Abstract
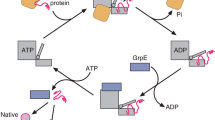
During and after protein translation, molecular chaperones require ATP hydrolysis to favor the native folding of their substrates and, under stress, to avoid aggregation and revert misfolding. Why do some chaperones need ATP, and what are the consequences of the energy contributed by the ATPase cycle? Here, we used biochemical assays and physical modeling to show that the bacterial chaperones GroEL (Hsp60) and DnaK (Hsp70) both use part of the energy from ATP hydrolysis to restore the native state of their substrates, even under denaturing conditions in which the native state is thermodynamically unstable. Consistently with thermodynamics, upon exhaustion of ATP, the metastable native chaperone products spontaneously revert to their equilibrium non-native states. In the presence of ATPase chaperones, some proteins may thus behave as open ATP-driven, nonequilibrium systems whose fate is only partially determined by equilibrium thermodynamics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Anfinsen, C. B., Haber, E., Sela, M. & White, F. H. Jr. The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain. Proc. Natl. Acad. Sci. USA 47, 1309–1314 (1961).
Onuchic, J. N. & Wolynes, P. G. Theory of protein folding. Curr. Opin. Struct. Biol. 14, 70–75 (2004).
Guo, Z. Y. & Thirumalai, D. Kinetics of protein-folding: nucleation mechanism, time scales, and pathways. Biopolymers 36, 83–102 (1995).
Knowles, T. P. J., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Finka, A., Mattoo, R. U. H. & Goloubinoff, P. Experimental milestones in the discovery of molecular chaperones as polypeptide unfolding enzymes. Annu. Rev. Biochem. 85, 715–742 (2016).
Gat-Yablonski, G. et al. Quantitative proteomics of rat livers shows that unrestricted feeding is stressful for proteostasis with implications on life span. Aging (Albany NY) 8, 1735–1758 (2016).
Goloubinoff, P., Christeller, J. T., Gatenby, A. A. & Lorimer, G. H. Reconstitution of active dimeric ribulose bisphosphate carboxylase from an unfoleded state depends on two chaperonin proteins and Mg-ATP. Nature 342, 884–889 (1989).
Jakob, U., Gaestel, M., Engel, K. & Buchner, J. Small heat shock proteins are molecular chaperones. J. Biol. Chem. 268, 1517–1520 (1993).
Koldewey, P., Stull, F., Horowitz, S., Martin, R. & Bardwell, J. C. A. Forces driving chaperone action. Cell 166, 369–379 (2016).
Frank, G. A. et al. Out-of-equilibrium conformational cycling of GroEL under saturating ATP concentrations. Proc. Natl. Acad. Sci. USA 107, 6270–6274 (2010).
De Los Rios, P. & Barducci, A. Hsp70 chaperones are non-equilibrium machines that achieve ultra-affinity by energy consumption. eLife 3, e02218 (2014).
Barducci, A. & De Los Rios, P. Non-equilibrium conformational dynamics in the function of molecular chaperones. Curr. Opin. Struct. Biol. 30, 161–169 (2015).
Goloubinoff, P., Gatenby, A. A. & Lorimer, G. H. GroE heat-shock proteins promote assembly of foreign prokaryotic ribulose bisphosphate carboxylase oligomers in Escherichia coli. Nature 337, 44–47 (1989).
Hayer-Hartl, M., Bracher, A. & Hartl, F. U. The GroEL-GroES chaperonin machine: a nano-cage for protein folding. Trends Biochem. Sci. 41, 62–76 (2016).
Hartman, D. J., Surin, B. P., Dixon, N. E., Hoogenraad, N. J. & Høj, P. B. Substoichiometric amounts of the molecular chaperones GroEL and GroES prevent thermal denaturation and aggregation of mammalian mitochondrial malate dehydrogenase in vitro. Proc. Natl. Acad. Sci. USA 90, 2276–2280 (1993).
Dobson, C. M. & Karplus, M. The fundamentals of protein folding: bringing together theory and experiment. Curr. Opin. Struct. Biol. 9, 92–101 (1999).
Peralta, D., Hartman, D. J., Hoogenraad, N. J. & Høj, P. B. Generation of a stable folding intermediate which can be rescued by the chaperonins GroEL and GroES. FEBS Lett. 339, 45–49 (1994).
Buchner, J. et al. GroE facilitates refolding of citrate synthase by suppressing aggregation. Biochemistry 30, 1586–1591 (1991).
Moran, U., Phillips, R. & Milo, R. SnapShot: key numbers in biology. Cell 141, 1262–1262.e1 (2010).
Priya, S. et al. GroEL and CCT are catalytic unfoldases mediating out-of-cage polypeptide refolding without ATP. Proc. Natl. Acad. Sci. USA 110, 7199–7204 (2013).
Horst, R. et al. Direct NMR observation of a substrate protein bound to the chaperonin GroEL. Proc. Natl. Acad. Sci. USA 102, 12748–12753 (2005).
Lin, Z., Madan, D. & Rye, H. S. GroEL stimulates protein folding through forced unfolding. Nat. Struct. Mol. Biol. 15, 303–311 (2008).
Sharma, S. et al. Monitoring protein conformation along the pathway of chaperonin-assisted folding. Cell 133, 142–153 (2008).
Libich, D. S., Tugarinov, V. & Clore, G. M. Intrinsic unfoldase/foldase activity of the chaperonin GroEL directly demonstrated using multinuclear relaxation-based NMR. Proc. Natl. Acad. Sci. USA 112, 8817–8823 (2015).
Weaver, J. et al. GroEL actively stimulates folding of the endogenous substrate protein PepQ. Nat. Commun. 8, 15934 (2017).
van der Vaart, A., Ma, J. & Karplus, M. The unfolding action of GroEL on a protein substrate. Biophys. J. 87, 562–573 (2004).
Stan, G., Lorimer, G. H., Thirumalai, D. & Brooks, B. R. Coupling between allosteric transitions in GroEL and assisted folding of a substrate protein. Proc. Natl. Acad. Sci. USA 104, 8803–8808 (2007).
Sharma, S. K., De Los Rios, P., Christen, P., Lustig, A. & Goloubinoff, P. The kinetic parameters and energy cost of the Hsp70 chaperone as a polypeptide unfoldase. Nat. Chem. Biol. 6, 914–920 (2010).
Kellner, R. et al. Single-molecule spectroscopy reveals chaperone-mediated expansion of substrate protein. Proc. Natl. Acad. Sci. USA 111, 13355–13360 (2014).
Lee, J. H. et al. Heterogeneous binding of the SH3 client protein to the DnaK molecular chaperone. Proc. Natl. Acad. Sci. USA 112, E4206–E4215 (2015).
Sekhar, A., Rosenzweig, R., Bouvignies, G. & Kay, L. E. Mapping the conformation of a client protein through the Hsp70 functional cycle. Proc. Natl. Acad. Sci. USA 112, 10395–10400 (2015).
Tehver, R. & Thirumalai, D. Kinetic model for the coupling between allosteric transitions in GroEL and substrate protein folding and aggregation. J. Mol. Biol. 377, 1279–1295 (2008).
Santra, M., Farrell, D. W. & Dill, K. A. Bacterial proteostasis balances energy and chaperone utilization efficiently. Proc. Natl. Acad. Sci. USA 114, E2654–E2661 (2017).
Priya, S., Sharma, S. K. & Goloubinoff, P. Molecular chaperones as enzymes that catalytically unfold misfolded polypeptides. FEBS Lett. 587, 1981–1987 (2013).
Yifrach, O. & Horovitz, A. Nested cooperativity in the ATPase activity of the oligomeric chaperonin GroEL. Biochemistry 34, 5303–5308 (1995).
Corrales, F. J. & Fersht, A. R. Toward a mechanism for GroEL.GroES chaperone activity: an ATPase-gated and -pulsed folding and annealing cage. Proc. Natl. Acad. Sci. USA 93, 4509–4512 (1996).
Todd, M. J., Lorimer, G. H. & Thirumalai, D. Chaperonin-facilitated protein folding: optimization of rate and yield by an iterative annealing mechanism. Proc. Natl. Acad. Sci. USA 93, 4030–4035 (1996).
Doyle, S. M. & Wickner, S. Hsp104 and ClpB: protein disaggregating machines. Trends Biochem. Sci. 34, 40–48 (2009).
Haslberger, T. et al. M domains couple the ClpB threading motor with the DnaK chaperone activity. Mol. Cell 25, 247–260 (2007).
Langer, T. et al. Successive action of DnaK, DnaJ and GroEL along the pathway of chaperone-mediated protein folding. Nature 356, 683–689 (1992).
Buchberger, A., Schröder, H., Hesterkamp, T., Schönfeld, H. J. & Bukau, B. Substrate shuttling between the DnaK and GroEL systems indicates a chaperone network promoting protein folding. J. Mol. Biol. 261, 328–333 (1996).
Sharma, S. K., De Los Rios, P. & Goloubinoff, P. Probing the different chaperone activities of the bacterial HSP70-HSP40 system using a thermolabile luciferase substrate. Proteins 79, 1991–1998 (2011).
Rajkowitsch, L. & Schroeder, R. Dissecting RNA chaperone activity. RNA 13, 2053–2060 (2007).
Quan, S. et al. Genetic selection designed to stabilize proteins uncovers a chaperone called Spy. Nat. Struct. Mol. Biol. 18, 262–269 (2011).
Lindquist, S. Protein folding sculpting evolutionary change. Cold Spring Harb. Symp. Quant. Biol. 74, 103–108 (2009).
Tokuriki, N. & Tawfik, D. S. Chaperonin overexpression promotes genetic variation and enzyme evolution. Nature 459, 668–673 (2009).
Bhaskaran, H. & Russell, R. Kinetic redistribution of native and misfolded RNAs by a DEAD-box chaperone. Nature 449, 1014–1018 (2007).
Yang, Q., Fairman, M. E. & Jankowsky, E. DEAD-box-protein-assisted RNA structure conversion towards and against thermodynamic equilibrium values. J. Mol. Biol. 368, 1087–1100 (2007).
Bershtein, S., Mu, W., Serohijos, A. W. R., Zhou, J. & Shakhnovich, E. I. Protein quality control acts on folding intermediates to shape the effects of mutations on organismal fitness. Mol. Cell 49, 133–144 (2013).
Schrödinger, E. What is life? (Cambridge University Press, Cambridge, 1945).
Ranson, N. A., Dunster, N. J., Burston, S. G. & Clarke, A. R. Chaperonins can catalyse the reversal of early aggregation steps when a protein misfolds. J. Mol. Biol. 250, 581–586 (1995).
Török, Z. et al. Evidence for a lipochaperonin: association of active protein-folding GroESL oligomers with lipids can stabilize membranes under heat shock conditions. Proc. Natl. Acad. Sci. USA 94, 2192–2197 (1997).
Gur, E. et al. The Escherichia coli DjlA and CbpA proteins can substitute for DnaJ in DnaK-mediated protein disaggregation. J. Bacteriol. 186, 7236–7242 (2004).
Woo, K. M., Kim, K. I., Goldberg, A. L., Ha, D. B. & Chung, C. H. The heat-shock protein ClpB in Escherichia coli is a protein-activated ATPase. J. Biol. Chem. 267, 20429–20434 (1992).
Diamant, S., Azem, A., Weiss, C. & Goloubinoff, P. Increased efficiency of GroE-assisted protein folding by manganese ions. J. Biol. Chem. 270, 28387–28391 (1995).
Finka, A. & Goloubinoff, P. Proteomic data from human cell cultures refine mechanisms of chaperone-mediated protein homeostasis. Cell Stress Chaperon. 18, 591–605 (2013).
Todd, M. J. & Lorimer, G. H. Stability of the asymmetric Escherichia coli chaperonin complex: guanidine chloride causes rapid dissociation. J. Biol. Chem. 270, 5388–5394 (1995).
Acknowledgements
We thank summer-project students P. Inns and I. Koog for setting up the citrate synthase enzymatic assay and performing key preliminary and control chaperone assays. This project was financed in part by Swiss National Science Foundation grants 140512/1 and 31003A_156948, and Swiss State Secretariat for Education Research and Innovation grant C15.0042 to P.G. and B.F., and by Swiss National Science Foundation grant 200020_163042 to P.D.L.R. and A.S.S. A.B. acknowledges support from the French Agence Nationale de la Recherche (ANR), under grant ANR-14-ACHN-0016. We thank H.-J. Schönfeld (F. Hoffmann-La Roche) for providing GrpE.
Author information
Authors and Affiliations
Contributions
P.G., A.B. and P.D.L.R. conceived the study. P.G. and B.F. performed the experiments. P.D.L.R., A.S.S. and A.B. developed the model. P.D.L.R. and A.S.S. performed the calculations. All authors contributed to the writing of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Note
Rights and permissions
About this article
Cite this article
Goloubinoff, P., Sassi, A.S., Fauvet, B. et al. Chaperones convert the energy from ATP into the nonequilibrium stabilization of native proteins. Nat Chem Biol 14, 388–395 (2018). https://doi.org/10.1038/s41589-018-0013-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0013-8
This article is cited by
-
Design of an Arabidopsis thaliana reporter line to detect heat-sensing and signaling mutants
Plant Methods (2023)
-
Mechanisms and pathology of protein misfolding and aggregation
Nature Reviews Molecular Cell Biology (2023)
-
A fluorescent multi-domain protein reveals the unfolding mechanism of Hsp70
Nature Chemical Biology (2023)
-
Maize heat shock proteins—prospection, validation, categorization and in silico analysis of the different ZmHSP families
Stress Biology (2023)
-
Dissipation-driven selection of states in non-equilibrium chemical networks
Communications Chemistry (2021)