Y., Monib, M. & Hazem, A. Chloramphenicol, a simultaneous carbon and nitrogen source for a Streptomyces sp. from Egyptain soil. Nature 189, 775–776 (1961).
Johnsen, J. Utilization of benzylpenicillin as carbon, nitrogen and energy source by a Pseudomonas fluorescens strain. Arch. Microbiol. 115, 271–275 (1977).
Beckman, W. & Lessie, T. G. Response of Pseudomonas cepacia to β-Lactam antibiotics: utilization of penicillin G as the carbon source. J. Bacteriol. 140, 1126–1128 (1979).
Johnsen, J. Presence of β-lactamase and penicillin acylase in a Pseudomonas sp. utilizing benzylpenicillin as a carbon source. J. Gen. Appl. Microbiol. 27, 499–503 (1981).
Wang, P. et al. Characterization and mechanism analysis of penicillin G biodegradation with Klebsiella pneumoniae Z1 isolated from waste penicillin bacterial residue. J. Ind. Eng. Chem. 27, 50–58 (2015).
Barnhill, A. E., Weeks, K. E., Xiong, N., Day, T. A. & Carlson, S. A. Identification of multiresistant Salmonella isolates capable of subsisting on antibiotics. Appl. Environ. Microbiol. 76, 2678–2680 (2010).
Dantas, G., Sommer, M. O. A., Oluwasegun, R. D. & Church, G. M. Bacteria subsisting on antibiotics. Science 320, 100–103 (2008).
Bello González, Tde.J., Zuidema, T., Bor, G., Smidt, H. & van Passel, M. W. Study of the aminoglycoside subsistence phenotype of bacteria residing in the gut of humans and zoo animals. Front. Microbiol. 6, 1550 (2016).
Xin, Z. et al. Isolation, identification and characterization of human intestinal bacteria with the ability to utilize chloramphenicol as the sole source of carbon and energy. FEMS Microbiol. Ecol. 82, 703–712 (2012).
Topp, E. et al. Accelerated biodegradation of veterinary antibiotics in agricultural soil following long-term exposure, and isolation of a sulfamethazine-degrading sp. J. Environ. Qual. 42, 173–178 (2013).
Tappe, W. et al. Degradation of sulfadiazine by Microbacterium lacus strain SDZm4, isolated from lysimeters previously manured with slurry from sulfadiazine-medicated pigs. Appl. Environ. Microbiol. 79, 2572–2577 (2013).
Walsh, F., Amyes, S. G. B. & Duffy, B. Challenging the concept of bacteria subsisting on antibiotics. Int. J. Antimicrob. Agents 41, 558–563 (2013).
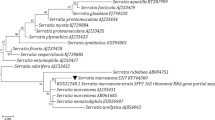
Crofts, T. S. et al. Draft genome sequences of three β-lactam-catabolizing soil Proteobacteria. Genome Announc. 5, e00653–17 (2017).
Teufel, R. et al. Bacterial phenylalanine and phenylacetate catabolic pathway revealed. Proc. Natl Acad. Sci. USA 107, 14390–14395 (2010).
Bush, K. & Jacoby, G. A. Updated functional classification of β-lactamases. Antimicrob. Agents Chemother. 54, 969–976 (2010).
Blair, J. M. A., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Ghebre-Sellassie, I., Hem, S. L. & Knevel, A. M. Epimerization of benzylpenicilloic acid in alkaline media. J. Pharm. Sci. 73, 125–128 (1984).
Hmelo, L. R. et al. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat. Protoc. 10, 1820–1841 (2015).
Valle, F., Balbás, P., Merino, E. & Bolivar, F. The role of penicillin amidases in nature and in industry. Trends Biochem. Sci. 16, 36–40 (1991).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Svedas, V., Guranda, D., van Langen, L., van Rantwijk, F. & Sheldon, R. Kinetic study of penicillin acylase from Alcaligenes faecalis. FEBS Lett. 417, 414–418 (1997).
Hammond, P. M., Price, C. P. & Scawen, M. D. Purification and properties of aryl acylamidase from Pseudomonas fluorescens ATCC 39004. Eur. J. Biochem. 132, 651–655 (1983).
Szewczuk, A., Siewiński, M. & Słowińska, R. Colorimetric assay of penicillin amidase activity using phenylacetyl-aminobenzoic acid as substrate. Anal. Biochem. 103, 166–169 (1980).
Oinonen, C. & Rouvinen, J. Structural comparison of Ntn-hydrolases. Protein Sci. 9, 2329–2337 (2000).
McVey, C. E., Walsh, M. A., Dodson, G. G., Wilson, K. S. & Brannigan, J. A. Crystal structures of penicillin acylase enzyme-substrate complexes: structural insights into the catalytic mechanism. J. Mol. Biol. 313, 139–150 (2001).
McDonough, M. A., Klei, H. E. & Kelly, J. A. Crystal structure of penicillin G acylase from the Bro1 mutant strain of Providencia rettgeri. Protein Sci. 8, 1971–1981 (1999).
Janes, L. E., Löwendahl, A. C. & Kazlauskas, R. J. Quantitative screening of hydrolase libraries using pH indicators: identifying active and enantioselective hydrolases. Chemistry 4, 2324–2331 (1998).
Batchelor, F. R., Chain, E. B., Hardy, T. L., Mansford, K. R. & Rolinson, G. N. 6-Aminopenicillanic acid. III. Isolation and purification. Proc. R. Soc. London. Ser. B, Biol. Sci. 154, 498–508 (1961).
Margolin, A. L., Svedas, V. K. & Berezin, I. V. Substrate specificity of penicillin amidase from E. coli. Biochim. Biophys. Acta 616, 283–289 (1980).
Alkema, W. B., Floris, R. & Janssen, D. B. The use of chromogenic reference substrates for the kinetic analysis of penicillin acylases. Anal. Biochem. 275, 47–53 (1999).
Henke, E. & Bornscheuer, U. T. Fluorophoric assay for the high-throughput determination of amidase activity. Anal. Chem. 75, 255–260 (2003).
Ferrández, A. et al. Catabolism of phenylacetic acid in Escherichia coli. Characterization of a new aerobic hybrid pathway. J. Biol. Chem. 273, 25974–25986 (1998).
Schumacher, G., Sizmann, D., Haug, H., Buckel, P. & Böck, A. Penicillin acylase from E. coli: unique gene-protein relation. Nucleic Acids Res. 14, 5713–5727 (1986).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Larsson, D. G. J., de Pedro, C. & Paxeus, N. Effluent from drug manufactures contains extremely high levels of pharmaceuticals. J. Hazard. Mater. 148, 751–755 (2007).
Pehrsson, E. C. et al. Interconnected microbiomes and resistomes in low-income human habitats. Nature 533, 212–216 (2016).
Rietsch, A., Vallet-Gely, I., Dove, S. L. & Mekalanos, J. J. ExsE, a secreted regulator of type III secretion genes in Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 102, 8006–8011 (2005).
Yoneda, A., Wittmann, B. J., King, J. D., Blankenship, R. E. & Dantas, G. Transcriptomic analysis illuminates genes involved in chlorophyll synthesis after nitrogen starvation in Acaryochloris sp. CCMEE 5410. Photosynth. Res. 129, 171–182 (2016).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Habegger, L. et al. RSEQtools: a modular framework to analyze RNA-Seq data using compact, anonymized data summaries. Bioinformatics 27, 281–283 (2011).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Huang, Y., Niu, B., Gao, Y., Fu, L. & Li, W. CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26, 680–682 (2010).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Gasteiger, E. et al. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31, 3784–3788 (2003).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Acknowledgements
This work is supported in part by awards to G.D. through the Edward Mallinckrodt, Jr. Foundation (Scholar Award), and from the NIH Director’s New Innovator Award (http://commonfund.nih.gov/newinnovator/), the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK: http://www.niddk.nih.gov/), the National Institute of General Medical Sciences (NIGMS: http://www.nigms.nih.gov/), and the National Institute of Allergy and Infectious Diseases (NIAID: https://www.niaid.nih.gov/) of the National Institutes of Health (NIH) under award numbers DP2DK098089, R01GM099538, and R01AI123394, respectively. T.S.C. received support from a National Institute of Diabetes and Digestive and Kidney Diseases Training Grant through award number T32 DK077653 (P.I. Tarr, Principal Investigator) and a National Institute of Child Health and Development Training Grant through award number T32 HD049305 (K.H. Moley, Principal Investigator). K.J.F. received support from the NHGRI Genome Analysis Training Program (T32 HG000045), the NIGMS Cellular and Molecular Biology Training Program (T32 GM007067), and the NSF as a graduate research fellow (award number DGE-1143954). M.K.G. received support as a Mr. and Mrs. Spencer T. Olin Fellow at Washington University and from the NSF as a graduate research fellow (DGE-1143954). Sequencing through the US Army Edgewood Chemical Biological Center was supported in part through funding provided by the Transformational Medical Technologies Initiative of the Defense Threat Reduction Agency, US Department of Defense. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies. We are thankful to J. Hoisington-Lopez in the Center for Genome Sciences and Systems Biology at Washington University in St. Louis School of Medicine for Illumina sequencing support, T. Wencewicz and B. Evans for their useful discussions regarding biochemistry and LC–MS and members of the Dantas lab for general helpful discussions regarding the manuscript.
Author information
Authors and Affiliations
Contributions
T.S.C., A.S., T.A.G., M.O.A.S., and G.D. conceived of experiments and design of work. T.S.C., B.W., A.S., and T.A.G. performed in vitro, microbial, and transcriptomic experiments. L.A.J., S.M.B., C.N.R., E.W.S., and H.S.G. sequenced strain genomes. T.S.C., A.S., T.A.G., K.J.F, and M.K.G. provided analyses. Article drafting was performed by T.S.C. with critical revision performed by T.S.C., B.W., A.S., K.J.F, M.K.G., M.O.A.S., and G.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4, Supplementary Figures 1–8
Supplementary Dataset 1
RNA-seq count dataset
Rights and permissions
About this article
Cite this article
Crofts, T.S., Wang, B., Spivak, A. et al. Shared strategies for β-lactam catabolism in the soil microbiome. Nat Chem Biol 14, 556–564 (2018). https://doi.org/10.1038/s41589-018-0052-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0052-1
This article is cited by
-
Sustainable remediation and redevelopment of brownfield sites
Nature Reviews Earth & Environment (2023)
-
Genome-resolved insight into the reservoir of antibiotic resistance genes in aquatic microbial community
Scientific Reports (2022)
-
Medicines as an emergent contaminant: the review of microbial biodegration potential
Folia Microbiologica (2022)
-
Assessment of the efficiency of synergistic photocatalysis on penicillin G biodegradation by whole cell Paracoccus sp
Journal of Biological Engineering (2021)
-
Enzyme-catalyzed biodegradation of penicillin fermentation residues by β-lactamase OtLac from Ochrobactrum tritici
Microbial Cell Factories (2021)