Abstract
Adult kidney organoids have been described as strictly tubular epithelia and termed tubuloids. While the cellular origin of tubuloids has remained elusive, here we report that they originate from a distinct CD24+ epithelial subpopulation. Long-term-cultured CD24+ cell-derived tubuloids represent a functional human kidney tubule. We show that kidney tubuloids can be used to model the most common inherited kidney disease, namely autosomal dominant polycystic kidney disease (ADPKD), reconstituting the phenotypic hallmark of this disease with cyst formation. Single-cell RNA sequencing of CRISPR–Cas9 gene-edited PKD1- and PKD2-knockout tubuloids and human ADPKD and control tissue shows similarities in upregulation of disease-driving genes. Furthermore, in a proof of concept, we demonstrate that tolvaptan, the only approved drug for ADPKD, has a significant effect on cyst size in tubuloids but no effect on a pluripotent stem cell-derived model. Thus, tubuloids are derived from a tubular epithelial subpopulation and represent an advanced system for ADPKD disease modeling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Processed gene expression values from the scRNA-seq are available at https://doi.org/10.6084/m9.figshare.11786238. Processed data from the ATAC-seq analysis are available for peer review under the private link at https://figshare.com/s/728705bc42446275044d in FigShare. These data have a reserved DOI (https://doi.org/10.6084/m9.figshare.11848281). All raw data are available in the controlled EGA access repository EGAS00001006551. Source data are provided with this paper.
Code availability
The code for reproducible analysis of the single-cell data is available at https://github.com/saezlab/Xu_tubuloid and https://github.com/KramannLab/kidney_human_organoids in GitHub repositories. The computer code to reproduce the analysis of ATAC-seq data is available at https://github.com/ATA82/ATAC_Seq_Xu in GitHub. The specific software and methods used for the analysis are described in the README file of the repository.
References
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015).
Morizane, R. et al. Nephron organoids derived from human pluripotent stem cells model kidney development and injury. Nat. Biotechnol. 33, 1193–1200 (2015).
Taguchi, A. et al. Redefining the in vivo origin of metanephric nephron progenitors enables generation of complex kidney structures from pluripotent stem cells. Cell Stem Cell 14, 53–67 (2014).
Freedman, B. S. et al. Modelling kidney disease with CRISPR-mutant kidney organoids derived from human pluripotent epiblast spheroids. Nat. Commun. 6, 8715 (2015).
Wu, H. et al. Comparative analysis and refinement of human PSC-derived kidney organoid differentiation with single-cell transcriptomics. Cell Stem Cell 23, 869–881 (2018).
Schutgens, F. et al. Tubuloids derived from human adult kidney and urine for personalized disease modeling. Nat. Biotechnol. 37, 303–313 (2019).
Lazzeri, E. et al. Regenerative potential of embryonic renal multipotent progenitors in acute renal failure. J. Am. Soc. Nephrol. 18, 3128–3138 (2007).
Romagnani, P. & Remuzzi, G. CD133+ renal stem cells always co-express CD24 in adult human kidney tissue. Stem Cell Res. 12, 828–829 (2014).
Romagnani, P. Family portrait. Am. J. Pathol. 178, 490–493 (2011).
Sagrinati, C. et al. Isolation and characterization of multipotent progenitor cells from the Bowman’s capsule of adult human kidneys. J. Am. Soc. Nephrol. 17, 2443–2456 (2006).
Smeets, B. et al. Proximal tubular cells contain a phenotypically distinct, scattered cell population involved in tubular regeneration. J. Pathol. 229, 645–659 (2013).
Teslaa, T. & Teitell, M. A. Pluripotent stem cell energy metabolism: an update. EMBO J. 34, 138–153 (2015).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Barker, N. et al. Lgr5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
El-Achkar, T. M. et al. A multimodal and integrated approach to interrogate human kidney biopsies with rigor and reproducibility: guidelines from the Kidney Precision Medicine Project. Physiol. Genomics 53, 1–11 (2021).
Cornec-Le Gall, E., Alam, A. & Perrone, R. D. Autosomal dominant polycystic kidney disease. Lancet 393, 919–935 (2019).
Guo, T. et al. Harnessing accurate non-homologous end joining for efficient precise deletion in CRISPR/Cas9-mediated genome editing. Genome Biol. 19, 170 (2018).
Cruz, N. M. et al. Organoid cystogenesis reveals a critical role of microenvironment in human polycystic kidney disease. Nat. Mater. 16, 1112–1119 (2017).
Czerniecki, S. M. et al. High-throughput screening enhances kidney organoid differentiation from human pluripotent stem cells and enables automated multidimensional phenotyping. Cell Stem Cell 22, 929–940 (2018).
Margaria, J. P., Campa, C. C., de Santis, M. C., Hirsch, E. & Franco, I. The PI3K/Akt/mTOR pathway in polycystic kidney disease: a complex interaction with polycystins and primary cilium. Cell. Signal. 66, 109468 (2020).
Qin, S. et al. Failure to ubiquitinate c-Met leads to hyperactivation of mTOR signaling in a mouse model of autosomal dominant polycystic kidney disease. J. Clin. Invest. 120, 3617–3628 (2010).
Yamaguchi, T. et al. Cyclic AMP activates B-Raf and ERK in cyst epithelial cells from autosomal-dominant polycystic kidneys. Kidney Int. 63, 1983–1994 (2003).
Cai, J. et al. A RhoA–YAP–c-Myc signaling axis promotes the development of polycystic kidney disease. Genes Dev. 32, 781–793 (2018).
Nagai, J. S., Leimkühler, N. B., Schaub, M. T., Schneider, R. K. & Costa, I. G. CrossTalkeR: analysis and visualization of ligand–receptor networks. Bioinformatics 37, 4263–4265 (2021).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
Baert, L. Hereditary polycystic kidney disease (adult form): a microdissection study of two cases at an early stage of the disease. Kidney Int. 13, 519–525 (1978).
Wilson, P. D. Polycystic kidney disease. N. Engl. J. Med. 350, 151–164 (2004).
Schäfer, K. et al. Characterization of the Han:SPRD rat model for hereditary polycystic kidney disease. Kidney Int. 46, 134–152 (1994).
Nagao, S. et al. Renal activation of extracellular signal-regulated kinase in rats with autosomal-dominant polycystic kidney disease. Kidney Int. 63, 427–437 (2003).
Islam, M. R. et al. Retinoic acid-dependent activation of the polycystic kidney disease-1 (PKD1) promoter. Am. J. Physiol. Ren. Physiol. 295, F1845–F1854 (2008).
Pritchard, L. et al. A human PKD1 transgene generates functional polycystin-1 in mice and is associated with a cystic phenotype. Hum. Mol. Genet. 9, 2617–2627 (2000).
Puri, S. et al. Ets factors regulate the polycystic kidney disease-1 promoter. Biochem. Biophys. Res. Commun. 342, 1005–1013 (2006).
Yu, A. S. L., Kanzawa, S. A., Usorov, A., Lantinga-van Leeuwen, I. S. & Peters, D. J. M. Tight junction composition is altered in the epithelium of polycystic kidneys. J. Pathol. 216, 120–128 (2008).
Russo, R. J. et al. Impaired formation of desmosomal junctions in ADPKD epithelia. Histochem. Cell Biol. 124, 487–497 (2005).
Rogers, K. K., Jou, T.-S., Guo, W. & Lipschutz, J. H. The Rho family of small GTPases is involved in epithelial cystogenesis and tubulogenesis. Kidney Int. 63, 1632–1644 (2003).
Wang, X., Wu, Y., Ward, C. J., Harris, P. C. & Torres, V. E. Vasopressin directly regulates cyst growth in polycystic kidney disease. J. Am. Soc. Nephrol. 19, 102–108 (2008).
Shimizu, T. et al. A novel ADPKD model using kidney organoids derived from disease-specific human iPSCs. Biochem. Biophys. Res. Commun. 529, 1186–1194 (2020).
Torres, V. E. et al. Effective treatment of an orthologous model of autosomal dominant polycystic kidney disease. Nat. Med. 10, 363–364 (2004).
Gattone, V. H. 2nd, Maser, R. L., Tian, C., Rosenberg, J. M. & Branden, M. G. Developmental expression of urine concentration-associated genes and their altered expression in murine infantile-type polycystic kidney disease. Dev. Genet. 24, 309–318 (1999).
Lawlor, K. T. et al. Cellular extrusion bioprinting improves kidney organoid reproducibility and conformation. Nat. Mater. 20, 260–271 (2021).
Subramanian, A. et al. Single cell census of human kidney organoids shows reproducibility and diminished off-target cells after transplantation. Nat. Commun. 10, 5462 (2019).
Uchimura, K., Wu, H., Yoshimura, Y. & Humphreys, B. D. Human pluripotent stem cell-derived kidney organoids with improved collecting duct maturation and injury modeling. Cell Rep. 33, 108514 (2020).
Kabadi, A. M., Ousterout, D. G., Hilton, I. B. & Gersbach, C. A. Multiplex CRISPR/Cas9-based genome engineering from a single lentiviral vector. Nucleic Acids Res. 42, e147 (2014).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Fujii, M., Matano, M., Nanki, K. & Sato, T. Efficient genetic engineering of human intestinal organoids using electroporation. Nat. Protoc. 10, 1474–1485 (2015).
Kuppe, C. et al. Decoding myofibroblast origins in human kidney fibrosis. Nature 589, 281–286 (2021).
Costales-Carrera, A. et al. Plocabulin displays strong cytotoxic activity in a personalized colon cancer patient-derived 3D organoid assay. Mar. Drugs 17, 648 (2019).
Daley, T. & Smith, A. D. Predicting the molecular complexity of sequencing libraries. Nat. Methods 10, 325–327 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and samtools. Bioinformatics 25, 2078–2079 (2009).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Lawrence, M., Gentleman, R. & Carey, V. rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Ibrahim, M. M. & Kramann, R. genesorteR: feature ranking in clustered single cell data. Preprint at bioRxiv https://doi.org/10.1101/676379 (2019).
Tung, P.-Y. et al. Batch effects and the effective design of single-cell gene expression studies. Sci. Rep. 7, 39921 (2017).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Chen, Y., Lun, A. T. L. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res. 5, 1438 (2016).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2016).
Liberzon, A. et al. Molecular Signatures Database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Holland, C. H. et al. Robustness and applicability of transcription factor and pathway analysis tools on single-cell RNA-seq data. Genome Biol. 21, 36 (2020).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Dimitrov, D. et al. Comparison of methods and resources for cell–cell communication inference from single-cell RNA-seq data. Nat. Commun. 13, 3224 (2022).
Acknowledgements
We thank M. Newen and S. Ernst of the Interdisziplinäres Zentrum für Klinische Forschung (IZKF) Aachen confocal microscopy facility and M. Vogt of the IZKF Aachen two-photon imaging facility for assistance with imaging and image analysis. We further thank the IZKF Aachen Genomics Facility for its support. This work was supported by grants from the German Research Foundation (DFG: SFBTRR219 to R.K. and J.F., CRU344-4288578857858, CRU5011-445703531), the European Research Council (ERC-StG CureCKDHeart 677448, ERC-Con TargetCKD 101043403), the Else Kroener Fresenius Foundation, the Dutch Kidney Foundation (DKF), TASKFORCE EP1805, NWO VIDI 09150172010072 and the Leducq Foundation (all to R.K.). This work was also supported by the BMBF eMed Consortia Fibromap (to I.C., R.K.S. and R.K.) and grants from the DFG (project IDs 322900939, 454024652, 432698239 and 445703531 to P.B.). Further funding came from the START-Program of the Faculty of Medicine, RWTH Aachen (to K.H. and T.S.), the German Society of Internal Medicine (to T.S.) and the DKF (14A3D10 to B.S.). We thank I. Gomez for his technical assistance. Work in the Freedman laboratory was supported by NIH (awards R01DK117914, U01DK127553, UG3TR002158 and UG3TR003288) and the Lara Nowak Macklin Fund.
Author information
Authors and Affiliations
Contributions
Y.X. designed and carried out experiments, analyzed results and wrote the manuscript; C.K. designed and performed some experiments, analyzed the data and contributed to writing; J.P.P., S.H., F.P., J.N. and H.K. analyzed the scRNA-seq data; J.K. and J.S. recruited patients, collected clinical data and reviewed the manuscript; A.T.A. analyzed the ATAC-seq data; B.F. performed the iPSC experiment and contributed to data interpretation and writing; M.H., T.S., K.H., J.J. and S.M. performed some experiments; M.L. and J.M. helped with the Seahorse experiments; E.M.B. and P.B. performed the electron microscopy experiments; C.K. and I.K. performed genotyping of the patient specimens; M.R. and C.B. consented patients and provided living donor biopsies; B.F. performed the iPSC experiments; R.H. and E.B. performed the sequencing and alignment of the scRNA-seq; F.P., J.J., I.K., J.S.R., Z.L., T.B.H., A.K., T.S., M.E.J.R., I.C., M.R., C.B., T.S., M.R., C.v.R., B.S., E.J.H., J.G., R.K.S. and J.F. contributed to interpretation of the results. All authors reviewed the manuscript. R.K. planned the project and experiments, analyzed and interpreted the data, wrote the manuscript and obtained funding. All authors revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.K. reports unrelated funding from Travere Therapeutics, Galapagos and Novo Nordisk and unrelated honoraria from Bayer, Pfizer, Grünenthal and Novo Nordisk. S.H. reports unrelated funding from Novo Nordisk. J.S.R. reports unrelated funding from GSK and Sanofi and unrelated fees from Travere Therapeutics and Astex Therapeutics. All other authors have nothing to disclose.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Metabolism and DNA accessibility in CD24+ cells as compared to CD13+ cells.
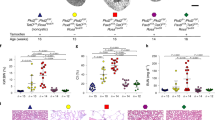
a-b, Cellular glycolysis and proton efflux rate (PER) of CD24+ cells as compared to CD13+ cells. Statistical analysis in n = 12 mean ± s.e.m. (a-b), 2-way ANOVA post-hoc Bonferroni, non-significant for post 2-deoxy-D-glucose (2-DG, a glucose analog that inhibits glycolysis) in CD24+ cell vs CD13+ cells, P = 0.1834 (b); Basal glycolysis and comp glycolysis (compensatory glycolysis) in CD24+ cells vs CD13+ cells, ****P < 0.0001. c, ATP-linked vs basal OCR between CD24+ cells and CD13+ cells in n = 12 mean ± s.e.m., P = 0.7041, non-significant using a two-tailed (also called two-sided) unpaired t-test with Welch’s correction. d, Spare respiratory capacity (SRC) of CD24+ cells compared to CD13+ cells, n = 12 mean ± s.e.m., ****P < 0.0001 by a two-tailed unpaired t-test with Welch’s correction. e-f, Extracellular acidification rates (EACR) of CD24+ cells compared to CD13+ cells during anaerobic glycolysis. n = 12 mean ± s.e.m.(e-f), 2-way-ANOVA with post-hoc Bonferroni, nonsignificant, P = 0.053 for Post 2-DG in CD24+ cells vs CD13+ cells, ****P < 0.0001 for Basal, Compensatory glycolysis and 2-DG in CD24+ cells vs CD13+ cells(f). g-h, Extracellular acidification rates (EACR) in CD24+ cells compared to CD13+ cells during aerobic mitochondrial respiration. n = 12 mean ± s.d. (g-h), ****P < 0.0001 for Basal, Oligomycin-ATP-linked, FCCP, and non-Mito-Rot/AA in CD24+ cells vs CD13+ cells by 2-way-ANOVA with post-hoc Bonferroni(h). i, FACS gating for sorting of CD13+ or CD24+ cells. j, Scaled heat map showing the chromatin accessibility of top 50 most accessible promoter regions in CD24+ cells or CD13+ cells. k, Top 25 upregulated and downregulated GO terms in CD24+ cells vs CD13+ cells based on the genome-wide measured chromatin accessibility by ATAC-seq.
Extended Data Fig. 2 Wnt3a/RSPO1 activated growth and expansion of CD24+ cells.
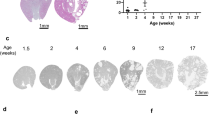
a-d, Representative images and quantification of cell-cluster sizes of Wnt3a conditioned medium (CM) treated CD24+ cells compared to CD13+ cells at 72 h. Statistical analysis in n = 7 mean ± s.e.m., **P = 0.0045 by two-tailed t-test with Welch’s correction(c); CD24+ cells formed large cell clusters before Wnt3a was removed at day 4 (d). Scale bars 100 µm-a and b; 50 µm-d. e-f, Expansion of CD24+ cells into early tubuloids at day 6, which subjected to 4-phase protocol under Wnt3a/RSPO1 conditioned medium (e); CD13+ cells that subjected to 4-phase protocol under Wnt3a/RSPO1 conditioned medium, did not produce tubuloids and presented as loose cell-aggregates with vacuole formation (f). Scale bar 50 µm: e and f. g, Representative images of CD 24+ cell derived organoid formation at day 7, 12, 17, 21 as well as organoid growth maintenance at day 30 and day 69, which were subjected to the 1-phase protocol published by Schutgens et al. Scale bar, 200 µm: day 21, 30, 69; 100 µm: day 0, 7, 17; 50 µm: day 12. h, Representative image of CD24+ cell derived organoid formation at day 21 that was subjected to the 4-phase protocol. Scare bar 50µm. i-j, Schematic (I) and representative images of CD24+ cells subjected to a tubuloid protocol without WNT3a/RSPO1(j). In the schematic, a FGF2, FGF10 stability factor (SF), heparin was used in this protocol, stability factor (SF) represents heparin (i). Scale bars 200 µm-m at day 30, 73; 100 µm- m at day 0, day 14; 50 µm-m at day 5, 8, 12 (j). k, Organoid formation rate comparison between CD24+ cell subjected to the 4-phase protocol without Wnt3a/RSPO1 (i-j) and the original 4-phase protocol, n = 4 mean ± s.e.m with two-tailed unpaired t-test with Welch’s correction, ***P = 0.0003. Arrows mark borders of tubuloids. Wnt-Wnt3a; RS- RSPO1. For details on statistics and reproducibility, see Methods.
Extended Data Fig. 3 CD24+ cells are the cellular origin of adult kidney tubuloids.
a, FACS gating for sorted CD24+/CD13+, CD24+/CD13-, CD24-/CD13+ and CD24-/CD13- cells. b-e, Representative bright field images of sorted CD24+/CD13+(b); CD24+/CD13-(c); CD24-/CD13+(d) and CD24-/CD13- cells (e) subjected to the 4-phase organoid formation and expansion protocol. Note that only the populations, that contain CD24+ cells were able to form tubuloids (b, c). Arrows mark cells, borders of nephrospheres or tubuloids. Scale bar 100 µm-b-e: sorted cells, 72h Wnt3a, Day 1 in 3D; 50 µm-b-e: Day 4 in 3D, Day 21, Day 30. f-l, Representative DIC (differential interference contrast) images exhibited growth and expansion of tubuloids from CD24+/CD13+ cells(f-j) and CD24+/CD13- cells(k-l). h represents a zoom from g. Red square marks tubuli in a tubuloid at day 52. Arrows mark cells, borders of tubuli or tubuloids. Scale bars, 200 µm: j;100 µm: g, l at -day 61; 50 µm: f, h, i, k at day 40, l at day 52, l -zoom in. m, Comparison of tubuloid formation rates in CD24+/CD13+, CD24+/CD13-, CD24-/CD13+ and CD24-/CD13- cells. Statistical analysis was performed by 2-way ANOVA with post-hoc Bonferroni correction in n = 4 mean ± s.e.m., non-significant, P > 0.9999 for CD24-/CD13- vs CD24-/CD13+; **P = 0.0077 for CD24+/CD13- vs CD24-/CD13+ or CD24-/CD13-; ***P = 0.0002 for CD24+/CD13+ vs CD24+/CD13-; ****P < 0.0001 for CD24+/CD13+ vs CD24-/CD13+ or CD24-/CD13- cells. Arrows mark cells, borders of cells or tubuloids.
Extended Data Fig. 4 Gene-editing resulted in a precise deletion within the human PKD1 or PKD2 gene.
a-d, Representative images of a horizontal agarose gel electrophoresis for PCR amplified DNA fragments of the target region using sorted GFP+ cells from PKD1−/−, PKD2−/− and control (lentibbCas9v2eGFP) tubuloids at day 7 of transduced tubuloids in 3D (day 10 after transduction, a and c), quantification of the knockout efficiency from paired gRNA gene editing of PKD1 gene (b) and paired gRNA gene editing of PKD2 gene (d) by measurement of the relative band intensity. Statistical analysis was performed by two-tailed unpaired t-test with n = 3 mean ± s.d., **P = 0.0041 in PKD1−/− tubuloids vs control (b); and n = 3 mean ± s.d., **P = 0.0013 in PKD2−/− tubuloids vs control (d). Uncropped DNA gel images in Supplementary Fig. 18. e-f, Sanger sequencing of the paired CRISPR/Cas9 targeting region revealed an accurate deletion of 265bp in the PKD1 gene (e) and 165bp in the PKD2 gene (f) at day 7 of the transduced kidney tubuloids in 3D (10 days after transduction). g, Quantification of the percentage of GFP+ cysts in control (EV) and PKD1−/−, PKD2−/− tubuloids at 10, 15 and 20 days after transduction that were treated with blebbistatin (Blebb), then transferred into suspension culture for 72 h. Statistical analysis using 2-way-ANOVA post-hoc Bonferroni in n = 4 mean ± s.e.m., P > 0.999, no significant in GFP+ cyst rate of EV tubuloids between day 20 and 15 or 10 each other; P = 0.0516 in GFP+ cyst rate of PKD1−/− +Blebb for day 20 vs 15; ***P = 0.0007 in GFP+ cyst rate of PKD1−/− +Blebb for day 15 vs 10; ****P < 0.0001 in GFP+ cyst rate of PKD1−/− +Blebb for day 20 vs 10; **P = 0.0020 in GFP+ cyst rate of PKD2−/− +Blebb for day 15 vs 10; ****P <0.0001 in GFP+ cyst rate of PKD2−/− +Blebb for day 20 vs 15 or 10. ###P <0.001 in PKD1−/− or PKD2−/− +Blebb for day 15 and 10; P = 0.402 in PKD1-/- and #P = 0.048 in PKD2−/− for day 20, by 2-way-ANOVA with post-hoc Bonferroni vs EV+Blebb on the same day.
Extended Data Fig. 5 Single nuclei ADPKD transcriptomic data.
a, Scaled gene expression of the top 3 differentially over-expressed genes (sorted by log2-fold-changes) in each cluster. b, Individual UMAP embeddings of nuclei from each of the 5 human kidney specimens. c, Number of genes, cell count and percentage of reads mapped to mitochondrial genes for each human sample. Box-whisker plots. d, Contribution of each sample toward the composition of the major cell types identified in human samples. e, Cell-type distribution was shown per condition and percentage of cells. f, Top 5 upregulated genes in human ADPKD vs donor biopsies. g, Heatmap of pathway activity using PROGENY comparing the integrated ADPKD kidney tissue data to the donor biopsies, indicating increased pathway activity in several cystic epithelial clusters as compared to their nephron counterparts in the healthy tissue. ***P < 0.001; **P < 0.01; *P < 0.05. P: Adjusted P-value by Benjamini & Hochberg method for multiple testing, from a nominal P-value obtained from an enrichment test with 10,000 permutations. h, Heatmap of gene-set enrichment of hallmark pathways of ADPKD comparing the integrated human ADPKD tissue data to the control data from donor biopsies. ***P < 0.001; **P <0.01; *P <0.05. P: Adjusted P-value by Benjamini & Hochberg method, two-tailed, from a nominal P value obtained from an enrichment test with 1,000 permutations.
Extended Data Fig. 6 Pathway analysis of human ADPKD and control kidney tissue.
a, Gene set enrichment with PID pathway database in human ADPKD kidney tissue compared to controls. b, Gene set enrichment with GO Molecular function terms in human ADPKD kidney tissue compared to controls. c, Gene set enrichment with KEGG pathways in human ADPKD kidney tissue compared to controls. d, Gene set enrichment with BIOCARTA gene sets in human ADPKD kidney tissue compared to controls.
Extended Data Fig. 7 Receptor ligand interaction analysis in human ADPKD kidney tissue snRNA-seq data compared to healthy human kidney tissue.
a, Cell-cell interaction based on CrossTalkeR for all cell-clusters in human ADPKD kidney tissue snRNA-seq data as compared to control tissue (donor biopsies) snRNA-seq data. Color indicates weight of interaction. Orange and cyan color signifies cell-cell interaction enrichment in ADPKD and controls, respectively. b, PageRank log ratio for cell-type ranking based on the in-coming and out-going interactions. Orange and cyan color signifies cell-cell interaction enrichment in ADPKD and controls, respectively. c, PageRank log ratio for cell-gene interaction ranking. Orange and cyan color signifies cell-cell interaction enrichment in ADPKD and controls, respectively. d, Heatmap of KEGG pathway enrichment using top 100 high-scoring dysregulated genes using the different ranking measures. e, Sankey plot summarizing top 50 out of 566 significant MET ligand-receptor interactions from major cell types. f, Sankey plot summarizing top 50 out of 291 significant ERBB4 ligand-receptor interactions from major cell types.
Extended Data Fig. 8 Subclustering analysis of selected populations in human ADPKD and control tissue snRNA-seq data.
a, Subclustering of principal cells of the collecting duct and epithelial cells of the connecting tubule (PC-CD/CNT), relative distribution of cells per subcluster in ADPKD and control tissue (donor biopsies) and top filtered marker genes of subclusters. b, Subclustering of cells from the distal convoluted tubule (DCT), relative distribution of cells per subcluster in ADPKD and control tissue (donor biopsies) and top filtered marker genes of subclusters. c, Subclustering of collecting duct intercalated cells (ICs), relative distribution of cells per subcluster in ADPKD and control tissue (donor biopsies) and top filtered marker genes of subclusters. d, Subclustering of PT epithelial cells, relative distribution of cells per subcluster in ADPKD and control tissue (donor biopsies) and top filtered marker genes of subclusters. e, Subclustering of cells from the LOH thick ascending limb (TAL), relative distribution of cells per subcluster in ADPKD and control tissue (donor biopsies) and top filtered marker genes of subclusters. f, Overrepresentation analyses with Reactome pathway database using up-regulated DE genes obtained from ADPKD vs control comparison.
Extended Data Fig. 9 Symphony mapping of tubuloid cells to human kidney snRNA-seq data.
a-e, Symphony mapping of tubuloid cell clusters from different tubuloids generated (a, early tubuloid 4-phase; b, late tubuloid 4-phase; c, tubuloid 1-phase; d, PKD1 gene edited tubuloid; e, PKD2 gene edited tubuloid) to the integrated human kidney snRNA-seq data from donor biopsies. f, Representative Immunofluorescent staining of the proximal tubular marker LTL (green) and the TAL marker SLC12A1 (white), non-ADPKD human kidney tissue. Scale bars 75 µm. g-h, Representative immunofluorescent staining of the proximal tubular marker LTL (green) and the TAL marker SLC12A1 (white) on human ADPKD tissue sections showing epithelial cysts lined by TAL marker SLC12A1+ cells (arrows). Nuclei were counterstained with DAPI. Scale bars 75 µm. For details on statistics and reproducibility, see Methods.
Extended Data Fig. 10 Gene expression in gene edited tubuloids as compared to human kidneys and AVPR2 expression in tubuloids.
a, Venn diagram of separately commonly up and downregulated genes in TAL cluster TAL_2 and PT_4 in human ADPKD kidney tissue versus healthy human kidney (donor biopsies) and the gene edited tubuloids as compared to control tubuloids. b, Commonly dysregulated pathways obtained from gene set enrichment with KEGG pathways using all differentially expressed (DE) genes in human ADPKD vs donor biopsies in PT-4 and TAL-2 cells with the cells of the tubuloids that mapped to PT-4 and TAL-2 using Symphony. c, Top selected commonly downregulated genes in human ADPKD vs donor biopsies and the cells that mapped to PT-4 and TAL-2 in PKD−/− vs Control. d, Results of the CellTiter-Glo 3D Cell Viability assay from control (EV), PKD1−/− and PKD2−/− tubuloids subjected to different doses of tolvaptan. 0.0 indicates 0.2% DMSO treated tubuloids as a control. 2-way ANOVA post-hoc Bonferroni in n = 3 mean ± s.e.m. No significant difference in cell viability for DMSO control (0.0 µM tolvaptan, 0.2% DMSO) compared to 0.1 to 20 µM tolvaptan treated EV tubuloids, PKD1−/−, PKD2−/− tubuloids, P > 0.9999; 40 µM tolvaptan vs DMSO control, P = 0.0846 for EV tubuloids, **P = 0.0019 for PKD1−/− tubuloids, and *P = 0.0215 for PKD2−/− tubuloids. e-f, Dotplots of AVPR1A and AVPR2 average scaled mRNA expression in tubuloids (late and 1-phase protocol) (e) and PKD1−/− as well as PKD2−/− (f). g, Relative mRNA expression for AVPR2 (V2R) in non-treated tubuloids as compared to tubuloids transduced with a lentivirus control (EV) and tubuloids were gene edited by lenti paired CRISPR/Cas9 for either PKD1 or PKD2. 1-way-ANOVA with post-hoc Bonferroni in n = 4 mean ± s.d., PKD1−/− tubuloids vs EV tubuloids, ****P < 0.0001; PKD2−/− tubuloids vs EV tubuloids, ***P = 0.0006; PKD1−/− tubuloids vs normal tubuloids, ****P < 0.0001; PKD2−/− tubuloids vs normal tubuloids, ***P = 0.0004; EV tubuloids vs normal tubuloids, P > 0.9999. h, Representative in-situ hybridization of AVPR2, human ADPKD2 kidney tissue section. AVPR2 mRNA was detected in cyst-lining epithelial cells. Scale bar 50 µm. i, Representative immunofluorescent staining of the LOH marker SLC12A1 and AVPR2 in collecting duct epithelial cells. Scale bars 50 µm. j, Representative immunofluorescent staining of the LOH marker SLC12A1 and AVPR2 in ADPKD cyst lining epithelial cells. Scale bars 50 µm. For details on statistics and reproducibility, see Methods.
Supplementary information
Supplementary Information
Supplementary Notes and Supplementary Figs. 1–18.
Supplementary Video 1
Brightfield time-lapse video of early CD24+ cell-derived nephrospheres at day 7 forming a lumen, magnification ×20.
Supplementary Video 2
Brightfield time-lapse video of CD24+ cells at day 9 of culture showing that the early lumina converge into tubules, magnification ×20.
Supplementary Video 3
Confocal z stack of CD24+ cell-derived kidney tubuloids at day 21, stained for LTL (far red) and CHD1 (green); magnification ×40, zoom 1.2, 26 images in 26-µm z stack.
Supplementary Video 4
Confocal z stack of a CD24+ cell-derived kidney tubuloids at day 21, stained for β-catenin (red) and CDH1 (green); magnification ×40, 36 images in 36-µm z stack.
Supplementary Video 5
Brightfield time-lapse video showing cyst formation at day 18 of PKD1−/− tubuloids after transduction, magnification ×20.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Y., Kuppe, C., Perales-Patón, J. et al. Adult human kidney organoids originate from CD24+ cells and represent an advanced model for adult polycystic kidney disease. Nat Genet 54, 1690–1701 (2022). https://doi.org/10.1038/s41588-022-01202-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01202-z
This article is cited by
-
Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
Nature Communications (2024)
-
Generation of proximal tubule-enhanced kidney organoids from human pluripotent stem cells
Nature Protocols (2023)