Abstract
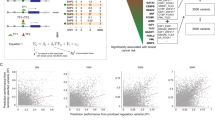
While many germline cancer risk variants have been identified through genome-wide association studies (GWAS), the mechanisms by which these variants operate remain largely unknown. Here we used 406 cancer ATAC-Seq samples across 23 cancer types to identify 7,262 germline allele-specific accessibility QTLs (as-aQTLs). Cancer as-aQTLs had stronger enrichment for cancer risk heritability (up to 145 fold) than any other functional annotation across seven cancer GWAS. Most cancer as-aQTLs directly altered transcription factor (TF) motifs and exhibited differential TF binding and gene expression in functional screens. To connect as-aQTLs to putative risk mechanisms, we introduced the regulome-wide associations study (RWAS). RWAS identified genetically associated accessible peaks at >70% of known breast and prostate loci and discovered new risk loci in all examined cancer types. Integrating as-aQTL discovery, motif analysis and RWAS identified candidate causal regulatory elements and their probable upstream regulators. Our work establishes cancer as-aQTLs and RWAS analysis as powerful tools to study the genetic architecture of cancer risk.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Full allelic imbalance results and all RWAS model weights are available at https://doi.org/10.5281/zenodo.6371439. ATAC-Seq data for all cancer samples are available at https://gdc.cancer.gov/about-data/publications/ATACseq-AWG. SNP-SELEX assay data is available at http://renlab.sdsc.edu/GVATdb/search.html. SuRE assay data is available at https://osf.io/6wev3/. The hg19 reference genome (human_g1k_v37) can be found at https://www.internationalgenome.org/category/grch37/. GTEx v.8 data can be found at https://www.gtexportal.org/home/datasets. Cancer eQTL data can be found at http://gong_lab.hzau.edu.cn/PancanQTL/. TWAS models can be found at http://gusevlab.org/projects/fusion/.
Code availability
Code to conduct allelic imbalance analyses, build RWAS models and conduct RWAS analyses is available at https://doi.org/10.5281/zenodo.6371678.
References
Zhang, H. et al. Genome-wide association study identifies 32 novel breast cancer susceptibility loci from overall and subtype-specific analyses. Nat. Genet. 52, 572–581 (2020).
Michailidou, K. et al. Association analysis identifies 65 new breast cancer risk loci. Nature 551, 92–94 (2017).
Conti, D. V. et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 53, 65–75 (2021).
McKay, J. D. et al. Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat. Genet. 49, 1126–1132 (2017).
Sud, A., Kinnersley, B. & Houlston, R. S. Genome-wide association studies of cancer: current insights and future perspectives. Nat. Rev. Cancer 17, 692–704 (2017).
Fachal, L. et al. Fine-mapping of 150 breast cancer risk regions identifies 191 likely target genes. Nat. Genet. 52, 56–73 (2020).
Gusev, A. et al. A transcriptome-wide association study of high-grade serous epithelial ovarian cancer identifies new susceptibility genes and splice variants. Nat. Genet. 51, 815–823 (2019).
Wu, L. et al. A transcriptome-wide association study of 229,000 women identifies new candidate susceptibility genes for breast cancer. Nat. Genet. 50, 968–978 (2018).
Mancuso, N. et al. Large-scale transcriptome-wide association study identifies new prostate cancer risk regions. Nat. Commun. 9, 4079 (2018).
Hormozdiari, F. et al. Leveraging molecular quantitative trait loci to understand the genetic architecture of diseases and complex traits. Nat. Genet. 50, 1041–1047 (2018).
Liu, X. et al. Functional architectures of local and distal regulation of gene expression in multiple human tissues. Am. J. Hum. Genet. 100, 605–616 (2017).
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Gaffney, D. J. et al. Dissecting the regulatory architecture of gene expression QTLs. Genome Biol. 13, R7 (2012).
Battle, A. & Montgomery, S. B. Determining causality and consequence of expression quantitative trait loci. Hum. Genet. 133, 727–735 (2014).
Brown, A. A. et al. Predicting causal variants affecting expression by using whole-genome sequencing and RNA-seq from multiple human tissues. Nat. Genet. 49, 1747–1751 (2017).
Gong, J. et al. PancanQTL: systematic identification of cis-eQTLs and trans-eQTLs in 33 cancer types. Nucleic Acids Res. 46, D971–D976 (2018).
Geeleher, P. et al. Cancer expression quantitative trait loci (eQTLs) can be determined from heterogeneous tumor gene expression data by modeling variation in tumor purity. Genome Biol. 19, 130 (2018).
Li, Q. et al. Expression QTL-based analyses reveal candidate causal genes and loci across five tumor types. Hum. Mol. Genet. 23, 5294–5302 (2014).
Gamazon, E. R. et al. Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation. Nat. Genet. 50, 956–967 (2018).
Mu, Z. et al. The impact of cell type and context-dependent regulatory variants on human immune traits. Genome Biol. 22, 122 (2021).
Yao, D. W., O’Connor, L. J., Price, A. L. & Gusev, A. Quantifying genetic effects on disease mediated by assayed gene expression levels. Nat. Genet. 52, 626–633 (2020).
Umans, B. D., Battle, A. & Gilad, Y. Where are the disease-associated eQTLs? Trends Genet. 37, 109–124 (2021).
Chun, S. et al. Limited statistical evidence for shared genetic effects of eQTLs and autoimmune-disease-associated loci in three major immune-cell types. Nat. Genet. 49, 600–605 (2017).
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015).
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Gate, R. E. et al. Genetic determinants of co-accessible chromatin regions in activated T cells across humans. Nat. Genet. 50, 1140–1150 (2018).
Liang, D. et al. Cell-type-specific effects of genetic variation on chromatin accessibility during human neuronal differentiation. Nat. Neurosci. 24, 941–953 (2021).
Pickrell, J. K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Yan, H., Yuan, W., Velculescu, V. E., Vogelstein, B. & Kinzler, K. W. Allelic variation in human gene expression. Science 297, 1143 (2002).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Castel, S. E. et al. A vast resource of allelic expression data spanning human tissues. Genome Biol. 21, 234 (2020).
Wang, A. T. et al. Allele-specific QTL fine mapping with PLASMA. Am. J. Hum. Genet. 106, 170–187 (2020).
Liang, Y., Aguet, F., Barbeira, A. N., Ardlie, K. & Im, H. K. A scalable unified framework of total and allele-specific counts for cis-QTL, fine-mapping, and prediction. Nat. Commun. 12, 1424 (2021).
Gutierrez-Arcelus, M. et al. Allele-specific expression changes dynamically during T cell activation in HLA and other autoimmune loci. Nat. Genet. 52, 247–253 (2020).
Houlahan, K. E. et al. Genome-wide germline correlates of the epigenetic landscape of prostate cancer. Nat. Med. 25, 1615–1626 (2019).
Gusev, A. et al. Allelic imbalance reveals widespread germline-somatic regulatory differences and prioritizes risk loci in renal cell carcinoma. Preprint at bioRxiv https://doi.org/10.1101/631150 (2019).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Kumasaka, N., Knights, A. J. & Gaffney, D. J. Fine-mapping cellular QTLs with RASQUAL and ATAC-seq. Nat. Genet. 48, 206–213 (2016).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Huyghe, J. R. et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat. Genet. 51, 76–87 (2019).
Schumacher, F. R. et al. Association analyses of more than 140,000 men identify 63 new prostate cancer susceptibility loci. Nat. Genet. 50, 928–936 (2018).
Scelo, G. et al. Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nat. Commun. 8, 15724 (2017).
Melin, B. S. et al. Genome-wide association study of glioma subtypes identifies specific differences in genetic susceptibility to glioblastoma and non-glioblastoma tumors. Nat. Genet. 49, 789–794 (2017).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
Gazal, S., Marquez-Luna, C., Finucane, H. K. & Price, A. L. Reconciling S-LDSC and LDAK functional enrichment estimates. Nat. Genet. 51, 1202–1204 (2019).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Watt, A. C. et al. CDK4/6 inhibition reprograms the breast cancer enhancer landscape by stimulating AP-1 transcriptional activity. Nat. Cancer 2, 34–48 (2020).
Eferl, R. & Wagner, E. F. AP-1: a double-edged sword in tumorigenesis. Nat. Rev. Cancer 3, 859–868 (2003).
Verde, P., Casalino, L., Talotta, F., Yaniv, M. & Weitzman, J. B. Deciphering AP-1 function in tumorigenesis: fra-ternizing on target promoters. Cell Cycle 6, 2633–2639 (2007).
Kharman-Biz, A. et al. Expression of activator protein-1 (AP-1) family members in breast cancer. BMC Cancer 13, 441 (2013).
Tang, Y., Shu, G., Yuan, X., Jing, N. & Song, J. FOXA2 functions as a suppressor of tumor metastasis by inhibition of epithelial-to-mesenchymal transition in human lung cancers. Cell Res. 21, 316–326 (2011).
Parolia, A. et al. Distinct structural classes of activating FOXA1 alterations in advanced prostate cancer. Nature 571, 413–418 (2019).
Radhakrishnan, S. K. & Gartel, A. L. FOXM1: the Achilles’ heel of cancer? Nature reviews. Cancer vol. 8 c1; author reply c2 (2008).
Chakrabarti, R. et al. Elf5 inhibits the epithelial-mesenchymal transition in mammary gland development and breast cancer metastasis by transcriptionally repressing Snail2. Nat. Cell Biol. 14, 1212–1222 (2012).
Peng, C. et al. Cyclin-dependent kinase 2 (CDK2) is a key mediator for EGF-induced cell transformation mediated through the ELK4/c-Fos signaling pathway. Oncogene 35, 1170–1179 (2016).
Cheng, M. et al. Transcription Factor ELF1 Activates MEIS1 Transcription and Then Regulates the GFI1/FBW7 Axis to Promote the Development of Glioma. Mol. Ther. Nucleic Acids 23, 418–430 (2021).
Jané-Valbuena, J. et al. An oncogenic role for ETV1 in melanoma. Cancer Res. 70, 2075–2084 (2010).
Pellecchia, A. et al. Overexpression of ETV4 is oncogenic in prostate cells through promotion of both cell proliferation and epithelial to mesenchymal transition. Oncogenesis 1, e20 (2012).
Miao, B. et al. The transcription factor FLI1 promotes cancer progression by affecting cell cycle regulation. Int. J. Cancer 147, 189–201 (2020).
Yan, J. Systematic analysis of binding of transcription factors to noncoding variants. Nature 591, 147–151 (2021).
van Arensbergen, J. et al. Genome-wide mapping of autonomous promoter activity in human cells. Nat. Biotechnol. 35, 145–153 (2017).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Dadaev, T. et al. Fine-mapping of prostate cancer susceptibility loci in a large meta-analysis identifies candidate causal variants. Nat. Commun. 9, 2256 (2018).
Baca, S. C. et al. Genetic determinants of chromatin reveal prostate cancer risk mediated by context-dependent gene regulation. Preprint at bioRxiv https://doi.org/10.1101/2021.05.10.443466 (2021).
Forbes, S. A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011).
Pasaniuc, B. et al. Enhanced statistical tests for GWAS in admixed populations: assessment using African Americans from CARe and a Breast Cancer Consortium. PLoS Genet. 7, e1001371 (2011).
Chimge, N.-O. & Frenkel, B. The RUNX family in breast cancer: relationships with estrogen signaling. Oncogene 32, 2121–2130 (2013).
Kalita, C. A. & Gusev, A. A novel method to identify cell-type specific regulatory variants and their role in cancer risk. bioRxiv https://doi.org/10.1101/2021.11.11.468278 (2021).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Boix, C. A., James, B. T., Park, Y. P., Meuleman, W. & Kellis, M. Regulatory genomic circuitry of human disease loci by integrative epigenomics. Nature 590, 300–307 (2021).
Mancuso, N. et al. Integrating gene expression with summary association statistics to identify genes associated with 30 complex traits. Am. J. Hum. Genet. 100, 473–487 (2017).
Bonder, M. J. et al. Identification of rare and common regulatory variants in pluripotent cells using population-scale transcriptomics. Nat. Genet. 53, 313–321 (2021).
Strober, B. J. et al. Dynamic genetic regulation of gene expression during cellular differentiation. Science 364, 1287–1290 (2019).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
Fulco, C. P. et al. Activity-by-contact model of enhancer-promoter regulation from thousands of CRISPR perturbations. Nat. Genet. 51, 1664–1669 (2019).
Nasser, J. et al. Genome-wide enhancer maps link risk variants to disease genes. Nature 593, 238–243 (2021).
Weissbrod, O. et al. Functionally informed fine-mapping and polygenic localization of complex trait heritability. Nat. Genet. 52, 1355–1363 (2020).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Broad Institute TCGA Genome Data Analysis Center. Analysis-ready standardized TCGA data from Broad GDAC Firehose 2016_01_28 run. (2016) https://doi.org/10.7908/C11G0KM9
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Zhou, D. et al. A unified framework for joint-tissue transcriptome-wide association and Mendelian randomization analysis. Nat. Genet. 52, 1239–1246 (2020).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Meyer, K. B. et al. Allele-specific up-regulation of FGFR2 increases susceptibility to breast cancer. PLoS Biol. 6, e108 (2008).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012). S1–S3.
Acknowledgements
We thank M. Freedman, B. Pasaniuc and N. Mancuso for providing feedback on the manuscript. D.G. and A.G. were supported by R01 CA227237 and R01 CA244569. A.G. was also supported by R01 MH115676 and R01 CA259200. D.G. was also supported by the IBM Ph.D. Fellowship Award.
Author information
Authors and Affiliations
Contributions
A.G. conceived and supervised the project and developed RWAS. D.G. conducted all analyses. D.G. and A.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Christopher Amos, Jason Stein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effects of population structure, sample size, peak definitions and CNVs on imbalance analysis.
(a) Inferred genetic ancestry of ATAC-seq samples; each sample is a point color-coded by self-reported race (unfilled circles represent “not reported”) plotted along with the top two genetic Principal Components (PC) shown on the x and y axes. Allelic imbalance is not inflated by population structure due to “canceling out” trans/environmental effects by comparing functional activity at the two alleles within an individual, and only testing heterozygous carriers (thus not biased by population differences in allele frequency). (b) The number of discovered significant as-aQTLs (y-axis) grows linearly with the number of (randomly downsampled) cancer samples (x-axis). (c) The discovery of as-aQTLs does not strongly depend on peak calling with MACS. Many of the same regions are identified without any previous peak calling. (d) The density of as-aQTLs (y-axis) increases exponentially in high-CNV genomic regions (x-axis). The segment mean values are calculated from TCGA somatic copy number calls (segment mean =| log2(CNV/2)|). The chosen maximum threshold of 0.6 corresponds to the gain of one additional allele copy (0.58 ≈| log2(3/2) |).
Extended Data Fig. 2 Cancer as-aQTLs are strongly enriched for cancer risk heritability.
(a) Meta-analysis of 7 cancer types shows that as-aQTLs are more strongly enriched for cancer risk heritability (x-axis) than any other evaluated annotation (y-axis/bars); error bars correspond to the estimated standard error. Data are presented as inverse-variance weighted mean values + /- s.e. (b) as-aQTLs discovered in all 7 cancer types are also enriched for cancer risk variants when the enrichment is quantified using a simpler strategy based on top GWAS associations that does not account for background annotations (see Supplementary Note). Z-scores for the significance of enrichment are shown.
Extended Data Fig. 3 Heritability enrichment at cancer as-aQTLs is cancer type-specific.
(a-c) Cancer type-specific as-aQTLs do not exhibit heritability enrichment for noncancer traits. Data are presented as mean values + /- s.e. (d-e) The cancer risk heritability enrichment at pancancer as-aQTLs is lower than at matching cancer type-specific as-aQTLs. Data are presented as mean values + /- s.e.
Extended Data Fig. 4 Cancer as-aQTLs and d-as-aQTLs are strongly enriched for eQTLs.
(a) The enrichment of cancer-specific accessible peaks (by cancer type: y-axis) for intersecting GTEx tissue eQTLs (by tissue: x-axis) relative to random genomic sequences. Each cell reports the corresponding enrichment (ratio of eQTLs in peaks to eQTLs in random regions) and is shaded by this value. The pancancer peak set (top row) is expected to have a higher fraction of rare/private peaks, which likely explains the apparent lower eQTL enrichment. (b) The fraction of as-aQTLs (by cancer type: y-axis) that intersect one or more GTEx tissue eQTLs (by tissue: x-axis). Each cell reports the fraction and is shaded by this value. For many cancer-tissue pairs, >50% of as-aQTLs contain eQTLs. (c) The enrichment of cancer d-as-aQTLs for intersecting GTEx tissue eQTLs relative to all cancer type-specific peaks. Each cell reports the corresponding enrichment (ratio) and is shaded by this value. Non-significant enrichments (Z < 2, see Methods) are shown as NA. (d) The average Z-scores for the enrichment of cancer as-aQTLs for GTEx tissue eQTLs (averaged across all GTEx tissues, y-axis) as a function of the number of samples in the corresponding cancer type (x-axis). Each point is labeled by the corresponding cancer type. eQTL enrichment z-scores correlate with the number of samples per cancer type.
Extended Data Fig. 5 Distribution of allelic fractions at as-aQTLs with functional and disrupted TF motifs.
(a-l) Violin plots for selected TF motifs representing 12 motif families. The distribution of allelic fractions at as-aQTLs is shown for functional (red) and disrupted (blue) TF motifs.
Extended Data Fig. 6 Additional TF motif scores calculations at as-aQTLs and correlations between as-aQTL allelic fractions and TF binding / gene expression.
(a) Allelic fraction differences (left) and HOMER motif scores (right) from an additional 30 TF motifs for which the difference in motif scores between sequences with high and lower allelic fractions was most significant. Alleles with higher allelic fractions are shown in blue and alleles with lower allelic fractions in orange. Note the reversed directionality for the LRF motif, which is known to act as a repressor of transcription (Constantinou, C. et al. The multi-faceted functioning portrait of LRF/ZBTB7A. Hum. Genomics 13, 66 (2019)). Data are presented as mean values + /- s.d. (b) The correlation between significant SNP-SELEX SNP-TF pair PBS values (p < 0.01) and as-aQTL allelic fractions (y-axis) does not significantly change with varying allelic fraction thresholds |AF-0.5 | (x-axis). Data are presented as Pearson correlations + /- s.e. Number of pairs used for correlation analysis are shown above each point. (c) The correlation between significant SuRE SNP ΔExpressionALT-REF values (p < 0.00173121) and as-aQTL allelic fractions (y-axis) changes only slightly with varying allelic fraction thresholds |AF-0.5 | (x-axis). Data are presented as Pearson correlations + /- s.e. Number of pairs used for correlation analysis are shown above each point.
Extended Data Fig. 7 Additional RWAS analyses.
(a) Numbers of significant RWAS associations between peaks with variants and cancer risk discovered for 7 cancer types using 6 model types. (b) Numbers of significant RWAS associations between peaks without variants and cancer risk discovered for 7 cancer types using 6 model types. In total, 491036 peaks were analyzed of which 337793 (~69%) had a variant and 153243 (~31%) did not. (c) TWAS conducted using GTEx normal tissue expression data yields a comparable number of GWAS risk loci with TWAS genes. The lower number of prostate cancer TWAS genes is due to the smaller sample size of the normal prostate RNA-Seq dataset (132 GTEx samples vs. 468 TCGA samples). (d) RWAS still outperforms TWAS when only top1.total and top1.lasso model types are used (the same model types used for TWAS). (e) The correlations between heritable (cross-validation P of < 0.05) and TWAS-significant genes and RWAS-significant peaks at breast and prostate cancer risk loci are significantly stronger than the correlations between heritable (cross-validation p-values of < 0.05) but non-significant TWAS genes and RWAS peaks at GWAS risk loci. Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)). (f) The correlation between heritable TWAS genes and RWAS peaks increases with decreasing distance. RWAS peak correlations with TWAS genes identified from expression in normal tissue are not significantly different from correlations with TWAS genes identified from the expression in cancer tissues. Matching TCGA breast normal and cancer samples were used for TWAS. Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)).
Extended Data Fig. 8 Prostate cancer risk-associated RWAS peaks are linked to risk-associated TWAS genes.
(a) Correlations between TWAS gene and RWAS peak pairs at 33 prostate cancer GWAS risk loci. Nodes representing TWAS genes are shown in red with gene names shown in brackets. Nodes representing RWAS peaks are shown in black. The color of the edges represents the strength of the correlations between models (absolute Pearson correlation). (b) Median absolute Pearson correlations between significant TWAS gene and RWAS peak pairs at each GWAS prostate cancer risk locus. Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)).
Extended Data Fig. 9 Prostate cancer risk-associated RWAS peaks are linked to risk-associated CWAS features.
(a) Overlap of significant RWAS and CWAS (H3K27ac) peaks with prostate cancer GWAS risk loci and significant RWAS and CWAS (AR) peaks with prostate cancer GWAS risk loci. (b) Correlations between prostate cancer risk-associated RWAS and CWAS peak pairs across all prostate cancer GWAS risk loci are similar to the RWAS-TWAS associations (Extended Data Fig. 8). Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)). (c) Correlations between CWAS (H3K27ac) and RWAS peak pairs at 42 prostate cancer GWAS risk loci. Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)). (d) Correlations between CWAS (AR) and RWAS peak pairs at 27 prostate cancer GWAS risk loci. Horizontal lines inside the boxes indicate the medians. Box bounds show Q1 and Q3. Whiskers are minima (Q1 - 1.5x(Q3-Q1)) and maxima (Q3 + 1.5x(Q3-Q1)).
Extended Data Fig. 10 RWAS associations can explain GWAS risk loci.
(a-f) Examples of RWAS-significant peaks near COSMIC genes that explain a large portion of the GWAS signal in conditional analyses. Each dot corresponds to a GWAS SNP with the significance of the association on the y-axis and physical position on the x-axis. The gray dots indicate marginal GWAS signals, and the blue dots show the same signals become less significant after conditioning on an as-aQTL/balanced peak identified in RWAS.
Supplementary information
Supplementary Information
Supplementary Notes and References
Supplementary Table 1
Supplementary Tables 1–8
Rights and permissions
About this article
Cite this article
Grishin, D., Gusev, A. Allelic imbalance of chromatin accessibility in cancer identifies candidate causal risk variants and their mechanisms. Nat Genet 54, 837–849 (2022). https://doi.org/10.1038/s41588-022-01075-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01075-2
This article is cited by
-
Allele-specific binding (ASB) analyzer for annotation of allele-specific binding SNPs
BMC Bioinformatics (2023)
-
Molecular quantitative trait loci
Nature Reviews Methods Primers (2023)
-
Leveraging molecular quantitative trait loci to comprehend complex diseases/traits from the omics perspective
Human Genetics (2023)