Abstract
Meta-analysis of whole genome sequencing/whole exome sequencing (WGS/WES) studies provides an attractive solution to the problem of collecting large sample sizes for discovering rare variants associated with complex phenotypes. Existing rare variant meta-analysis approaches are not scalable to biobank-scale WGS data. Here we present MetaSTAAR, a powerful and resource-efficient rare variant meta-analysis framework for large-scale WGS/WES studies. MetaSTAAR accounts for relatedness and population structure, can analyze both quantitative and dichotomous traits and boosts the power of rare variant tests by incorporating multiple variant functional annotations. Through meta-analysis of four lipid traits in 30,138 ancestrally diverse samples from 14 studies of the Trans Omics for Precision Medicine (TOPMed) Program, we show that MetaSTAAR performs rare variant meta-analysis at scale and produces results comparable to using pooled data. Additionally, we identified several conditionally significant rare variant associations with lipid traits. We further demonstrate that MetaSTAAR is scalable to biobank-scale cohorts through meta-analysis of TOPMed WGS data and UK Biobank WES data of ~200,000 samples.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
This paper used the TOPMed Freeze 5 WGS data and lipids phenotype data. Genotype and phenotype data are both available in database of Genotypes and Phenotypes. The TOPMed WGS data were from the following fourteen study cohorts (accession numbers provided in parentheses): Framingham Heart Study (phs000974.v1.p1); Old Order Amish (phs000956.v1.p1); Jackson Heart Study (phs000964.v1.p1); and Multi-Ethnic Study of Atherosclerosis (phs001416.v1.p1); Atherosclerosis Risk in Communities Study (phs001211); Cleveland Family Study (phs000954); Cardiovascular Health Study (phs001368); Diabetes Heart Study (phs001412); Genetic Study of Atherosclerosis Risk (phs001218); Genetic Epidemiology Network of Arteriopathy (phs001345); Genetics of Lipid Lowering Drugs and Diet Network (phs001359); San Antonio Family Heart Study (phs001215); Genome-wide Association Study of Adiposity in Samoans (phs000972) and Women’s Health Initiative (phs001237). The sample sizes, ancestry and phenotype summary statistics of these cohorts are given in Supplementary Table 1. The UK Biobank analyses were conducted using the UK Biobank resource under application 52008.
The functional annotation data are publicly available and were downloaded from the following links: GRCh38 CADD v1.4 (https://cadd.gs.washington.edu/download); ANNOVAR dbNSFP v3.3a (https://annovar.openbioinformatics.org/en/latest/user-guide/download); LINSIGHT (https://github.com/CshlSiepelLab/LINSIGHT); FATHMM-XF (http://fathmm.biocompute.org.uk/fathmm-xf); FANTOM5 CAGE (https://fantom.gsc.riken.jp/5/data); GeneCards (https://www.genecards.org; v4.7 for hg38); and Umap/Bismap (https://bismap.hoffmanlab.org; ‘before March 2020’ version). In addition, recombination rate and nucleotide diversity were obtained from Gazal et al. ref. 51). The whole genome individual functional annotation data was assembled from a variety of sources and the computed annotation PCs are available at the Functional Annotation of Variant-Online Resource (FAVOR) site (https://favor.genohub.org) and the FAVOR database (https://doi.org/10.7910/DVN/1VGTJI)52. The tissue-specific functional annotations were downloaded from ENCODE (https://www.encodeproject.org/report/?type=Experiment).
Code availability
MetaSTAAR is implemented as an open-source R package available at https://github.com/xihaoli/MetaSTAAR and https://content.sph.harvard.edu/xlin/software.html. Data analysis was performed in R (3.5.1). STAAR v0.9.6 and MetaSTAAR v0.9.6 were used in simulation and real data analysis and implemented as open-source R packages available at https://github.com/xihaoli/STAAR (ref. 53) and https://github.com/xihaoli/MetaSTAAR (ref. 54). The scripts used to generate the results have been archived on Zenodo using https://doi.org/10.5281/zenodo.6668274 (ref. 55). RareMetal v4.15.1 (https://github.com/statgen/raremetal) and GMMAT v1.3.2 (https://cran.r-project.org/web/packages/GMMAT/index.html) were used for comparison. The assembled functional annotation data were downloaded from FAVOR using Wget (https://www.gnu.org/software/wget/wget.html).
References
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Van Hout, C. V. et al. Exome sequencing and characterization of 49,960 individuals in the UK Biobank. Nature 586, 749–756 (2020).
Szustakowski, J. D. et al. Advancing human genetics research and drug discovery through exome sequencing of the UK Biobank. Nat. Genet. 53, 942–948 (2021).
Hindy, G. et al. Rare coding variants in 35 genes associate with circulating lipid levels—a multi-ancestry analysis of 170,000 exomes. Am. J. Hum. Genet. 109, 81–96 (2022).
Flannick, J. et al. Exome sequencing of 20,791 cases of type 2 diabetes and 24,440 controls. Nature 570, 71–76 (2019).
Jurgens, S. J. et al. Analysis of rare genetic variation underlying cardiometabolic diseases and traits among 200,000 individuals in the UK Biobank. Nat. Genet. 54, 240–250 (2022).
Wainschtein, P. et al. Assessing the contribution of rare variants to complex trait heritability from whole-genome sequence data. Nat. Genet. 54, 263–273 (2022).
Lee, S., Abecasis, Gonçalo, R., Boehnke, M. & Lin, X. Rare-variant association analysis: study designs and statistical tests. Am. J. Hum. Genet. 95, 5–23 (2014).
Li, B. & Leal, S. M. Methods for detecting associations with rare variants for common diseases: application to analysis of sequence data. Am. J. Hum. Genet. 83, 311–321 (2008).
Madsen, B. E. & Browning, S. R. A groupwise association test for rare mutations using a weighted sum statistic. PLoS Genet. 5, e1000384 (2009).
Morris, A. P. & Zeggini, E. An evaluation of statistical approaches to rare variant analysis in genetic association studies. Genet. Epidemiol. 34, 188–193 (2010).
Wu, M. C. et al. Rare-variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011).
Liu, Y. et al. ACAT: a fast and powerful P value combination method for rare-variant analysis in sequencing studies. Am. J. Hum. Genet. 104, 410–421 (2019).
McCarthy, M. I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat. Rev. Genet. 9, 356–369 (2008).
Evangelou, E. & Ioannidis, J. P. A. Meta-analysis methods for genome-wide association studies and beyond. Nat. Rev. Genet. 14, 379–389 (2013).
Buniello, et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Lin, D. Y. & Zeng, D. Meta-analysis of genome-wide association studies: no efficiency gain in using individual participant data. Genet. Epidemiol. 34, 60–66 (2010).
Lin, D. Y. & Zeng, D. On the relative efficiency of using summary statistics versus individual-level data in meta-analysis. Biometrika 97, 321–332 (2010).
Liu, D. J. et al. Meta-analysis of gene-level tests for rare variant association. Nat. Genet. 46, 200–204 (2014).
Feng, S., Liu, D., Zhan, X., Wing, M. K. & Abecasis, G. R. RAREMETAL: fast and powerful meta-analysis for rare variants. Bioinformatics 30, 2828–2829 (2014).
Lee, S., Teslovich, Tanya, M., Boehnke, M. & Lin, X. General framework for meta-analysis of rare variants in sequencing association studies. Am. J. Hum. Genet. 93, 42–53 (2013).
Hu, Y.-J. et al. Meta-analysis of gene-level associations for rare variants based on single-variant statistics. Am. J. Hum. Genet. 93, 236–248 (2013).
Yang, J., Chen, S. & Abecasis, G., IAMDGC. Improved score statistics for meta-analysis in single-variant and gene-level association studies. Genet. Epidemiol. 42, 333–343 (2018).
Chen, H. et al. Efficient variant set mixed model association tests for continuous and binary traits in large-scale whole-genome sequencing studies. Am. J. Hum. Genet. 104, 260–274 (2019).
Chen, M.-H., Pitsillides, A. & Yang, Q. An evaluation of approaches for rare variant association analyses of binary traits in related samples. Sci. Rep. 11, 3145 (2021).
Li, X. et al. Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale. Nat. Genet. 52, 969–983 (2020).
Gogarten, S. M. et al. Genetic association testing using the GENESIS R/Bioconductor package. Bioinformatics 35, 5346–5348 (2019).
Chen, H. et al. Control for population structure and relatedness for binary traits in genetic association studies via logistic mixed models. Am. J. Hum. Genet. 98, 653–666 (2016).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Natarajan, P. et al. Deep-coverage whole genome sequences and blood lipids among 16,324 individuals. Nat. Commun. 9, 3391 (2018).
Stilp, A. M. et al. A system for phenotype harmonization in the national heart, lung, and blood institute Trans-omics for Precision Medicine (TOPMed) program. Am. J. Epidemiol. 190, 1977–1992 (2021).
Forrest, A. R. et al. A promoter-level mammalian expression atlas. Nature 507, 462 (2014).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Fishilevich, S. et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database 2017, bax028 (2017).
Li, Z. et al. A framework for detecting noncoding rare-variant associations of large-scale whole-genome sequencing studies. Nat. Methods (2022). https://doi.org/10.1038/s41592-022-01640-x
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Huang, Y.-F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Nat. Genet. 49, 618–624 (2017).
Rogers, M. F. et al. FATHMM-XF: accurate prediction of pathogenic point mutations via extended features. Bioinformatics 34, 511–513 (2017).
Dong, C. et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 24, 2125–2137 (2014).
Zhou, H. et al. FAVOR: functional annotation of variants online resource and annotator for variation across the human genome. Nucleic Acids Res. gkac966, https://doi.org/10.1093/nar/gkac966 (2022).
Schaffner, S. F. et al. Calibrating a coalescent simulation of human genome sequence variation. Genome Res. 15, 1576–1583 (2005).
Lee, P. H. et al. Principles and methods of in-silico prioritization of non-coding regulatory variants. Hum. Genet. 137, 15–30 (2018).
Morrison, A. C. et al. Practical approaches for whole-genome sequence analysis of heart-and blood-related traits. Am. J. Hum. Genet. 100, 205–215 (2017).
Li, Z. et al. Dynamic scan procedure for detecting rare-variant association regions in whole-genome sequencing studies. Am. J. Hum. Genet. 104, 802–814 (2019).
The All of Us Research Program Investigators The “All of Us” Research Program. N Engl J Med 381, 668–676 (2019).
Klarin, D. et al. Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat. Genet. 50, 1514–1523 (2018).
Breslow, N. E. & Clayton, D. G. Approximate inference in generalized linear mixed models. J. Am. Stat. Assoc. 88, 9–25 (1993).
Jiang, L. et al. A resource-efficient tool for mixed model association analysis of large-scale data. Nat. Genet. 51, 1749–1755 (2019).
Jiang, L., Zheng, Z., Fang, H. & Yang, J. A generalized linear mixed model association tool for biobank-scale data. Nat. Genet. 53, 1616–1621 (2021).
Quick, C. et al. A versatile toolkit for molecular QTL mapping and meta-analysis at scale. Preprint at bioRxiv https://doi.org/10.1101/2020.12.18.423490 (2020).
Gazal, S. et al. Linkage disequilibrium–dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Zhou, H., Arapoglou, T., Li, X., Li, Z. & Lin, X. FAVOR Essential Database. V1 Edition (Harvard Dataverse, 2022).
Li, X., Li, Z. & Chen, H. xihaoli/STAAR: STAAR_v0.9.6. Version 0.9.6 https://doi.org/10.5281/zenodo.6960622 (2022)
Li, X. & Li, Z. xihaoli/MetaSTAAR: MetaSTAAR_v0.9.6. Version 0.9.6 https://doi.org/10.5281/zenodo.6960606 (2022)
Li, X., Li, Z. & Lin, X. MetaSTAAR. Version 1 https://doi.org/10.5281/zenodo.6668274 (2022)
Acknowledgements
This work was supported by grants R35-CA197449, U19-CA203654, R01-HL113338, U01-HG012064 and U01-HG009088 (X. Lin), NHLBI BioData Catalyst Fellowship (Z.L.), R01-HL142711 and R01-HL127564 (P.N. and G.M.P.), 75N92020D00001, HHSN268201500003I, N01-HC-95159, 75N92020D00005, N01-HC-95160, 75N92020D00002, N01-HC-95161, 75N92020D00003, N01-HC-95162, 75N92020D00006, N01-HC-95163, 75N92020D00004, N01-HC-95164, 75N92020D00007, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169, UL1-TR-000040, UL1-TR-001079, UL1-TR-001420, UL1-TR001881, DK063491, R01-HL071051, R01-HL071205, R01-HL071250, R01-HL071251, R01-HL071258, R01-HL071259 and UL1-RR033176 (J.I.R. and X.G.), R35-HL135824 (C.J.W.), U01-HL72518, HL087698, HL49762, HL59684, HL58625, HL071025, HL112064, NR0224103 and M01-RR000052 (to the Johns Hopkins General Clinical Research Center), NO1-HC-25195, HHSN268201500001I, 75N92019D00031 and R01-HL092577-06S1 (R.S.V. and L.A.C.), the Evans Medical Foundation and the Jay and Louis Coffman Endowment from the Department of Medicine, Boston University School of Medicine (R.S.V.), HHSN268201800001I and U01-HL137162 (K.M.R.), R01-HL093093 and R01-HL133040 (S.T.M.), R35-HL135818, R01-HL113338 and HL436801 (S.R.), KL2TR002490 (L.M.R.), R01-HL92301, R01-HL67348, R01-NS058700, R01-AR48797 and R01-AG058921 (N.D.P. and D.W.B.), R01-DK071891 (N.D.P., B.I.F. and D.W.B.), M01-RR07122 and F32-HL085989 (to the General Clinical Research Center of the Wake Forest University School of Medicine), the American Diabetes Association, P60-AG10484 (to the Claude Pepper Older Americans Independence Center of Wake Forest University Health Sciences), U01-HL137181 (J.R.O.), HHSN268201600018C, HHSN268201600001C, HHSN268201600002C, HHSN268201600003C and HHSN268201600004C (C.K.), R01-HL113323, U01-DK085524, R01-HL045522, R01-MH078143, R01-MH078111 and R01-MH083824 (H.H.H.G., R.D., J.E.C. and J.B.), 18CDA34110116 from American Heart Association (P.S.d.V.), HHSN268201800010I, HHSN268201800011I, HHSN268201800012I, HHSN268201800013I, HHSN268201800014I and HHSN268201800015I (A.C.), R01-HL153805, R03-HL154284 (B.E.C.), HHSN268201700001I, HHSN268201700002I, HHSN268201700003I, HHSN268201700005I and HHSN268201700004I (E.B.), U01-HL072524, R01-HL104135-04S1, U01-HL054472, U01-HL054473, U01-HL054495, U01-HL054509 and R01-HL055673-18S1 (D.K.A.). Molecular data for the Trans Omics in Precision Medicine (TOPMed) program was supported by the National Heart, Lung and Blood Institute (NHLBI). Core support including centralized genomic read mapping and genotype calling, along with variant quality metrics and filtering were provided by the TOPMed Informatics Research Center (3R01HL-117626-02S1; contract HHSN268201800002I). Core support including phenotype harmonization, data management, sample-identity QC and general program coordination was provided by the TOPMed Data Coordinating Center (R01HL-120393; U01HL-120393; contract HHSN268201800001I). We gratefully acknowledge the studies and participants who provided biological samples and data for TOPMed and UK Biobank. The full study-specific acknowledgements and NHLBI BioData Catalyst acknowledgement are detailed in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
X. Li, C.Q., G.M.P., Z.L. and X.L. designed the experiments. X.Li, C.Q., Z.L. and X. Lin performed the experiments. X. Li, C.Q., H.Z., S.M.G., Y.L., H.C., M.S.S., R.S., R.D., D.K.A., L.F.B., J.C.B., J.B., E.B., D.W.B., J.A.B., B.E.C., A.C., L.A.C., J.E.C., P.S.d.V., R.D., B.I.F., H.H.H.G., X.G., J.H., R.R.K., C.K., B.G.K., L.A.L., A.M., L.W.M., S.T.M., B.D.M., M.E.M., A.C.M., T.N., J.R.O., N.D.P., P.A.P., B.M.P., L.M.R., S.R., A.P.R., M.S.R., K.M.R., S.S.R., C.M.S, J.A.S., K.D.T., R.S.V., C.J.W., J.G.W., L.R.Y., W.Z., J.I.R., P.N., G.M.P., Z.L. and X. Lin acquired, analyzed or interpreted data. G.M.P., P.N. and the NHLBI TOPMed Lipids Working Group provided administrative, technical or material support. X. Li, Z.L. and X. Lin drafted the manuscript and revised it according to suggestions by the co-authors. All authors critically reviewed the manuscript, suggested revisions as needed and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
S.M.G. is now an employee of Regeneron Genetics Center. For B.D.M., The Amish Research Program receives partial support from Regeneron Pharmaceuticals. M.E.M. reports grant from Regeneron Pharmaceutical unrelated to the present work. B.M.P. serves on the Steering Committee of the Yale Open Data Access Project funded by Johnson & Johnson. L.M.R. is a consultant for the TOPMed Administrative Coordinating Center (through Westat). For S.R., Jazz Pharma, Eli Lilly, Apnimed, unrelated to the present work. The spouse of C.J.W. works at Regeneron Pharmaceuticals. P.N. reports investigator-initiated grants from Amgen, Apple, AstraZeneca, Boston Scientific and Novartis, personal fees from Apple, AstraZeneca, Blackstone Life Sciences, Foresite Labs, Novartis, Roche/Genentech, is a cofounder of TenSixteen Bio, is a shareholder of geneXwell and TenSixteen Bio, and spousal employment at Vertex, all unrelated to the present work. X. Lin is a consultant of AbbVie Pharmaceuticals and Verily Life Sciences. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Christoph Lippert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quantile-quantile plots for gene-centric unconditional meta-analysis of lipid traits LDL-C, HDL-C, TG and TC using TOPMed WGS data (n = 30,138).
MetaSTAAR-O is a two-sided test. Different symbols represent the MetaSTAAR-O P values of different functional categories of individual genes (putative loss-of-function, missense, synonymous, promoter and enhancer). The promoter and enhancer of a gene are the promoter and the GeneHancer region that overlap with CAGE sites for a given gene, respectively (Methods). Four lipid traits were analyzed using MetaSTAAR-O: LDL-C, low-density lipoprotein cholesterol; HDL-C, high-density lipoprotein cholesterol; TG, triglycerides; and TC, total cholesterol.
Extended Data Fig. 2 Manhattan plots for gene-centric unconditional meta-analysis of lipid traits LDL-C, HDL-C, TG and TC using TOPMed WGS data (n = 30,138).
The horizontal line indicates the genome-wide MetaSTAAR-O P value threshold of 5.00 × 10−7. The significant threshold is defined by multiple comparisons using the Bonferroni correction (0.05/(20,000×5) = 5.00 × 10−7). MetaSTAAR-O is a two-sided test. Different symbols represent the MetaSTAAR-O P values of different functional categories of individual genes (putative loss-of-function, missense, synonymous, promoter and enhancer). The promoter and enhancer of a gene are the promoter and the GeneHancer region that overlap with CAGE sites for a given gene, respectively (Methods). Four lipid traits were analyzed using MetaSTAAR-O: LDL-C, low-density lipoprotein cholesterol; HDL-C, high-density lipoprotein cholesterol; TG, triglycerides; TC, total cholesterol.
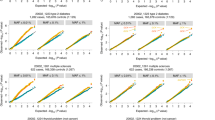
Extended Data Fig. 3 Scatterplots comparing gene-centric unconditional meta-analysis P values from MetaSTAAR-O with STAAR-O from the joint analysis of pooled individual-level data (STAAR-O-Pooled) of lipid traits LDL-C, HDL-C, TG and TC using TOPMed WGS data (n = 30,138).
Each dot represents a functional category of a gene with x-axis label being the −log10(P) of STAAR-O-Pooled and y-axis label being the −log10(P) of MetaSTAAR-O (n = 30,138). The horizontal and vertical lines indicate the genome-wide P value threshold of 5.00 × 10−7. The significant threshold is defined by multiple comparisons using the Bonferroni correction (0.05/(20,000×5) = 5.00 × 10−7). Both MetaSTAAR and STAAR are two-sided tests. LDL-C, low-density lipoprotein cholesterol; HDL-C, high-density lipoprotein cholesterol; TG, triglycerides; TC, total cholesterol.
Extended Data Fig. 4 Scatterplot of P values comparing MetaSTAAR-O to Burden-MS, SKAT-MS and ACAT-V-MS (MS is short for MetaSTAAR) for quantitative and dichotomous traits when 15% of rare variants are causal variants.
In each simulation replicate, a 2-kb region was randomly selected as the signal region. Within each signal region, variants were randomly generated to be causal based on a multiple logistic model and on average there were 15% causal variants in the signal region. The effect sizes of causal variants were βj = c0|log10MAFj|. For quantitative traits, c0 = 0.07; for dichotomous traits, c0 = 0.11. All causal variants had positive effect sizes. Power was estimated as the proportion of the P values less than α = 10−7 based on 104 replicates. Burden-MS, SKAT-MS, ACAT-V-MS and MetaSTAAR-O are two-sided tests. Five studies were included in meta-analysis, each with a sample size of 10,000.
Supplementary information
Supplementary information
Supplementary Figs. 1–4 and Supplementary note
Supplementary Table 1
Supplementary Tables 1–9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Quick, C., Zhou, H. et al. Powerful, scalable and resource-efficient meta-analysis of rare variant associations in large whole genome sequencing studies. Nat Genet 55, 154–164 (2023). https://doi.org/10.1038/s41588-022-01225-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01225-6
This article is cited by
-
Enhancing prediction accuracy of coronary artery disease through machine learning-driven genomic variant selection
Journal of Translational Medicine (2024)
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
Accurate and efficient estimation of local heritability using summary statistics and the linkage disequilibrium matrix
Nature Communications (2023)