Abstract
Characterization of genetic regulatory variants acting on livestock gene expression is essential for interpreting the molecular mechanisms underlying traits of economic value and for increasing the rate of genetic gain through artificial selection. Here we build a Cattle Genotype–Tissue Expression atlas (CattleGTEx) as part of the pilot phase of the Farm animal GTEx (FarmGTEx) project for the research community based on 7,180 publicly available RNA-sequencing (RNA-seq) samples. We describe the transcriptomic landscape of more than 100 tissues/cell types and report hundreds of thousands of genetic associations with gene expression and alternative splicing for 23 distinct tissues. We evaluate the tissue-sharing patterns of these genetic regulatory effects, and functionally annotate them using multiomics data. Finally, we link gene expression in different tissues to 43 economically important traits using both transcriptome-wide association and colocalization analyses to decipher the molecular regulatory mechanisms underpinning such agronomic traits in cattle.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All raw data analyzed in this study are publicly available for download without restrictions from SRA (https://www.ncbi.nlm.nih.gov/sra/) and BIGD (https://bigd.big.ac.cn/bioproject/) databases. Details of RNA-seq, WGBS and WGS can be found in Supplementary Tables 1, 2 and 15, respectively. All processed data, the full summary statistics of QTL mapping are available at https://cgtex.roslin.ed.ac.uk/.
Code availability
All the computational scripts and codes for RNA-seq and DNA methylation data quantification, quality control, gene expression normalization, genotype imputation, QTL mapping, functional enrichment, TWAS and colocalization are available at both the web portal of CattleGTEx (https://cgtex.roslin.ed.ac.uk/) and the github website (https://github.com/shuliliu/cattleGTEx, https://doi.org/10.5281/zenodo.6510550)57.
References
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Hu, Z. L., Park, C. A. & Reecy, J. M. Building a livestock genetic and genomic information knowledgebase through integrative developments of Animal QTLdb and CorrDB. Nucleic Acids Res. 47, D701–D710 (2019).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Fang, L. et al. Comprehensive analyses of 723 transcriptomes enhance genetic and biological interpretations for complex traits in cattle. Genome Res. 30, 790–801 (2020).
Xiang, R. et al. Quantifying the contribution of sequence variants with regulatory and evolutionary significance to 34 bovine complex traits. Proc. Natl Acad. Sci. USA 116, 19398–19408 (2019).
Prowse-Wilkins, C. P. et al. Putative causal variants are enriched in annotated functional regions from six bovine tissues. Front. Genet. 12, 664379 (2021).
Xiang, R. et al. Genome variants associated with RNA splicing variations in bovine are extensively shared between tissues. BMC Genomics 19, 521 (2018).
Cesar, A. S. M. et al. Identification of putative regulatory regions and transcription factors associated with intramuscular fat content traits. BMC Genomics 19, 499 (2018).
Littlejohn, M. D. et al. Sequence-based association analysis reveals an MGST1 eQTL with pleiotropic effects on bovine milk composition. Sci. Rep. 6, 25376 (2016).
Hayes, B. J. & Daetwyler, H. D. 1000 Bull Genomes Project to map simple and complex genetic traits in cattle: applications and outcomes. Annu. Rev. Anim. Biosci. 7, 89–102 (2019).
Deelen, P. et al. Calling genotypes from public RNA-sequencing data enables identification of genetic variants that affect gene-expression levels. Genome Med. 7, 30 (2015).
Hombach, S. & Kretz, M. Non-coding RNAs: classification, biology and functioning. Adv. Exp. Med. Biol. 937, 3–17 (2016).
GTEx Consortium et al.Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Tenesa, A. et al. Estimation of linkage disequilibrium in a sample of the United Kingdom dairy cattle population using unphased genotypes. J. Anim. Sci. 81, 617–623 (2003).
Fang, L. et al. Functional annotation of the cattle genome through systematic discovery and characterization of chromatin states and butyrate-induced variations. BMC Biol. 17, 68 (2019).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rosen, B.D. et al. De novo assembly of the cattle reference genome with single-molecule sequencing. Gigascience 9, giaa021 (2020).
Jiang, J. et al. Functional annotation and Bayesian fine-mapping reveals candidate genes for important agronomic traits in Holstein bulls. Commun. Biol. 2, 212 (2019).
Freebern, E. et al. GWAS and fine-mapping of livability and six disease traits in Holstein cattle. BMC Genomics 21, 41 (2020).
Fang, L. et al. Genetic and epigenetic architecture of paternal origin contribute to gestation length in cattle. Commun. Biol. 2, 100 (2019).
Gao, Y. et al. Single-cell transcriptomic analyses of dairy cattle ruminal epithelial cells during weaning. Genomics 113, 2045–2055 (2021).
Kim-Hellmuth, S. et al. Cell type-specific genetic regulation of gene expression across human tissues. Science 369, eaaz8528 (2020).
Clark, E. L. et al. From FAANG to fork: application of highly annotated genomes to improve farmed animal production. Genome Biol. 21, 285 (2020).
Xiang, R. D. Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations. Nat. Commun. 12, 860 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using LeafCutter. Nat. Genet. 50, 151–158 (2018).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Castel, S. E., Mohammadi, P., Chung, W. K., Shen, Y. & Lappalainen, T. Rare variant phasing and haplotypic expression from RNA sequencing with phASER. Nat. Commun. 7, 12817 (2016).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Song, Q. et al. A reference methylome database and analysis pipeline to facilitate integrative and comparative epigenomics. PLoS ONE 8, e81148 (2013).
Liu, H. et al. Systematic identification and annotation of human methylation marks based on bisulfite sequencing methylomes reveals distinct roles of cell type-specific hypomethylation in the regulation of cell identity genes. Nucleic Acids Res. 44, 75–94 (2016).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Ongen, H., Buil, A., Brown, A. A., Dermitzakis, E. T. & Delaneau, O. Fast and efficient QTL mapper for thousands of molecular phenotypes. Bioinformatics 32, 1479–1485 (2016).
Mohammadi, P., Castel, S. E., Brown, A. A. & Lappalainen, T. Quantifying the regulatory effect size of cis-acting genetic variation using allelic fold change. Genome Res. 27, 1872–1884 (2017).
Wen, X., Lee, Y., Luca, F. & Pique-Regi, R. Efficient integrative multi-SNP association analysis via deterministic approximation of posteriors. Am. J. Hum. Genet. 98, 1114–1129 (2016).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS ONE 7, e30377 (2012).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Saha, A. & Battle, A. False positives in trans-eQTL and co-expression analyses arising from RNA-sequencing alignment errors. F1000Res. 7, 1860 (2018).
Yang, J., Zaitlen, N. A., Goddard, M. E., Visscher, P. M. & Price, A. L. Advantages and pitfalls in the application of mixed-model association methods. Nat. Genet. 46, 100–106 (2014).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat. Commun. 9, 1825 (2018).
Barbeira, A. N. et al. Integrating predicted transcriptome from multiple tissues improves association detection. PLoS Genet. 15, e1007889 (2019).
Wen, X., Pique-Regi, R. & Luca, F. Integrating molecular QTL data into genome-wide genetic association analysis: probabilistic assessment of enrichment and colocalization. PLoS Genet. 13, e1006646 (2017).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Heger, A., Webber, C., Goodson, M., Ponting, C. P. & Lunter, G. GAT: a simulation framework for testing the association of genomic intervals. Bioinformatics 29, 2046–2048 (2013).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Liu, S. et al. A multi-tissue atlas of regulatory variants in cattle. Code resource at github website. GitHub: https://github.com/shuliliu/cattleGTEx; Zenodo: https://doi.org/10.5281/zenodo.6510550
Acknowledgements
This work was supported in part by Agriculture and Food Research Initiative (AFRI) grant numbers 2016-67015-24886, 2019-67015-29321 and 2021-67015-33409 from the United States Department of Agriculture (USDA) National Institute of Food and Agriculture (NIFA) Animal Genome and Reproduction Programs, and US–Israel Binational Agricultural Research and Development (BARD) grant number US-4997-17 from the BARD Fund. L.F. was partially funded through Health Data Research UK (HDRUK) award HDR-9004 and the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No. 801215. A.T. acknowledged funding from the Biotechnology and Biological Sciences Research Council through program grants BBS/E/D/10002070 and BBS/E/D/30002275, Medical Research Council research grant MR/P015514/1 and HDRUK award HDR-9004. O.C.-X. was supported by MR/R025851/1. R.X. was supported by Australian Research Council’s Discovery Projects (DP200100499). Y. Yu. was supported by the National Science Foundation of China-Pakistan Science Foundation Joint Project (31961143009) and National Key R&D Program of China (2021YFD1200900 and 2021YFD1200903). L.M. was supported in part by AFRI grant numbers 2020-67015-31398 and 2021-67015-33409 from the NIFA. G.E.L., B.D.R. and C.P.V.T. were supported by appropriated project 8042-31000-001-00-D, ‘Enhancing Genetic Merit of Ruminants Through Improved Genome Assembly, Annotation, and Selection’ of the Agricultural Research Service (ARS) of the USDA. C.-J.L. was supported by appropriated project 8042-31310-078-00-D, ‘Improving Feed Efficiency and Environmental Sustainability of Dairy Cattle through Genomics and Novel Technologies’ of ARS-USDA. J.B.C. was supported by appropriated project 8042-31000-002-00-D, ‘Improving Dairy Animals by Increasing Accuracy of Genomic Prediction, Evaluating New Traits, and Redefining Selection Goals’ of ARS-USDA. This research used resources provided by the SCINet project of the ARS-USDA project number 0500-00093-001-00-D. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA. The USDA is an equal opportunity provider and employer. All the funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank US dairy producers for providing phenotypic, genomic and pedigree data through the Council on Dairy Cattle Breeding under ARS-USDA Material Transfer Research Agreement 58-8042-8-007. Access to 1000 Bull Genomes Project data was provided under ARS-USDA Data Transfer Agreement 15443. International genetic evaluations were calculated by the International Bull Evaluation Service (Interbull; Uppsala, Sweden).
Author information
Authors and Affiliations
Contributions
L.F., A.T. and G.E.L. conceived and designed the project. S.L., Y.G., O.C.-X., S.W., L.F., R.X., W.C., B.L., C.X., Y. Yao, Z.Y. and X.L. performed bioinformatic analyses. O.C.-X., L.F., Y. Yu, E.P.-C., K.D., K.R., C.L., A.J.C., P.N., D.R., B.D.R., C.P.V.T., P.M.V., S.Z., L.M., J.B.C., G.E.L. and A.T. contributed to the resource generation. S.L., L.F., Y.G., G.E.L. and A.T. wrote the manuscript. All authors read, edited and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Ben Hayes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Data summary of publicly available RNA-Seq data.
(a) The number of publicly available RNA-Seq samples increases rapidly over years by fitting a second order polynominal model. (b) Distribution of sequence platforms of all 8,536 RNA-Seq samples. (c) Percentage of RNA-seq with single or paired reads. (d) Distribution of numbers of clean reads across all samples. (e) Distribution of read lengths. (f) Distribution of sexes. (g) Distribution of ages (Year-old). (h) Distribution of uniquely mapping rates. (i) Distribution of major tissues and breeds/ancestries in the 7,180 high quality RNA-Seq datasets (clean read > 500,000 & mapping rate > 60%).
Extended Data Fig. 2 Gene expression and alternative splicing profiles across samples.
(a) Number of expressed genes (Transcripts per Million, TPM > 0.1) increases rapidly with the increasing number of clean reads across all 8,536 samples, reaching a plateau at 50 million reads. The black line is the smoothed curve fitted by a generalized additive model using geom_smooth function from ggplot2 (v3.3.6) in R (v3.4.1). The shaded area around the lines represents the 95% confidence interval for the fitted values (the line). (b) The percentage of unexpressed genes (TPM < 0.1 across all samples) on known chromosomes (Known) and unplaced scaffolds (Unplaced, 54.10%). (c-f) Compared to expressed genes, the unexpressed genes have shorter gene length (df = 21,921, P = 2.2 × 10−4) (c), fewer exons (df = 27,675, P = 2.5 × 10−5) (d), higher CG density (df = 21,921, P = 1.5 × 10−103) (e), and higher dN/dS ratio (df = 19,718, P = 5.4 × 10−21) (f). (g) The number of spliced introns increases rapidly with the increasing number of clean reads across samples, reaching a plateau at 100 million reads. The smoothed curve and the shaded band are obtained using the same method as in (a). (h-k) Compared to all genes, genes without spliced introns in any tissues have shorter gene length (df = 22,320, P = 2.9 × 10−18) (h), fewer exons (df = 17,690, P = 7.4 × 10−52) (i), lower expression levels (median gene expression levels across samples, df = 28,479, P = 0.35) (j), and higher dN/dS ratio (df = 19,921, P = 3.7 × 10−32) (k). All the P values above are obtained based on the two-sided Welch two sample t-test, and * indicates P < 0.05. (l) Distribution of gene types for those without spliced introns. (m) Significant terms (P < 0.05) of Gene Ontology for genes without spliced introns based on the hypergeometric test.
Extended Data Fig. 3 DNA methylation profiles of 144 WGBS samples.
(a) The percentage of covered CpGs (read depth ≥ 5× or ≥ 10×) in the entire genome increases rapidly with the increasing number of reads used for methylation extraction, approximately reaching a plateau at 200 million reads. The black solid line and dash line are the smoothed curves fitted by a generalized additive model using geom_smooth function from ggplot2 (v3.3.6) in R (v3.4.1) for read depth ≥ 5× and ≥ 10×, respectively. The shaded area around the lines represents the 95% confidence interval for the fitted values (the lines). (b) Compared to covered CpGs (Covered), the uncovered CpGs (read depth < 5× across all samples, Uncovered) tend to be located within gene deserts (df = 15,074,753, P < 2.2×10−308) and regions with higher CG density (df = 15,074,753, P < 2.2×10−308). All the P values above are obtained based on the two-sided Welch two sample t-test, and * indicates P < 0.05. (c) Distribution of uncovered CpGs (< 5×) along the entire genome.
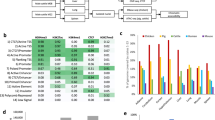
Extended Data Fig. 4 Genotyping and imputation of variants from RNA-Seq data.
(a) Distribution of numbers of SNPs directly called from RNA-Seq data across all 8,536 samples. (b) Concordance rates between genotypes (mean = 78,587, range = 47,407–113,868) called from RNA-Seq data and imputed genotypes (mean = 2.50 million, range = 1.20–2.73 million) in three tissues and those called from whole genome sequencing (WGS) data across four Holstein (HOL) animals. (c) Proportion of variants within functional categories using different imputation accuracy cutoffs. These results are derived from 109 Holstein animals with both RNA-seq and 50 K SNP array. ‘All.SNPs’ are those 31,377,923 imputed variants common in the two imputation processes (that is, the genotype imputation based on RNA-Seq SNPs and that based on SNP array). ‘imp.acc>=0.80.Aus’ are those imputed based on 50 K SNP array genotypes (Australian HOL animals) and variants with imputation accuracy DR2 > 0.80 were selected (n = 16,501,943). ‘imp.acc>=0.80.GTEx’ are those in the CattleGTEx data where the imputation was based on RNA-seq SNPs and variants with imputation accuracy DR2 > 0.80 were selected (n = 5,292,828). (d) Comparison of DR2 of SNPs imputed from SNP array (50 K) and those imputed from RNA-Seq SNPs along 1 Mb up-/down- stream of gene body. The up-/down-stream is divided into windows of 100 kb length, while the gene body region of each gene is evenly divided into 10 windows. The DR2 values of SNPs within each window are then averaged for plotting. (e) Pearson correlations of genotype counts between variants imputed from RNA-Seq SNPs and those from 50 K SNP arrays across different imputation quality cutoffs and chromosomes. The horizontal dashed line in each graph indicates the mean of correlations across chromosomes. (f) Distribution of identity by state (IBS) distance between all sample pairs. The IBS distance is calculated using PLINK v1.90 to measure the average proportion of alleles shared between samples. The sample pairs with IBS distance > 0.85 are considered as duplicated samples.
Extended Data Fig. 5 Functional annotation of tissue-specific genes and their sequence conservation in mammals.
(a) Significant Gene Ontology (GO) terms for genes with tissue-specific expression, based on the one-sided Fisher’s exact test using ClusterProfiler v3.0.4. FDR is obtained after the Benjamini-Hochberg correction for the raw P value. (b) Significant GO terms for genes with tissue-specific hypomethylated promoters (P < 0.05). (c) Significant GO terms for genes with brain-specific spliced introns (Benjamini-Hochberg corrected P (FDR) < 0.05 after correction). (d) dN/dS ratio (between cattle and humans) of orthologous genes with tissue-specific expression across tissues. The red dash line indicates median value of 0.154. (e) PhastCons scores of regions with tissue-specific hypomethylation across tissues. PhastCons scores were obtained from UCSC website and calculated on the basis of DNA sequences of 46 placental mammals. The red dash line indicates the median value of 0.1. (f) The Pearson’s correlation (r = −0.68, the two-sided Student’s t-test: P = 0.0036) between PhastCons scores of tissue-specific DNA methylation regions and dN/dS ratios of tissue-specific expressed genes across 16 common tissues. (f) has the same color key as (d).
Extended Data Fig. 6 Characterization of cis-eQTLs across tissues.
(a) Factor weight variance as a function of PEER factors computed up to 75 factors for each of 23 distinct tissues. Factor weight variances become small for most of tissues when the number of inferred hidden PEER factors reaches 10. (b) Pearson’s correlation between inferred factors and known covariates in adipose. The color in each cell denotes −log10(FDR) after the Benjamini-Hochberg correlation of P values (the two-sided Student’s t-test). Only significant correlations (FDR < 0.05) are shown in cells. (c) The proportion of cis-eQTLs with |log2(aFC)| ≤ 1 over all cis-eQTLs as a function of sample size across 23 distinct tissues. |log2(aFC)|, that is, the log2 transformed allelic fold change, which is used to measure the effect size of cis-eQTL. (d) The cis-eQTL cumulative proportion plot of |log2(aFC)| distribution across 9 tissues with variable sample sizes. The arrow indicates tissues in legend were listed from largest to smallest sample size. (e) Distribution of cis-eQTLs around TSS (1 Mb up- and down-stream) in adipose. All gene-variant pairs tested as null; ‘Significant’ indicates the top eQTLs for significant eGenes; ‘Top, Not significant’ indicates the top associated SNP for non-significant genes (non-eGenes). (f) Distribution of cis-eQTLs around the TSS (1 Mb up- and down-stream) across all 23 distinct tissues. (g) Correlation of effect sizes (fastQTL slope) of cis-eQTLs and aFC of matched loci tested by allelic specific expression (ASE) analysis in adipose (Spearman’s rho = 0.75, the two-sided Student’s t-test: P < 2.2 × 10−308) and muscle (Spearman’s rho = 0.68, the two-sided Student’s t-test: P = 2.1 × 10−162). (h) Percentage of cis-eQTLs in the combined muscle data that are replicated in multi-subspecies meta-analysis at different P-value cutoffs used for defining cis-eQTLs. The cis-eQTLs with higher significant levels are more likely to be specifically detected in the combined population. (i) Effects sizes (|log2(aFC)|) of cis-eQTLs specifically detected in combined population are significantly (the two-sided Welch two sample t-test: P = 1.25 × 10−26) smaller than those that are replicated in multi-breed meta-analysis.
Extended Data Fig. 7 Sharing of QTLs across ancestries and tissues.
(a) Distribution of minor allele frequency (MAF) of loci with bos taurus specific (n = 202,583) and overlapped (n = 459) cis-eQTLs in bos indicus population. (b) Distribution of MAF of loci with bos indicus specific (n = 437) and overlapped cis-eQTLs in bos taurus population. (c) Tissue-sharing patterns of cis-sQTL. (d) The gene co-expression patterns across tissues. (e) The co-splicing patterns of spliced introns across tissues. The π1 values are calculated to measure the replication rates between tissues. (f) The Pearson’s correlation of π1 values of matched tissue-pairs (that is, values in the tissue-sharing heatmaps above) across four data types. The P value is obtained by the two-sided Student’s t-test.
Extended Data Fig. 8 Characterization and internal replications of trans-eQTLs.
(a) Comparison of imputation quality (DR2) of trans-eQTLs and all tested SNPs. Dashed lines are median values of DR2. (b) Inter-chromosomal linkage disequilibrium (LD) (the genotype correlation in absolute values) between trans-eQTLs and cis-eQTLs of the same genes in muscle. (c) Comparison of LD of cis-eQTLs vs. trans-eQTLs of the same genes and that of cis-eQTLs vs. random SNPs with matched minor allele frequency (MAF) and chromosomes. The comparisons of all tissues are statistically significant (P < 0.05, the two-sided Student’s t-test). Box plots depict the interquartile range (IQR), whiskers depict 1.5 × IQR. (d) Tissue-sharing patterns (π1 statistics) of trans-eQTLs across tissues and replicates. Muscle_G1 (n = 435) and Muscle_G2 (n = 435) are two replicates of muscle samples by dividing the whole muscle samples randomly into two groups. Similarly, Blood_G1 (n = 349) and Blood_G2(n = 349) are two replicates of blood samples. (e) Pearson correlations of effect sizes (beta values) of trans-eQTLs in one tissue (x-axis) and those of matched SNPs in another tissue (y-axis). (f) Pearson correlation of effect sizes (beta values) of trans-eQTLs (n = 5,782) in blood tissue in Group1 (G1, n = 349) and those of matched SNPs in Group2 (G2, n = 349) (r = 0.66, the two-sided Student’s t-test: P < 2.2 × 10−308). (g) Pearson correlation of effect sizes (beta values) of trans-eQTLs (n = 4,344) in muscle tissue in Group1 (G1, n = 435) and those of matched SNPs in Group2 (G2, n = 435) (r = 0.75, the two-sided Student’s t-test: P < 2.2 × 10−308).
Extended Data Fig. 9 Associations of cis-eQTLs and GWAS loci for important agronomic traits in cattle.
(a) cis-eQTLs discovered in monocytes, and mammary gland show enrichments for top SNPs (top 10%) associated with age at first calving (AFC) (P = 0.001, the two-sided permutation test with 1,000 times), and somatic cell score (SCS) (P = 0.001, the two-sided permutation test with 1,000 times) respectively, compared to the null expectation (shown in gray) defined by ‘Not eQTLs’. (b) Pearson correlation between z-scores from permutation tests (1000 times) and sample sizes of cis-eQTL tissues for milk yield trait (top, r = 0.70, the two-sided Student’s t-test: P = 0.0002) and ketosis trait (bottom, r = 0.49, the two-sided Student’s t-test: P = 0.02). (c) An example of a colocalization of cis-eQTLs of TIGAR gene in muscle and GWAS loci of strength in cattle on chromosome 5. Four independent GWAS signals (that is, rs210875465, rs381714832, rs1115089453 and rs135928140) are located within the region. The All-but-One conditional analysis across the individual GWAS signals shows that only rs135928140 in strength GWAS is colocalized with cis-eQTLs of TIGAR in muscle, when conditioning on the remaining three signals. The colocalized SNP (that is, rs132865837) of TIGAR in muscle is in LD (r2 = 0.49) with the GWAS loci rs135928140. (d) Colocalization between GWAS loci of sire calving ease (Sire_Calv_Ease) in cattle and cis-eQTLs across 23 distinct tissues.
Extended Data Fig. 10 Locuscompare plots for six colocalized events detected by two TWAS methods (S-PrediXcan and MetaXcan), fastENLOC and Coloc simultaneously.
(a) eQTLs of DGAT1 colocalized with GWAS signals of fat percentage in liver. (b) eQTLs of CACNB3 colocalized with GWAS signals of protein percentage in blood. (c) eQTLs of MRPL45 colocalized with GWAS signals of somatic cell score (SCS) in blood. (d) eQTLs of P2RY2 colocalized with GWAS signals of protein percentage in muscle. (e) eQTLs of TSPAN32 colocalized with GWAS signals of Teat length in uterus. (f) eQTLs of DGAT1 colocalized with GWAS signals of udder depth in liver.
Supplementary information
Supplementary Tables
Supplementary Table 1. Detailed information of all RNA-seq data being analyzed. Supplementary Table 2. Detailed information of 144 whole-genome bisulfite sequence (WGBS) data. Supplementary Table 3. Imputation accuracy of SNPs generated from RNA-seq compared with those from WGS. Supplementary Table 4. Summary of cis-eGenes and cis-sGenes across 23 distinct tissues. Supplementary Table 5. GO term enrichments (clusterProfiler v.3.0.4 in R, one-sided Fisher’s exact test) of genes with no cis-eQTLs or no cis-sQTLs in any tissues. Supplementary Table 6. Genes are not significant cis-eGenes in any tissues. Supplementary Table 7. Genes are not significant cis-sGenes in any tissues. Supplementary Table 8. Significant interactions between SNP and subspecies (SNP × Subspecies) for 437 Bos indicus-specific cis-eQTLs in muscle tissue. Supplementary Table 9. Trans-eQTLs and Trans-sQTLs for 15 tissues (Sample size > 100) detected using a simple linear model implemented in MatrixEQTL and a mixed linear regression model (mlma) implemented in the GCTA software. Supplementary Table 10. Detailed information of 43 agronomic traits in cattle. Supplementary Table 11. Summary of significant transcriptome-wide association studies (TWAS) results from S-PrediXcan for 43 traits in cattle. Supplementary Table 12. Significant colocalizations (fastENLOC, Regional colocalization probability, rcp > 0.5) between cis-eQTLs from 22 tissues and 25 agronomic traits. Supplementary Table 13. Significant colocalizations (coloc, posterior probability of ‘both traits are associated and share a single causal variant’, PP.H4 > 0.8) between cis-eQTLs from 6 tissues and 32 agronomic traits. Supplementary Table 14. Significant TWAS results from S-MultiXcan (Bonferroni corrected P < 4×10−6) aggregated across tissues. Supplementary Table 15. Sample information of 207 public WGS for the reference genotype imputation panel.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, S., Gao, Y., Canela-Xandri, O. et al. A multi-tissue atlas of regulatory variants in cattle. Nat Genet 54, 1438–1447 (2022). https://doi.org/10.1038/s41588-022-01153-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01153-5
This article is cited by
-
Genomic dissection of the correlation between milk yield and various health traits using functional and evolutionary information about imputed sequence variants of 34,497 German Holstein cows
BMC Genomics (2024)
-
Molecular quantitative trait loci in reproductive tissues impact male fertility in cattle
Nature Communications (2024)
-
Genome-wide association analysis of heifer livability and early first calving in Holstein cattle
BMC Genomics (2023)
-
Sequence-based GWAS meta-analyses for beef production traits
Genetics Selection Evolution (2023)
-
Sequenced-based GWAS for linear classification traits in Belgian Blue beef cattle reveals new coding variants in genes regulating body size in mammals
Genetics Selection Evolution (2023)