Abstract
Glioma intratumoral heterogeneity enables adaptation to challenging microenvironments and contributes to therapeutic resistance. We integrated 914 single-cell DNA methylomes, 55,284 single-cell transcriptomes and bulk multi-omic profiles across 11 adult IDH mutant or IDH wild-type gliomas to delineate sources of intratumoral heterogeneity. We showed that local DNA methylation disorder is associated with cell–cell DNA methylation differences, is elevated in more aggressive tumors, links with transcriptional disruption and is altered during the environmental stress response. Glioma cells under in vitro hypoxic and irradiation stress increased local DNA methylation disorder and shifted cell states. We identified a positive association between genetic and epigenetic instability that was supported in bulk longitudinally collected DNA methylation data. Increased DNA methylation disorder associated with accelerated disease progression and recurrently selected DNA methylation changes were enriched for environmental stress response pathways. Our work identified an epigenetically facilitated adaptive stress response process and highlights the importance of epigenetic heterogeneity in shaping therapeutic outcomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All de-identified, nonprotected access somatic variant calls, single-cell gene expression profiles, regional scDNAme data and scDNAme disorder data are accessible via Synapse (https://synapse.org/singlecellglioma). Raw bulk and single-cell sequencing data and methylation microarray data are available through the European Genome-phenome Archive under accession no. EGAS00001005300. The GRCh37 (hg19) reference genome was obtained from GATK (https://gatk.broadinstitute.org/).
Code availability
Major analysis scripts are available on GitHub (https://github.com/TheJacksonLaboratory/singlecellglioma-verhaaklab) and Zenodo (https://doi.org/10.5281/zenodo.4967364).
References
Brat, D. J. et al. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563 (2016).
Louis, D. N. et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 131, 803–820 (2016).
Barthel, F. P. et al. Longitudinal molecular trajectories of diffuse glioma in adults. Nature 576, 112–120 (2019).
Klughammer, J. et al. The DNA methylation landscape of glioblastoma disease progression shows extensive heterogeneity in time and space. Nat. Med. 24, 1611–1624 (2018).
Körber, V. et al. Evolutionary trajectories of IDHWT glioblastomas reveal a common path of early tumorigenesis instigated years ahead of initial diagnosis. Cancer Cell 35, 692–704.e12 (2019).
Mazor, T. et al. DNA methylation and somatic mutations converge on the cell cycle and define similar evolutionary histories in brain tumors. Cancer Cell 28, 307–317 (2015).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56.e6 (2017).
Flavahan, W. A., Gaskell, E. & Bernstein, B. E. Epigenetic plasticity and the hallmarks of cancer. Science 357, eaal2380 (2017).
Bhaduri, A. et al. Outer radial glia-like cancer stem cells contribute to heterogeneity of glioblastoma. Cell Stem Cell 26, 48–63.e6 (2020).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849.e21 (2019).
Wang, L. et al. The phenotypes of proliferating glioblastoma cells reside on a single axis of variation. Cancer Discov. 9, 1708–1719 (2019).
Yuan, J. et al. Single-cell transcriptome analysis of lineage diversity in high-grade glioma. Genome Med. 10, 57 (2018).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Venteicher, A. S. et al. Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355, eaai8478 (2017).
Easwaran, H., Tsai, H.-C. & Baylin, S. B. Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol. Cell 54, 716–727 (2014).
Liau, B. B. et al. Adaptive chromatin remodeling drives glioblastoma stem cell plasticity and drug tolerance. Cell Stem Cell 20, 233–246.e7 (2017).
Gaiti, F. et al. Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia. Nature 569, 576–580 (2019).
Hernando-Herraez, I. et al. Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells. Nat. Commun. 10, 4361 (2019).
Johnson, K. C. et al. 5-Hydroxymethylcytosine localizes to enhancer elements and is associated with survival in glioblastoma patients. Nat. Commun. 7, 13177 (2016).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Argelaguet, R. et al. Multi-omics profiling of mouse gastrulation at single-cell resolution. Nature 576, 487–491 (2019).
Farlik, M. et al. DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19, 808–822 (2016).
Bian, S. et al. Single-cell multiomics sequencing and analyses of human colorectal cancer. Science 362, 1060–1063 (2018).
Guo, H. et al. Profiling DNA methylome landscapes of mammalian cells with single-cell reduced-representation bisulfite sequencing. Nat. Protoc. 10, 645–659 (2015).
Guo, H. et al. Single-cell methylome landscapes of mouse embryonic stem cells and early embryos analyzed using reduced representation bisulfite sequencing. Genome Res. 23, 2126–2135 (2013).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012).
Kelsey, G., Stegle, O. & Reik, W. Single-cell epigenomics: recording the past and predicting the future. Science 358, 69–75 (2017).
Landau, D. A. et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell 26, 813–825 (2014).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Zhu, J., Tsai, H.-J., Gordon, M. R. & Li, R. Cellular stress associated with aneuploidy. Dev. Cell 44, 420–431 (2018).
Hughes, L. A. E. et al. The CpG island methylator phenotype: what’s in a name? Cancer Res. 73, 5858–5868 (2013).
Luo, Y., Lu, X. & Xie, H. Dynamic Alu methylation during normal development, aging, and tumorigenesis. Biomed. Res. Int. 2014, 784706 (2014).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
MacLeod, G. et al. Genome-wide CRISPR–Cas9 screens expose genetic vulnerabilities and mechanisms of temozolomide sensitivity in glioblastoma stem cells. Cell Rep. 27, 971–986.e9 (2019).
Jin, X. et al. Targeting glioma stem cells through combined BMI1 and EZH2 inhibition. Nat. Med. 23, 1352–1361 (2017).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887.e17 (2019).
Orso, F. et al. Identification of functional TFAP2A and SP1 binding sites in new TFAP2A-modulated genes. BMC Genomics 11, 355 (2010).
Li, Z. et al. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell 15, 501–513 (2009).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010).
Peng, C. et al. Cyclin-dependent kinase 2 (CDK2) is a key mediator for EGF-induced cell transformation mediated through the ELK4/c-Fos signaling pathway. Oncogene 35, 1170–1179 (2016).
Kent, L. N. & Leone, G. The broken cycle: E2F dysfunction in cancer. Nat. Rev. Cancer 19, 326–338 (2019).
Koren, A. et al. Differential relationship of DNA replication timing to different forms of human mutation and variation. Am. J. Hum. Genet. 91, 1033–1040 (2012).
deCarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Morton, A. R. et al. Functional enhancers shape extrachromosomal oncogene amplifications. Cell 179, 1330–1341.e13 (2019).
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat. Genet. 52, 891–897 (2020).
Verhaak, R. G. W., Bafna, V. & Mischel, P. S. Extrachromosomal oncogene amplification in tumour pathogenesis and evolution. Nat. Rev. Cancer 19, 283–288 (2019).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Verburg, N. et al. Spatial concordance of DNA methylation classification in diffuse glioma. Neuro. Oncol., https://doi.org/10.1093/neuonc/noab134 (2021).
de Souza, C. F. et al. A distinct DNA methylation shift in a subset of glioma CpG island methylator phenotypes during tumor recurrence. Cell Rep. 23, 637–651 (2018).
Landan, G. et al. Epigenetic polymorphism and the stochastic formation of differentially methylated regions in normal and cancerous tissues. Nat. Genet. 44, 1207–1214 (2012).
Losman, J.-A. & Kaelin, W. G. Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 27, 836–852 (2013).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17, 510–522 (2010).
Thienpont, B. et al. Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 537, 63–68 (2016).
Heddleston, J. M. et al. Hypoxia inducible factors in cancer stem cells. Br. J. Cancer 102, 789–795 (2010).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Bhat, K. P. L. et al. Mesenchymal differentiation mediated by NF-κB promotes radiation resistance in glioblastoma. Cancer Cell 24, 331–346 (2013).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Hui, T. et al. High-resolution single-cell DNA methylation measurements reveal epigenetically distinct hematopoietic stem cell subpopulations. Stem Cell Rep. 11, 578–592 (2018).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Forrest, A. R. R. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Hunt, S. E. et al. Ensembl variation resources. Database (Oxford) 2018, bay119 (2018).
Raney, B. J. et al. Track data hubs enable visualization of user-defined genome-wide annotations on the UCSC Genome Browser. Bioinformatics 30, 1003–1005 (2014).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Garvin, T. et al. Interactive analysis and assessment of single-cell copy-number variations. Nat. Methods 12, 1058–1060 (2015).
Galili, T. dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31, 3718–3720 (2015).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Chakravarthy, A. et al. Pan-cancer deconvolution of tumour composition using DNA methylation. Nat. Commun. 9, 3220 (2018).
Sheffield, N. C. & Bock, C. LOLA: enrichment analysis for genomic region sets and regulatory elements in R and Bioconductor. Bioinformatics 32, 587–589 (2016).
Köster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 34, 3600 (2018).
Blokzijl, F., Janssen, R., van Boxtel, R. & Cuppen, E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med. 10, 33 (2018).
Deshwar, A. G. et al. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 16, 35 (2015).
Ha, G. et al. TITAN: inference of copy number architectures in clonal cell populations from tumor whole-genome sequence data. Genome Res. 24, 1881–1893 (2014).
Favero, F. et al. Sequenza: allele-specific copy number and mutation profiles from tumor sequencing data. Ann. Oncol. 26, 64–70 (2015).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Deshpande, V. et al. Exploring the landscape of focal amplifications in cancer using AmpliconArchitect. Nat. Commun. 10, 392 (2019).
Acknowledgements
We thank the patients and their families for their generous donation to biomedical research. We also thank the staff in the following groups at The Jackson Laboratory for Genomic Medicine: single-cell biology laboratory; flow cytometry core; and genomic technology core for assistance in data generation. We thank M. Wimsatt and Z. Reifsnyder for assistance in graphic design. We thank the University of Texas MD Anderson Epigenomics Profiling Core for their assistance in helping troubleshoot the scRRBS protocol. We thank the Henry Ford Hospital for sharing the patient-derived glioma spheroids. This work was supported by National Institutes of Health grants R01 CA237208 and R21 NS114873, Cancer Center Support Grant P30 CA034196, Department of Defense grant no. W81XWH1910246 (to R.G.W.V) and Jackson Laboratory Cancer Center Fast Forward funds. F.S.V. is supported by a postdoctoral fellowship from The Jane Coffin Childs Memorial Fund for Medical Research. F.P.B. is supported by the National Cancer Institute (grant no. K99 CA226387). E.Y. is a fellow of the American Brain Tumor Association. K.C.J. is the recipient of an American Cancer Society Fellowship (no. 130984-PF-17-141-01-DMC). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
K.C.J. and R.G.W.V. conceived the project and designed the experiments. K.B. and S.D. curated the patient samples and patient annotation. N.V. and P.C.d.W.H. provided input on the multi-sector analyses. K.C.J., M.R.H.E., M.T., N.E.N., R.M., C.Y.N., M.L.S. and P.R. performed the single-cell library optimization and sequencing. K.C.J. led the data production and performed the experiments with A.D.G., D.Z., D.L., E.Y. and E.T.C. K.C.J. and K.J.A. led the data analysis in collaboration with F.P.B., F.S.V., M.S., E.Y. and H.K. K.C.J., K.J.A. and R.G.W.V. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
R.G.W.V. is a cofounder of and has received research support from Boundless Bio. The other authors declare no conflicts of interest.
Additional information
Peer review information Nature Genetics thanks David Brocks and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
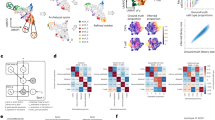
Extended Data Fig. 1 Integrated molecular profiles of patient samples.
Each patient is in a single column with data presented to indicate clinical features (top), followed by genetic alterations defined from bulk whole genome sequencing data, bulk RNA sequencing based subtype classification probabilities (Wang et al., n = 8 available), single-cell RNA tumor cellular state proportions, bulk DNAme microarray subtype classification probabilities (Capper et al.), and boxplots of single-cell DNAme disorder with samples colored by clinical timepoint. Each box spans the 25th and 75th percentile, center lines indicate the median, and the whiskers represent the absolute range (minima/maxima), excluding outliers.
Extended Data Fig. 2 Sample pre-processing and metrics related to single-cell DNAme data assessment.
a, Representative fluorescence activated cell sorting (FACS) data and strategy for viable cell enrichment for both single-cell protocols, and tumor cell enrichment in scRRBS. b, The number of unique CpGs detected per single cell, with the red line indicating the threshold (minimum 40,000 unique CpGs) for inclusion in the dataset presented herein. c, Representative distribution of single locus DNAme estimates for a single cell. DNAme percentage of 0 represents an unmethylated locus, while a percentage of 100 represents a methylated locus. d, The CpG count per genomic features across tumor single cells. e, Histogram representing the cell-to-cell CpG overlap of all single cells in this dataset. f, Upset plot of patient-level unique gene promoter overlap. The top bar plot represents the promoter intersection number measured across all patients (center portion) indicated by a filled bullet point. The right histogram represents the total number of unique promoters measured across all cells from a given tumor with IDH mutation status indicated by color.
Extended Data Fig. 3 Somatic copy number alteration examples estimated from whole genome sequencing, single-cell Reduced Representation Bisulfite Sequencing, and single-cell RNA-sequencing.
a-c, Representative images of copy number alterations derived from SM012 (IDHwt initial) whole genome sequencing (WGS) data. a, Depth ratio for each segment with copy number status determined as compared with germline (normal blood) WGS data. b, SM012 Single-cell DNAme-based copy number estimates (n = 69 tumor cells) with copy number integer depicted by color (blue = CN loss, white = neutral CN, and red = CN gain). Each row is a single cell with annotation for DNAme disorder provided. c, SM012 Single-cell RNAseq based copy number inference (n = 5,489) identifying major copy number events found in WGS with labelled subclones as presented in Fig. 6a. d-f, Similar example profiles as presented in a-c, for tumor sample SM006 (IDHwt initial, n = 82 scRRBS cells, n = 3,310 scRNAseq cells). g-i, Similar example profiles as presented in a-c for tumor sample SM001 (IDHmut recurrence, n = 181 scRRBS cells, n = 5,713 scRNAseq cells).
Extended Data Fig. 4 Distribution and relationship of DNAme and DNAme disorder throughout the glioma genome.
a, Visualization of inter-tumoral and intra-tumoral variation in DNAme (10 kb tiled DNAme). Genome-level and chromosome-level DNAme across 844 single tumor cells. Each row represents a single cell clustered based on pairwise dissimilarity between methylomes as presented in main Fig. 1b and each column represents a single 10 kb tile over which DNAme has been averaged as indicated by heatmap color (methylated = red, unmethylated = blue). The tile color for a cell that does not have a measurement for a given tile is represented by white and a tile without a measurement across any cells is represented by grey. Row annotation both patient identifier and IDH mutation status are presented for each cell. b, Boxplots highlighting the single-cell DNAme disorder estimates calculated across different genomic contexts with Kruskal-Wallis p-values indicating the differences in distributions across the groups. Each box spans the 25th and 75th percentile, center lines indicate the median, and the whiskers represent the absolute range (minima/maxima), excluding outliers. c, The dominant Catalogue of Somatic Mutations in Cancer (COSMIC v3) mutational signatures are presented for each subject. The stacked bar plots represent the relative contribution of each mutational signature to the tumor’s mutational burden. Colors indicate distinct mutational signatures, which are further annotated with their proposed etiology. d, Scatterplots and linear regression lines with standard error showing the relationship between genomic context-specific single-cell DNAme disorder (sample-specific scRRBS average) and genomic context-specific mutation burden derived from whole genome sequencing (n = 10 excluding hypermutant sample). Panels are separated into global (that is, all regions), promoter, gene body, and intergenic regions (Spearman correlations ρ > 0.05 for all comparisons). e, Scatterplot of the context-specific DNAme disorder (x-axis) vs. the average DNAme value (beta-value) for each genomic compartment. Subtype level Spearman correlation coefficients and p-values are presented. f, The median absolute deviation of DNAme across all cells from the same subtype (inter-patient heterogeneity) and g, all cells from the same patient (intra-patient heterogeneity). Two-sided Wilcoxon rank sum tests comparing median absolute deviation levels between IDHmut and IDHwt are presented for intra-patient DNAme heterogeneity.
Extended Data Fig. 5 Association between DNAme disorder and disrupted transcriptional programs.
a, Boxplots of gene expression values, in log(counts per million+1), from single-cell RNAseq data across different sets of gene body regions defined by gene-derived DNAme disorder groups. Each box spans the 25th and 75th percentile, center lines indicate the median, and the whiskers represent the absolute range (minima/maxima), excluding outliers. Surrounding violins represent the distribution for each group. Gene DNAme disorder groups are defined by the determining the mean DNAme disorder value across a single gene. Color indicates IDH1 mutation status. b-c, Scatterplots depicting single-cell gene-level DNAme disorder average plotted against the gene-level methylation estimates in both b, promoter regions and c, gene body regions. d-e, Gene Ontology enrichment analyses with false discovery rate correction for high DNAme genes and low DNAme disorder genes using gene body estimates. f, Representative density curves of distribution of epiallele CpGs in each patient overlapping a specific TFBS motif (for example, SOX2), with curves annotated by patient identifier. g, Gene Ontology enrichment analysis of TFs with high DNAme disorder in their binding sites with false discovery rate correction. h, Scatterplot depicting the association between average single-cell DNAme disorder estimate and single-sample Gene Set Enrichment Score for stress response, hypoxia, and random genes from bulk RNAseq data. Spearman correlation coefficient and p-values are indicated.
Extended Data Fig. 6 Pan-glioma cell state assignment and characteristics.
a, UMAP dimensionality reduction plot of all scRNAseq data, including tumor and non-tumor cells (n = 55,248 cells). Each dot depicts a single cell and colors represent the tumor of origin. Shaded regions represent cell state classification. b, Stacked violin plots of average single-cell gene expression for cells presented in Supplementary Fig. 6a. Selected genes presented are informative for cell state classification. c, Stacked bar plots representing the proportion of non-tumor cellular states d, Stacked bar plots representing the proportion of tumor cellular states per tumor for pan-glioma classification (top row) and previously published classifications (lower left row; Venteicher et al. and lower right Neftel et al.) e, Sankey plot representing the proportion of IDHmut tumor cells with pan-glioma classification and associated classification described in Venteicher et al. (left). Sankey plot representing the proportion of IDHwt tumor cells with pan-glioma classification and associated classification described in Neftel et al.(right). f, scRNAseq area under the curve estimates for selected gene sets (that is, proportion of expressed genes in signature per cell). The AUC estimates are presented for response to stress, hypoxia, and random gene set signatures summarized by pan-glioma cell state and separated by IDH mutation subtype. All cells from a single patient are normalized to its median AUC value for a given signature. Higher relative values indicate greater enrichment score for each signature. P-values represent two-sided Wilcoxon rank sum tests comparing differentiated-like tumor cells with stem-like and proliferating stem-like. g, Density plots representing TFBS motif DNAme disorder (scRRBS data) in IDHmut (left) and IDHwt (right) single-cell DNAme data for TFs whose activity (scRNAseq based SCENIC analysis in Fig. 6c,d) characterizes a specific cell state (n = 15 TFs per cell state). Kolmogorov-Smirnov p-value tests for differences in TFBS DNAme disorder across the cellular states. Dotted lines represent the median TFBS motif DNAme disorder value for cell state defining TFs.
Extended Data Fig. 7 LIGER integrated tumor-specific clustering of single-cell RNA and single-cell DNAme data.
a, Schematic diagram representing LIGER workflow to jointly cluster single-cell RNAseq and DNAme data generated from the same tumor dissociation. b, Joint single-cell RNAseq (scRNA) and single-cell DNAme (scDNAm) clustering and UMAP projections highlighting similar cellular state distributions across platforms. Sample annotation is presented on the left of each paired UMAP plot, each dot is an individual single cell, and cell number for each technology is presented in the lower-left hand corner. UMAP coordinate space remains the same for both scRNA and scDNAm visualizations with cellular states for that platform represented by a colored dot and data for the other platform represented by a gray dot. Stacked bar plots enumerating the proportion of cellular states detected by each platform are presented to the right of each paired UMAP plot. ‘*‘ indicate specimens in which the cellular proportions across the two platforms are significantly different (two-sided Fisher’s Exact test, p < 0.05). c, Promoter DNAme for samples with sufficient number of cells in each state. Each box spans the 25th and 75th percentile, center lines indicate the median, and the whiskers represent the absolute range (minima/maxima), excluding outliers. Surrounding violins represent the distribution for each condition. Two-sided Wilcoxon rank sum test p-values are presented for each tumor.
Extended Data Fig. 8 Stress-associated changes in DNAme disorder are associated with altered population-level transcriptional dynamics and not related with genetic changes.
a, Relative gene expression levels for two patient-derived glioma sphere-forming cells for candidate gene cell state (SOX2, POU5F1) and cell stress (JUN, EPAS1 (HIF2A), VEGFA) via RT-PCR. Normoxia and varying levels of hypoxia (2% and 1% oxygen, n = 4 per group) were assessed. Statistical significance (p < 0.05, Tukey HSD) is indicated by an asterisk. b, Relative DNAme disorder in hypoxia conditions (2% and 1%) compared with normoxia. P-values for Kruskal-Wallis tests are presented across specific genomic contexts (n = 4 per group). c, Upset plot of shared mutations for a randomly selected replicate from cell line HF3016 cultured under normoxia and irradiation (10 Gy). Mutations were determined in reference to patient normal blood. The mutational overlap is presented by the black bar with the mutations called private to irradiation and control also presented. d, Heatmap representing transcription factors that were determined to have consistently different TFBS motif DNAme disorder levels in stress conditions (hypoxia and irradiation) compared with controls across both cell lines (p < 0.1 two-sided Wilcoxon rank sum test across all cell lines and two stressors) are presented with their change in inferred TF activity (SCENIC, methods). e-f, ELK4 and TFDP1 are presented for TFBS motif DNAme disorder (RRBS) and TF activity (scRNAseq), which demonstrated consistent changes in TFBS motif DNAme disorder and stress altered TF activity. Two-sided Wilcoxon rank sum test p-values are presented. g-h, scRNAseq scaled gene expression heatmaps for the top 5 differentially expressed genes per stress exposure and time point. i-j, Stacked bar plots comparing the cell state proportions for the Neftel et al. proliferation-independent IDHwt classifier across different stress conditions, time points, and cell lines. Statistical differences are presented for Chi-Square test (*** = p < 0.001). Oligodendrocyte progenitor cell-like (OPC-like), Neural progenitor cell-like (NPC-like), Mesenchymal-like (MES-like), and Astrocyte-like (AC-like) cell states are presented.
Extended Data Fig. 9 Whole genome sequencing phylogenetic inference of tumor samples.
a, Stacked bar plots representing the proportion of whole-genome sequencing (WGS) derived somatic copy number alteration (SCNA) burden attributed to clonal vs. subclonal events. b-i, Phylogenetic trees constructed from whole genome sequencing data (mutations and somatic copy number alterations) using phyloWGS and further annotated using single-cell inferred copy number alterations (scRRBS + scRNAseq). Tree nodes represent alterations specific to the given clone, with node size corresponding to the fraction of cells with the associated alterations. Branch length scales with the number of mutations attributed to that clone. Clonal alterations are colored in blue, with subclonal alterations colored in gold. Genes considered significantly mutated in TCGA analyses2 and chromosomal arm-level events are presented. Arm-level events are defined as spanning at least 80 percent of the chromosome arm, while partial events span at least 40 percent.
Extended Data Fig. 10 Genetic influences on epigenetic and transcriptional diversity in glioma cells.
a-c, SCNA phylogenetic trees annotated with scRRBS-derived cell state. Adjacent boxplots are presented for DNAme and DNAme disorder across cuts in the dendrograms. d-e, Extrachromosomal DNA circular amplicon reconstruction displaying genomic rearrangements predicted from whole genome sequencing. Coverage depth is represented as a histogram across a genomic interval with segment copy number (CN) estimation provided on the right y-axis. Discordant read pair clusters are indicated by arcs and colors highlight read pair orientation (for example, brown = everted read pairs61). Amplicon intervals are provided at the bottom of the plot with annotation for known oncogenes (for example, EGFR). f, EGFR copy number estimation from single-cell RRBS data in ecDNA+ tumors. Cells with EGFR copy number greater than 6 were classified as EGFR ecDNA+ (blue). g, Single-cell 10-kb tiled DNAme separated by EGFR ecDNA status. Single cells with inferred copy number status greater than 6 were classified as ecDNA+ (blue). Two-sided Wilcoxon rank sum test p-values comparing DNAme across ecDNA status are reported for each patient tumor. h, Boxplots depicting transcriptional diversity using gene count signatures calculated in scRNAseq data for each tumor, with cells separated based on inferred EGFR copy number status (gray = EGFR ecDNA-, blue = EGFR ecDNA+). Transcriptional diversity was compared based on predicted ecDNA status within each tumor subclone. Stars (*) indicate statistically significant differences based on two-sided Wilcoxon rank sum test (p < 0.05). Each box plot in this figure spans the 25th and 75th percentile, center lines indicate the median, and the whiskers represent the absolute range (minima/maxima), excluding outliers. Surrounding violins represent the distribution for each condition.
Supplementary information
Rights and permissions
About this article
Cite this article
Johnson, K.C., Anderson, K.J., Courtois, E.T. et al. Single-cell multimodal glioma analyses identify epigenetic regulators of cellular plasticity and environmental stress response. Nat Genet 53, 1456–1468 (2021). https://doi.org/10.1038/s41588-021-00926-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00926-8
This article is cited by
-
Revealing the role of SPP1+ macrophages in glioma prognosis and therapeutic targeting by investigating tumor-associated macrophage landscape in grade 2 and 3 gliomas
Cell & Bioscience (2024)
-
Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Nature Communications (2024)
-
Hypoxia drives shared and distinct transcriptomic changes in two invasive glioma stem cell lines
Scientific Reports (2024)
-
Machine learning-based investigation of regulated cell death for predicting prognosis and immunotherapy response in glioma patients
Scientific Reports (2024)
-
A multidimensional atlas of human glioblastoma-like organoids reveals highly coordinated molecular networks and effective drugs
npj Precision Oncology (2024)