Abstract
Enlargement or aneurysm of the aorta predisposes to dissection, an important cause of sudden death. We trained a deep learning model to evaluate the dimensions of the ascending and descending thoracic aorta in 4.6 million cardiac magnetic resonance images from the UK Biobank. We then conducted genome-wide association studies in 39,688 individuals, identifying 82 loci associated with ascending and 47 with descending thoracic aortic diameter, of which 14 loci overlapped. Transcriptome-wide analyses, rare-variant burden tests and human aortic single nucleus RNA sequencing prioritized genes including SVIL, which was strongly associated with descending aortic diameter. A polygenic score for ascending aortic diameter was associated with thoracic aortic aneurysm in 385,621 UK Biobank participants (hazard ratio = 1.43 per s.d., confidence interval 1.32–1.54, P = 3.3 × 10−20). Our results illustrate the potential for rapidly defining quantitative traits with deep learning, an approach that can be broadly applied to biomedical images.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
UK Biobank data is made available to researchers from universities and other research institutions with genuine research inquiries, following IRB and UK Biobank approval. Full GWAS summary statistics for ascending and descending thoracic aortic measurements are available at the Broad Institute Cardiovascular Disease Knowledge Portal (http://www.broadcvdi.org). Single nucleus RNA sequencing data are publicly available at the Broad Institute’s Single Cell Portal (accession no. SCP1265, https://singlecell.broadinstitute.org/single_cell) and at the National Center for Biotechnology Information’s Gene Expression Omnibus Database (accession no. GSE165824). The dbGAP accession number for aortic phenotypes used in FHS replication is phs000007.v30.p11. All other data are contained within the article and its supplementary information, or are available upon reasonable request to the corresponding author.
Code availability
The code used to identify connected components is available as a Go library at https://github.com/carbocation/genomisc/tree/master/overlay and a README is provided in that folder to demonstrate library usage.
Change history
10 December 2021
In the version of this article initially published online, the link for Supplementary Tables 1–21 was missing and has been restored as of 10 December 2021.
References
Benjamin, E. J. et al. Heart disease and stroke statistics—2019 update: a report from the American Heart Association. Circulation 139, e56–e528 (2019).
Isselbacher, E. M. Thoracic and abdominal aortic aneurysms. Circulation 111, 816–828 (2005).
Owens, D. K. et al. Screening for abdominal aortic aneurysm: US Preventive Services Task Force Recommendation Statement. JAMA 322, 2211–2218 (2019).
Fann, J. I. Descending thoracic and thoracoabdominal aortic aneurysms. Coron. Artery Dis. 13, 93–102 (2002).
Guo, D.-C., Papke, C. L., He, R. & Milewicz, D. M. Pathogenesis of thoracic and abdominal aortic aneurysms. Ann. N. Y. Acad. Sci. 1085, 339–352 (2006).
Vapnik, J. S. et al. Characteristics and outcomes of ascending versus descending thoracic aortic aneurysms. Am. J. Cardiol. 117, 1683–1690 (2016).
Jondeau, G. & Boileau, C. Familial thoracic aortic aneurysms. Curr. Opin. Cardiol. 29, 492–498 (2014).
Pinard, A., Jones, G. T. & Milewicz, D. M. Genetics of thoracic and abdominal aortic diseases. Circ. Res. 124, 588–606 (2019).
Verstraeten, A., Luyckx, I. & Loeys, B. Aetiology and management of hereditary aortopathy. Nat. Rev. Cardiol. 14, 197–208 (2017).
Lindsay, M. E. & Dietz, H. C. Lessons on the pathogenesis of aneurysm from heritable conditions. Nature 473, 308–316 (2011).
Majesky, M. W. Developmental basis of vascular smooth muscle diversity. Arterioscler. Thromb. Vasc. Biol. 27, 1248–1258 (2007).
Hagan, P. G. et al. The International Registry of Acute Aortic Dissection (IRAD): new insights into an old disease. JAMA 283, 897–903 (2000).
Howard, J. & Gugger, S. Fastai: a layered API for deep learning. Information 11, 108 (2020).
Ronneberger, O., Fischer, P. & Brox, T. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. MICCAI 2015 (eds Navab, N. et al.) (Lecture Notes in Computer Science, Vol. 9351, Springer, 2015); https://doi.org/10.1007/978-3-319-24574-4_28
Deng, J. et al. ImageNet: a large-scale hierarchical image database. In Proc. 2009 IEEE Conference on Computer Vision and Pattern Recognition 248–255 (IEEE, 2009); https://doi.org/10.1109/CVPR.2009.5206848
Rosenfeld, A. & Pfaltz, J. L. Sequential operations in digital picture processing. JACM 13, 471–494 (1966).
Turkbey, E. B. et al. Determinants and normal values of ascending aortic diameter by age, gender and race/ethnicity in the Multi-Ethnic Study of Atherosclerosis (MESA). J. Magn. Reson. Imaging 39, 360–368 (2014).
Kaplan, S. et al. Prevalence of an increased ascending and descending thoracic aorta diameter diagnosed by multislice cardiac computed tomography in men versus women and in persons aged 23 to 50 years, 51 to 65 years, 66 to 80 years, and 81 to 88 years. Am. J. Cardiol. 100, 1598–1599 (2007).
Campens, L. et al. Reference values for echocardiographic assessment of the diameter of the aortic root and ascending aorta spanning all age categories. Am. J. Cardiol. 114, 914–920 (2014).
Wu, P. et al. Mapping ICD-10 and ICD-10-CM codes to phecodes: workflow development and initial evaluation. JMIR Med. Inform. 7, e14325 (2019).
Bradley, T. J., Bowdin, S. C., Morel, C. F. J. & Pyeritz, R. E. The expanding clinical spectrum of extracardiovascular and cardiovascular manifestations of heritable thoracic aortic aneurysm and dissection. Can. J. Cardiol. 32, 86–99 (2016).
Avdic, T. et al. Reduced long‐term risk of aortic aneurysm and aortic dissection among individuals with type 2 diabetes mellitus: a nationwide observational study. J. Am. Heart Assoc. 7, e007618 (2018).
Prakash, S. K., Pedroza, C., Khalil, Y. A. & Milewicz, D. M. Diabetes and reduced risk for thoracic aortic aneurysms and dissections: a nationwide case–control study. J. Am. Heart Assoc. 1, e000323 (2012).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Guo, D. et al. Genetic variants in LRP1 and ULK4 are associated with acute aortic dissections. Am. J. Hum. Genet. 99, 762–769 (2016).
van ’t Hof, F. N. G. et al. Shared genetic risk factors of intracranial, abdominal, and thoracic aneurysms. J. Am. Heart Assoc. 5, e002603 (2016).
LeMaire, S. A. et al. Genome-wide association study identifies a susceptibility locus for thoracic aortic aneurysms and aortic dissections spanning FBN1 at 15q21.1. Nat. Genet. 43, 996–1000 (2011).
Vasan, R. S. et al. Genetic variants associated with cardiac structure and function: a meta-analysis and replication of genome-wide association data. JAMA 302, 168–178 (2009).
Wild, P. S. et al. Large-scale genome-wide analysis identifies genetic variants associated with cardiac structure and function. J. Clin. Invest. 127, 1798–1812 (2017).
Rogers, I. S. et al. Distribution, determinants, and normal reference values of thoracic and abdominal aortic diameters by computed tomography (from the Framingham Heart Study). Am. J. Cardiol. 111, 1510–1516 (2013).
Qazi, S. et al. Increased aortic diameters on multidetector computed tomographic scan are independent predictors of incident adverse cardiovascular events: the Framingham Heart Study. Circ. Cardiovasc. Imaging 10, e006776 (2017).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed model association for biobank-scale data sets. Nat. Genet. 50, 906–908 (2018).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Wain, L. V. et al. Novel blood pressure locus and gene discovery using genome-wide association study and expression data sets from blood and the kidney. Hypertension https://doi.org/10.1161/HYPERTENSIONAHA.117.09438 (2017).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Tsutsui, K. et al. ADAMTSL-6 is a novel extracellular matrix protein that binds to fibrillin-1 and promotes fibrillin-1 fibril formation. J. Biol. Chem. 285, 4870–4882 (2010).
Chou, C.-K. et al. The regulations of deubiquitinase USP15 and its pathophysiological mechanisms in diseases. Int. J. Mol. Sci. 18, 483 (2017).
Eichhorn, P. J. A. et al. USP15 stabilizes TGF-β receptor I and promotes oncogenesis through the activation of TGF-β signaling in glioblastoma. Nat. Med. 18, 429–435 (2012).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Bhuwania, R. et al. Supervillin couples myosin-dependent contractility to podosomes and enables their turnover. J. Cell Sci. 125, 2300–2314 (2012).
Linder, S., Wiesner, C. & Himmel, M. Degrading devices: invadosomes in proteolytic cell invasion. Annu. Rev. Cell Dev. Biol. 27, 185–211 (2011).
Elbitar, S. et al. Pathogenic variants in THSD4, encoding the ADAMTS-like 6 protein, predispose to inherited thoracic aortic aneurysm. Genet. Med. 23, 111–122 (2021).
Maddika, S. et al. WWP2 is an E3 ubiquitin ligase for PTEN. Nat. Cell Biol. 13, 728–733 (2011).
Chen, H. et al. WWP2 regulates pathological cardiac fibrosis by modulating SMAD2 signaling. Nat. Commun. 10, 3616 (2019).
Pirruccello, J. P. et al. Analysis of cardiac magnetic resonance imaging in 36,000 individuals yields genetic insights into dilated cardiomyopathy. Nat. Commun. 11, 2254 (2020).
Teslovich, T. M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Renard, M. et al. Clinical validity of genes for heritable thoracic aortic aneurysm and dissection. J. Am. Coll. Cardiol. 72, 605–615 (2018).
Dietz, H. C. et al. Marfan syndrome caused by a recurrent de novo missense mutation in the fibrillin gene. Nature 352, 337–339 (1991).
Falak, S. et al. Protease inhibitor 15, a candidate gene for abdominal aortic internal elastic lamina ruptures in the rat. Physiol. Genomics 46, 418–428 (2014).
Parrott, A. et al. Cantu syndrome: a longitudinal review of vascular findings in three individuals. Am. J. Med. Genet. A 182, 1243–1248 (2020).
Heuzé, M. L. et al. ASB2 targets filamins A and B to proteasomal degradation. Blood 112, 5130–5140 (2008).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Petersen, S. E. et al. Imaging in population science: cardiovascular magnetic resonance in 100,000 participants of UK Biobank – rationale, challenges and approaches. J. Cardiovasc. Magn. Reson. 15, 46 (2013).
Petersen, S. E. et al. UK Biobank’s cardiovascular magnetic resonance protocol. J. Cardiovasc. Magn. Reson. 18, 8 (2016).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Preprint at https://arxiv.org/abs/1512.03385 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun. ACM 60, 84–90 (2017).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2017).
Smith, L. N. Cyclical learning rates for training neural networks. Preprint at https://arxiv.org/abs/1506.01186 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. in Computer Vision – ECCV 2014. ECCV 2014 (eds Fleet, D. et al.) 346–361 (Lecture Notes in Computer Science, Vol. 8691, Springer, 2014).
Horn, B. Robot Vision (The MIT Press, 1986).
Feltz, C. J. & Miller, G. E. An asympotic test for the equality of coefficients of variation from k populations. Stat. Med. 15, 647–658 (1996).
Marwick, B. & Krishnamoorthy, K. cvequality. R package version 0.2.0 (2019).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Yang, J. et al. FTO genotype is associated with phenotypic variability of body mass index. Nature 490, 267–272 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Pers, T. H., Timshel, P. & Hirschhorn, J. N. SNPsnap: a Web-based tool for identification and annotation of matched SNPs. Bioinformatics 31, 418–420 (2015).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
de Leeuw, C. A., Neale, B. M., Heskes, T. & Posthuma, D. The statistical properties of gene-set analysis. Nat. Rev. Genet. 17, 353–364 (2016).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Segrè, A. V. et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Van Hout, C. V. et al. Exome sequencing and characterization of 49,960 individuals in the UK Biobank. Nature 586, 749–756 (2020).
Regier, A. A. et al. Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects. Nat. Commun. 9, 4038 (2018).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Haas, M. E. et al. Genetic association of albuminuria with cardiometabolic disease and blood pressure. Am. J. Hum. Genet. 103, 461–473 (2018).
Bellenguez, C. et al. A robust clustering algorithm for identifying problematic samples in genome-wide association studies. Bioinformatics 28, 134–135 (2012).
Kent, K. C. et al. Analysis of risk factors for abdominal aortic aneurysm in a cohort of more than 3 million individuals. J. Vasc. Surg. 52, 539–548 (2010).
Acknowledgements
This work was supported by the Fondation Leducq grant no. 14CVD01 (P.T.E.); by grants from the National Institutes of Health no.1R01HL092577 (P.T.E.), no. R01HL128914 (P.T.E.), no. K24HL105780 (P.T.E.), no. R01HL134893 (J.E.H.), no. R01HL140224 (J.E.H.), no. 5K01HL140187 (N.R.T.), no. T32HL007208 (S.K.), no. R01HL128914 (E.J.B.), no. 2R01HL092577 (E.J.B.), no. 1R01HL141434 (E.J.B.), no. 2U54HL120163 (E.J.B.), no. 1R01HL139731 (S.A.L.), no. T32HL007208 (E.L.C.), no. K08HL159346 (J.P.P.); by a grant from the American Heart Association Strategically Focused Research Networks (P.T.E.); by the American Heart Association grants no. 18SFRN34110082 (E.J.B.), no. 18SFRN34110082 (A.W.H.), no. 18SFRN34110082 (L.-C.W.), no. 18SFRN34250007 (S.A.L.); by a John S LaDue Memorial Fellowship (J.P.P.); by a Sarnoff Scholar Award (J.P.P.); by a Career Award for Medical Scientists from the Burroughs Wellcome Fund (A.G.B.); and by the Fredman Fellowship for Aortic Disease (M.E.L.) and the Toomey Fund for Aortic Dissection Research (M.E.L.). The Precision Cardiology Laboratory is a joint effort between the Broad Institute and Bayer AG. The rapid autopsy effort was funded by the Susan Eid Tumor Heterogeneity Initiative.
Author information
Authors and Affiliations
Contributions
J.P.P. and P.T.E. conceived of the study. J.P.P. and M.N. annotated images. J.P.P., M.D.C., S.J.F., S.F.F., S.H.C., H.L., E.L.C. and M.N. conducted bioinformatic analyses. E.L.C., A.A., A.-D.A., N.R.T., D.J. and J.R.S. contributed to the rapid autopsy human aorta analysis. H.L., R.S.V., E.J.B. and U.H. contributed to the GWAS replication. J.P.P., M.E.L. and P.T.E. wrote the paper. S.K., A.G.B., L.-C.W., P.B., A.W.H., C.R., S.K.V., R.M.G., C.M.S., J.E.H., S.A.L. and A.A.P. contributed to the analysis plan or provided critical revisions.
Corresponding author
Ethics declarations
Competing interests
J.P.P. and A.G.B. have served as consultants for Maze Therapeutics. A.-D.A. and C.M.S. are employees of Bayer US LLC (a subsidiary of Bayer AG), and may own stock in Bayer AG. D.J. is supported by grants from Genentech, Eisai, EMD Serono, Takeda, Amgen, Celgene, Placon Therapeutics, Syros, Petra Pharma, InventisBio, Infinity Pharmaceuticals and Novartis. D.J. has also received personal fees from Genentech, Eisai, EMD Serono, Ipsen, Syros, Relay Therapeutics, MapKure, Vibliome, Petra Pharma and Novartis. A.A.P. is employed as a Venture Partner at GV; he is also supported by a grant from Bayer AG to the Broad Institute focused on machine learning for clinical trial design. J.E.H. is supported by a grant from Bayer AG focused on machine learning and cardiovascular disease and a research grant from Gilead Sciences. J.E.H. has received research supplies from EcoNugenics. P.B. is supported by grants from Bayer AG and IBM applying machine learning in cardiovascular disease. P.T.E. is supported by a grant from Bayer AG to the Broad Institute focused on the genetics and therapeutics of cardiovascular diseases. P.T.E. has also served on advisory boards or consulted for Bayer AG, Quest Diagnostics, MyoKardia and Novartis. S.A.L. receives sponsored research support from Bristol Myers Squibb/Pfizer, Bayer AG, Boehringer Ingelheim and Fitbit, and has consulted for Bristol Myers Squibb/Pfizer and Bayer AG, and participates in a research collaboration with IBM. The Broad Institute has filed for a patent on an invention from P.T.E., M.E.L. and J.P.P. related to a genetic risk predictor for aortic disease.
Additional information
Peer review information Nature Genetics thanks Chayakrit Krittanawong, Julie De Backer and Richard Redon for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Aortic size by age and sex.
The length of the minor elliptical axis of aorta at its maximum size during the cardiac cycle (that is, the diameter) is shown for the ascending aorta (left) and the descending aorta (right). The x-axis represents the participant’s age at the time of cardiac MRI, and the y-axis represents the size of aorta. Each point represents one person’s measurements; men are plotted in turquoise and women in red. Sex-specific locally weighted scatterplot smoothing (LOESS) curves are overplotted. Each point represents one of the 42,518 participants who passed imaging quality control for at least one of the ascending or descending aorta measurements: 40,363 had accepted measurements for ascending aorta, and 41,415 had accepted measurements for descending aorta.
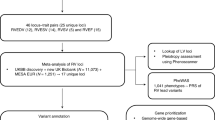
Extended Data Fig. 2 GWAS sample flow diagram.
The GWAS sample flow diagram depicts the sample filtering process that led to the specific samples being chosen for the ascending and descending aortic diameter GWAS.
Extended Data Fig. 3 GWAS QQ plots.
Quantile-quantile plots showing the theoretical distribution of P values under a uniform distribution (x-axis) versus the observed distribution within the sample (y-axis) are displayed for the ascending and descending aorta GWAS summary statistics. The plots are stratified by minor allele frequency (‘maf_bin’): ‘common’ denotes SNPs with MAF > 0.05, low frequency with 0.005 < MAF ≤ 0.05, and rare with 0.001 < MAF ≤ 0.005. Variants with MAF < 0.001 were excluded from the analysis.
Extended Data Fig. 4 GWAS replication in the Framingham Heart Study.
a,b, For lead SNPs from the main UK Biobank GWAS that could be identified in a GWAS from FHS, each SNP is plotted based on the UK Biobank Z score (x-axis) and the FHS Z score (y-axis). 72 SNPs for ascending aortic diameter (a) and 41 SNPs for descending aortic diameter (b) could be identified in FHS and are plotted here. SNPs where the direction of effect is in agreement between FHS and UK Biobank are plotted in blue, while those with opposite direction of effect are marked in red.
Extended Data Fig. 5 Genetic correlation with continuous traits.
The genetic correlation between continuous traits and the ascending (top) and descending (bottom) thoracic aorta in the UK Biobank are represented in volcano plots. Of the 281 tested traits, genetic correlation with 257 traits was computable in the ascending aorta and with 256 traits in the descending aorta. The x-axis represents the magnitude of genetic correlation, while the y-axis represents the -log10 of the genetic correlation P value, based on ldsc. Traits achieving Bonferroni significance are colored red (for positive genetic correlation) or blue (for negative genetic correlation). The top 10 positively and negatively associated traits are labeled. The underlying data are available in Supplementary Table 10.
Extended Data Fig. 6 Cell type-specific gene expression at the WWP2 locus.
Cell-type specificity of genes with expression data within 500 kb of the lead SNP near WWP2. As with Fig. 4, the size of each square represents the average log2(Expr) for a gene across all nuclei in a given cluster. The color represents the log fold-change comparing the expression of the given gene in each cluster to all other clusters based on a formal differential expression model. A dot represents significant up- or down-regulation in the given cluster based on a Benjamini-Hochberg correction for multiple testing at FDR < 0.01. Expr, normalized nucleus-level expression calculated as the number of counts of a gene divided by the total number of counts in the nucleus and multiplied by 10,000; FC, fold-change.
Extended Data Fig. 7 MAGMA gene set associations.
Gene sets enriched in MAGMA analysis of the GWAS of the ascending (top) and descending (bottom) thoracic aorta are represented in volcano plots. The x-axis represents the magnitude of estimated effect of a pathway-based gene set on the aortic trait, while the y-axis represents the -log10 of the MAGMA association P value. Pathways achieving Bonferroni significance are colored red and labeled. The underlying data are available in Supplementary Tables 17 and 18.
Supplementary information
Supplementary Information
Supplementary Note, Figs. 1–5.
Supplementary Tables
Supplementary Tables 1–21.
Rights and permissions
About this article
Cite this article
Pirruccello, J.P., Chaffin, M.D., Chou, E.L. et al. Deep learning enables genetic analysis of the human thoracic aorta. Nat Genet 54, 40–51 (2022). https://doi.org/10.1038/s41588-021-00962-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00962-4
This article is cited by
-
Protein interaction networks in the vasculature prioritize genes and pathways underlying coronary artery disease
Communications Biology (2024)
-
Unsupervised ensemble-based phenotyping enhances discoverability of genes related to left-ventricular morphology
Nature Machine Intelligence (2024)
-
Sex-specific genetic architecture of blood pressure
Nature Medicine (2024)
-
Genetic analysis of cardiac dynamic flow volumes identifies loci mapping aortic root size
Nature Genetics (2024)
-
Genetic architecture of cardiac dynamic flow volumes
Nature Genetics (2024)