Abstract
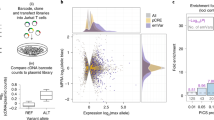
We report the largest and most diverse genetic study of type 1 diabetes (T1D) to date (61,427 participants), yielding 78 genome-wide-significant (P < 5 × 10−8) regions, including 36 that are new. We define credible sets of T1D-associated variants and show that they are enriched in immune-cell accessible chromatin, particularly CD4+ effector T cells. Using chromatin-accessibility profiling of CD4+ T cells from 115 individuals, we map chromatin-accessibility quantitative trait loci and identify five regions where T1D risk variants co-localize with chromatin-accessibility quantitative trait loci. We highlight rs72928038 in BACH2 as a candidate causal T1D variant leading to decreased enhancer accessibility and BACH2 expression in T cells. Finally, we prioritize potential drug targets by integrating genetic evidence, functional genomic maps and immune protein–protein interactions, identifying 12 genes implicated in T1D that have been targeted in clinical trials for autoimmune diseases. These findings provide an expanded genomic landscape for T1D.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All univariable summary statistics for genotype association with T1D (including imputed variants) are available through the NHGRI-EBI GWAS catalog (GCST90013445 and GCST90013446). The caQTL summary statistics are available through the Type 1 Diabetes Knowledge Portal (https://t1d.hugeamp.org).
Publicly available ATAC-seq: Raw FASTQ files were obtained from Gene Expression Omnibus (GEO) accession number GSE118189. These data included four individuals and 25 immune-cell types under resting conditions as well as after stimulation with anti-human CD3/CD28 Dynabeads and human IL-2 (for 24 h; T lymphocytes), F(ab)′2 anti-human IgG/IgM38 and human IL-4 (for 24 h; B lymphocytes), human IL-2 (for 48 h; natural killer cells) or lipopolysaccharide (for 6 h; monocytes)37.
The ATAC-seq data from the pancreatic islets of five donors without glucose intolerance and five EndoCβH1 cell line replicates, under resting conditions and after stimulation with IFN-γ and IL-1β for 48 h, were downloaded from the GEO (accession number GSE123404)35.
The ATAC-seq data from cardiac fibroblasts (two fetal and three adult) were downloaded from the European Nucleotide Archive (https://www.ebi.ac.uk/ena/data/view/SRX2843570 and https://www.ebi.ac.uk/ena/data/view/SRX2843571) as a control cell type that we did not expect to be involved in the etiology of T1D40.
Epigenome annotation tracks: We obtained chromHMM100 tracks from diverse primary human cells from the NIH Epigenome Roadmap, http://dcc.blueprint-epigenome.eu/#/md/secondary_analysis/Segmentation_of_ChIP-Seq_data_20140811, and additional immune-specific human primary and cell lines from the Blueprint consortium, https://egg2.wustl.edu/roadmap/data/byFileType/chromhmmSegmentations/ChmmModels/coreMarks/jointModel/final.
Whole-blood eQTL summary statistics: Summary statistics from whole-blood cis-eQTL analysis from 31,683 individuals42 were downloaded from https://eqtlgen.org.
Additional databases used in the priority-index drug target prioritization analysis were obtained through the relational database provided in the R package Pi (http://pi.well.ox.ac.uk:3010/download).
Code availability
Code used to generate the results presented in this paper is available at https://github.com/ccrobertson/t1d-immunochip-2020. The pipelines for processing ATAC-seq data are available at https://github.com/dfloresDIL/MEGA and http://pepatac.databio.org.
References
Barrett, J. C. et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat. Genet. 41, 703–707 (2009).
Todd, J. A. et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 39, 857–864 (2007).
Onengut-Gumuscu, S. et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 47, 381–386 (2015).
Inshaw, J. R. J., Walker, N. M., Wallace, C., Bottolo, L. & Todd, J. A. The chromosome 6q22.33 region is associated with age at diagnosis of type 1 diabetes and disease risk in those diagnosed under 5 years of age. Diabetologia 61, 147–157 (2018).
Fortune, M. D. et al. Statistical colocalization of genetic risk variants for related autoimmune diseases in the context of common controls. Nat. Genet. 47, 839–846 (2015).
Evangelou, M. et al. A method for gene-based pathway analysis using genomewide association study summary statistics reveals nine new type 1 diabetes associations. Genet. Epidemiol. 38, 661–670 (2014).
Rewers, M. & Ludvigsson, J. Environmental risk factors for type 1 diabetes. Lancet 387, 2340–2348 (2016).
Sharp, S. A. et al. Development and standardization of an improved type 1 diabetes genetic risk score for use in newborn screening and incident diagnosis. Diabetes Care 42, 200–207 (2019).
Krischer, J. P. et al. Predicting islet cell autoimmunity and type 1 diabetes: an 8-year TEDDY Study progress report. Diabetes Care 42, 1051–1060 (2019).
Onengut-Gumuscu, S. et al. Type 1 diabetes risk in African-ancestry participants and utility of an ancestry-specific genetic risk score. Diabetes Care 42, 406–415 (2019).
Skyler, J. S. Hope vs hype: where are we in type 1 diabetes? Diabetologia 61, 509–516 (2018).
Herold, K. C. et al. An anti-CD3 antibody, teplizumab, in relatives at risk for type 1 diabetes. N. Engl. J. Med. 381, 603–613 (2020).
King, E. A., Davis, J. W. & Degner, J. F. Are drug targets with genetic support twice as likely to be approved? Revised estimates of the impact of genetic support for drug mechanisms on the probability of drug approval. PLoS Genet. 15, e1008489 (2019).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Cooper, J. D. et al. Meta-analysis of genome-wide association study data identifies additional type 1 diabetes risk loci. Nat. Genet. 40, 1399–1401 (2008).
Hakonarson, H. et al. A novel susceptibility locus for type 1 diabetes on Chr12q13 identified by a genome-wide association study. Diabetes 57, 1143–1146 (2008).
Grant, S. F. A. et al. Follow-up analysis of genome-wide association data identifies novel loci for type 1 diabetes. Diabetes 58, 290–295 (2009).
Bradfield, J. P. et al. A genome-wide meta-analysis of six type 1 diabetes cohorts identifies multiple associated loci. PLoS Genet. 7, e1002293 (2011).
Huang, J., Ellinghaus, D., Franke, A., Howie, B. & Li, Y. 1000 Genomes-based imputation identifies novel and refined associations for the Wellcome Trust Case Control Consortium phase 1 Data. Eur. J. Hum. Genet. 20, 801–805 (2012).
Zhu, M. et al. Identification of novel T1D risk loci and their association with age and islet function at diagnosis in autoantibody-positive T1D individuals: based on a two-stage genome-wide association study. Diabetes Care 42, 1414–1421 (2019).
Divers, J. et al. Trends in incidence of type 1 and type 2 diabetes among youths—selected counties and Indian reservations, United States, 2002–2015. Morb. Mortal. Wkly Rep. 69, 161–165 (2020).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Cortes, A. et al. Promise and pitfalls of the Immunochip. Arthritis Res. Ther. 13, 101 (2011).
Ferreira, R. C. et al. Functional IL6R 358Ala allele impairs classical IL-6 receptor signaling and influences risk of diverse inflammatory diseases. PLoS Genet. 9, e1003444 (2013).
Okada, Y., Wu, D., Trynka, G. & Towfique, R. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under depencency. Ann. Stat. 29, 1165–1188 (2001).
Crouch, D. J. M. et al. Enhanced genetic analysis of type 1 diabetes by selecting variants on both effect size and significance, and by integration with autoimmune thyroid disease. Preprint at bioRxiv https://doi.org/10.1101/2021.02.05.429962 (2021).
Wallace, C. et al. Dissection of a complex disease susceptibility region using a Bayesian stochastic search approach to fine mapping. PLoS Genet. 11, e1005272 (2015).
Asimit, J. L. et al. Stochastic search and joint fine-mapping increases accuracy and identifies previously unreported associations in immune-mediated diseases. Nat. Commun. 10, 3216 (2019).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Kichaev, G. & Pasaniuc, B. Leveraging functional-annotation data in trans-ethnic fine-mapping studies. Am. J. Hum. Genet. 97, 260–271 (2015).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Lizio, M. et al. Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol. 16, 22 (2015).
Ramos-rodríguez, M. et al. The impact of proinflammatory cytokines on the β-cell regulatory landscape provides insights into the genetics of type 1 diabetes. Nat. Genet. 51, 1588–1595 (2019).
Ward, L. D. & Kellis, M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 44, 877–881 (2016).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Calderon, D. et al. Landscape of stimulation-responsive chromatin across diverse human immune cells. Nat. Genet. 51, 1494–1505 (2019).
Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc. Natl Acad. Sci. USA 114, 2301–2306 (2017).
Jonsson, M. K. B. et al. A transcriptomic and epigenomic comparison of fetal and adult human cardiac fibroblasts reveals novel key transcription factors in adult cardiac fibroblasts. JACC Basic Transl. Sci. 1, 590–602 (2016).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Võsa, U. et al. Unraveling the polygenic architecture of complex traits using blood eQTL meta-analysis. Preprint at bioRxiv https://doi.org/10.1101/447367 (2018).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 (2016).
Schmiedel, B. J. et al. Impact of genetic polymorphisms on human immune cell gene expression. Cell 175, 1701–1715 (2018).
Westra, H. J. et al. Fine-mapping and functional studies highlight potential causal variants for rheumatoid arthritis and type 1 diabetes. Nat. Genet. 50, 1366–1374 (2018).
Fang, H. et al. A genetics-led approach defines the drug target landscape of 30 immune-related traits. Nat. Genet. 51, 1082–1091 (2019).
Chun, S. et al. Limited statistical evidence for shared genetic effects of eQTLs and autoimmune-disease-associated loci in three major immune-cell types. Nat. Genet. 49, 600–605 (2017).
Hukku, A. et al. Probabilistic colocalization of genetic variants from complex and molecular traits: promise and limitations. Am. J. Hum. Genet. 108, 25–35 (2021).
Chiou, J. et al. Interpreting type 1 diabetes risk with genetics and single-cell epigenomics. Nature https://doi.org/10.1038/s41586-021-03552-w (2021).
Benaglio, P. et al. Mapping genetic effects on cell type-specific chromatin accessibility and annotating complex trait variants using single nucleus ATAC-seq. Preprint at bioRxiv https://doi.org/10.1101/2020.12.03.387894 (2020).
Kundu, K. et al. Genetic associations at regulatory phenotypes improve fine-mapping of causal variants for twelve immune-mediated diseases. Preprint at bioRxiv https://doi.org/10.1101/2020.01.15.907436 (2020).
Danko, C. G. et al. Dynamic evolution of regulatory element ensembles in primate CD4+ T cells. Nat. Ecol. Evol. 2, 537–548 (2018).
Tsukumo, S. et al. Bach2 maintains T cells in a naive state by suppressing effector memory-related genes. Proc. Natl Acad. Sci. USA 110, 10735–10740 (2013).
Roychoudhuri, R. et al. BACH2 regulates CD8+ T cell differentiation by controlling access of AP-1 factors to enhancers. Nat. Immunol. 17, 851–860 (2016).
Afzali, B. et al. BACH2 immunodeficiency illustrates an association between super-enhancers and haploinsufficiency. Nat. Immunol. 18, 813–823 (2017).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Faegan, B. G. et al. Risankizumab in patients with moderate to severe Crohn’s disease: an open-label extension study. Lancet Gastroenterol. Hepatol. 3, 671–680 (2018).
Fotiadou, C., Lazaridou, E., Sotiriou, E. & Ioannides, D. Targeting IL-23 in psoriasis: current perspectives. Psoriasis Targets Ther. 8, 1–5 (2018).
Wollenhaupt, J. et al. Safety and efficacy of tofacitinib for up to 9.5 years in the treatment of rheumatoid arthritis: final results of a global, open-label, long-term extension study. Arthritis Res. Ther. 21, 89 (2019).
Sandborn, W. J. et al. Tofacitinib as induction and maintenance therapy for ulcerative colitis. N. Engl. J. Med. 376, 1723–1736 (2017).
Gaglia, J. & Kissler, S. Anti-CD3 antibody for the prevention of type 1 diabetes: a story of perseverance. Biochemistry 58, 4107–4111 (2019).
Aylward, A., Chiou, J., Okino, M.-L., Kadakia, N. & Gaulton, K. J. Shared genetic risk contributes to type 1 and type 2 diabetes etiology. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddy314 (2018).
Dooley, J. et al. Genetic predisposition for beta cell fragility underlies type 1 and type 2 diabetes. Nat. Genet. 48, 519–527 (2016).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Price, A. L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 127–147 (2008).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Spielman, R. S., McGinnis, R. E. & Ewens, W. J. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am. J. Hum. Genet. 52, 506–516 (1993).
Taub, M. A., Schwender, H., Beaty, T. H., Louis, T. A. & Ruczinski, I. Incorporating genotype uncertainties into the genotypic TDT for main effects and gene-environment interactions. Genet. Epidemiol. 36, 225–234 (2012).
Kazeem, G. R. & Farrall, M. Integrating case-control and TDT studies. Ann. Hum. Genet. 69, 329–335 (2005).
Bottolo, L. & Richardson, S. Evolutionary stochastic search for Bayesian model exploration. Bayesian Anal. 5, 583–618 (2010).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Burren, O. S. et al. Chromosome contacts in activated T cells identify autoimmune disease candidate genes. Genome Biol. 18, 165 (2017).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Harrow, J. et al. GENCODE: the reference human genome annotation for the ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Gaspar, J. M. Improved peak-calling with MACS2. Preprint at bioRxiv https://doi.org/10.1101/496521 (2018).
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Trynka, G. et al. Disentangling the effects of colocalizing genomic annotations to functionally prioritize non-coding variants within complex-trait loci. Am. J. Hum. Genet. 97, 139–152 (2015).
Smith, J. P. et al. PEPATAC: an optimized ATAC-seq pipeline with serial alignments. Preprint at bioRxiv https://doi.org/10.1101/2020.10.21.347054 (2020).
Jiang, H., Lei, R., Ding, S. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform. 15, 182 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Robinson, M. D., Mccarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Fort, A. et al. MBV: a method to solve sample mislabeling and detect technical bias in large combined genotype and sequencing assay datasets. Bioinformatics 33, 1895–1897 (2017).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Liu, B., Gloudemans, M. J., Rao, A. S., Ingelsson, E. & Montgomery, S. B. Abundant associations with gene expression complicate GWAS follow-up. Nat. Genet. 51, 768–769 (2019).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014).
Fairfax, B. P. et al. Genetics of gene expression in primary immune cells identifies cell type-specific master regulators and roles of HLA alleles. Nat. Genet. 44, 502–510 (2012).
Andiappan, A. K. et al. Genome-wide analysis of the genetic regulation of gene expression in human neutrophils. Nat. Commun. 6, 7971 (2015).
Kasela, S. et al. Pathogenic implications for autoimmune mechanisms derived by comparative eQTL analysis of CD4+ versus CD8+ T cells. PLoS Genet. 13, e1006643 (2017).
Westra, H. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368 (2017).
R Core Team. R: A Language and Environment for Computing, https://www.R-project.org/ (R Foundation for Statistical Computing, 2020).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–2492 (2017).
Acknowledgements
We thank the investigators and their studies for contributing samples and/or data to the current work, and the participants in those studies who made this research possible. These studies include the T1DGC, British 1958 Birth Cohort, Genetic Resource Investigating Diabetes (GRID), Consortium for the Longitudinal Evaluation of African-Americans with Early Rheumatoid Arthritis (CLEAR), Epidemiology of Diabetes Interventions and Complications (EDIC), Genetics of Kidneys and Diabetes Study (GoKinD), New York Cancer Project (NYCP), SEARCH for Diabetes in Youth study (SEARCH), Type 1 Diabetes TrialNet study (TrialNet), Tyypin 1 Diabetekseen Sairastuneita Perheenjäsenineen (IDDMGEN), Tyypin 1 Diabeteksen Genetiikka (T1DGEN), Northern Ireland GRID Collection, Northern Ireland Young Hearts Project, Hvidoere Study Group on Childhood Diabetes (HSG) and International HapMap Project. Additional institutions contributing samples are: British Diabetes Association (BDA), NIHR Cambridge BioResource, UK Blood Service (UKBS), Benaroya Research Institute (BRI), National Institute of Mental Health (NIMH), University of Alabama at Birmingham (UAB), University of Colorado, University of California San Francisco (UCSF), Medical College of Wisconsin (MCW) and Steno Diabetes Center. Samples and data from the T1DGC, EDIC and GoKinD can be obtained from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) Central Repository. This research utilizes resources provided by the T1DGC, a collaborative clinical study sponsored by the NIDDK, National Institute of Allergy and Infectious Diseases (NIAID), National Human Genome Research Institute (NHGRI), National Institute of Child Health and Human Development (NICHD) and Juvenile Diabetes Research Foundation (JDRF) and is supported by grant no. U01 DK062418 to S.S.R. The generation of chromatin-accessibility data on T1DGC samples was supported by grants from the NIDDK (grant nos DP3 DK111906 to S.S.R. and R01 DK115694 to P.C.). Further support was provided by the NIAID (grant no. P01 AI042288 to M.A.A.). The JDRF/Wellcome Diabetes and Inflammation Laboratory was supported by grants from the JDRF (grant no. 4-SRA-2017-473-A-A) and the Wellcome Trust (grant no. 107212/A/15/Z). Computation used the Oxford Biomedical Research Computing (BMRC) facility, a joint development between the Wellcome Centre for Human Genetics and the Big Data Institute supported by Health Data Research UK and the NIHR Oxford Biomedical Research Centre. Financial support was provided by a Wellcome Core Award (grant no. 203141/Z/16/Z). The views expressed are those of the author(s) and not necessarily those of the NHS, NIHR or Department of Health. While working on this project, C.C.R. was supported by a training grant from the US National Library of Medicine (grant no. T32 LM012416) and the Wagner Fellowship from the UVA. This work made use of data and samples generated by the 1958 Birth Cohort (NCDS), which is managed by the Centre for Longitudinal Studies at the UCL Institute of Education, funded by the Economic and Social Research Council (grant no. ES/M001660/1). Access to these resources was enabled via the MRC and Wellcome: 58FORWARDS grant no. 108439/Z/15/Z (The 1958 Birth Cohort: Fostering new Opportunities for Research via Wider Access to Reliable Data and Samples). Before 2015, biomedical resources were maintained under the Wellcome and Medical Research Council 58READIE Project (grant nos WT095219MA and G1001799). We acknowledge use of DNA samples from the NIHR Cambridge BioResource. We thank volunteers for their support and participation in the Cambridge BioResource and members of the Cambridge BioResource Scientific Advisory Board and Management Committee for their support of our study. We thank the NIHR Cambridge Biomedical Research Centre for funding. Access to Cambridge BioResource volunteers and their data and samples are governed by the Cambridge BioResource Scientific Advisory Board. Documents describing access arrangements and contact details are available at http://www.cambridgebioresource.org.uk/. The ethics for GRID were processed by the NRES Committee East of England Cambridge South MREC 00/5/44. We thank the following CLEAR investigators who performed recruiting: D. Conn (Grady Hospital and Emory University), B. Jonas and L. Callahan (University of North Carolina at Chapel Hill), E. Smith (Medical University of South Carolina), R. Brasington (Washington University) and L. W. Moreland (University of Pittsburgh). The CLEAR Registry and Repository was funded by the NIH Office of the Director (grant nos N01-AR-0-2247 (30 September 2000–29 September 2006) and N01-AR-6-2278 (30 September 2006–31 March 2012); S.L.B. Jr, principal investigator). Bio-samples and/or data for this publication were obtained from NIMH Repository and Genomics Resource, a centralized national biorepository for genetic studies of psychiatric disorders. The SEARCH for Diabetes in Youth Study (www.searchfordiabetes.org) is indebted to the many youth and their families, as well as their healthcare providers, whose participation made this study possible. SEARCH for Diabetes in Youth is funded by the Centers for Disease Control and Prevention (PA numbers 00097, DP-05-069 and DP-10-001) and supported by the NIDDK. The SEARCH site contract numbers are: Kaiser Permanente Southern California, U48/CCU919219, U01 DP000246 and U18DP002714; University of Colorado Denver, U48/CCU819241-3, U01 DP000247 and U18DP000247-06A1; Children’s Hospital Medical Center (Cincinnati), U48/CCU519239, U01 DP000248 and 1U18DP002709; University of North Carolina at Chapel Hill, U48/CCU419249, U01 DP000254 and U18DP002708; University of Washington School of Medicine, U58/CCU019235-4, U01 DP000244 and U18DP002710-01; and Wake Forest University School of Medicine, U48/CCU919219, U01 DP000250 and 200-2010-35171. We acknowledge the support of the TrialNet group (https://www.trialnet.org), which identified study participants and provided samples and follow-up data for this study. The TrialNet group is a clinical trials network funded by the NIH through the NIDDK, NIAID and The Eunice Kennedy Shriver National Institute of Child Health and Human Development—through the cooperative agreements U01 DK061010, U01 DK061016, U01 DK061034, U01 DK061036, U01 DK061040, U01 DK061041, U01 DK061042, U01 DK061055, U01 DK061058, U01 DK084565, U01 DK085453, U01 DK085461, U01 DK085463, U01 DK085466, U01 DK085499, U01 DK085505 and U01 DK085509—and the JDRF. The contents of this article are solely the responsibility of the authors and do not necessarily represent the official views of the NIH or JDRF. Further support was provided by grants from the NIDDK (grant nos U01 DK103282 and U01 DK127404 to C.J.G.). DNA samples from the UAB were recruited, in part, with the support of grant nos P01-AR49084, UL1-TR001417 and UL1-TR003096 (to R.P.K.). We acknowledge the involvement of the Barbara Davis Center for Diabetes at the University of Colorado, supported by the following grants from the NIH NIDDK to M.J.R.: DRC P30 DK116073 and R01 DK032493. The collection of DNA samples at UCSF was supported by grant funding from the National Multiple Sclerosis Society (grant no. SI-2001-35701 to J.R.O.). Whole-genome-sequencing data production and variant calling was funded by an NHGRI Center for Common Disease Genomics award to Washington University in St. Louis (grant no. UM1 HG008853). This study used the TOPMed program imputation panel (version TOPMed-r2) supported by the NHLBI (www.nhlbiwgs.org). The TOPMed study investigators contributed data to the reference panel, which can be accessed through the Michigan Imputation Server (https://imputationserver.sph.umich.edu). The panel was constructed and implemented by the TOPMed Informatics Research Center at the University of Michigan (3R01HL-117626-02S1; contract HHSN268201800002I). The TOPMed Data Coordinating Center (3R01HL-120393-02S1; contract HHSN268201800001I) provided additional data management, sample identity checks and overall program coordination and support. We thank the studies and participants who provided biological samples and data for TOPMed. The individual members of the T1DGC and the SEARCH for Diabetes in Youth Study are listed in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
This study was conceptually designed by P.C., J.A.T. and S.S.R. The study was implemented by S.O.-G., P.C., J.A.T. and S.S.R. DNA samples for genotyping were managed by S.O.-G and E.F. L.S.W. contributed to data interpretation. D.J.M.C. provided statistical advice. Frozen T1DGC peripheral blood mononuclear cell samples for chromatin-accessibility profiling (ATAC-seq) were managed by P.C. and S.O.-G. ATAC-seq data generation at the UVA was led by S.O.-G. The generation of ATAC-seq data at the University of Oxford was led by A.J.C. Genotype data processing, quality control, imputation and statistical analyses were performed by W.-M.C., S.O.-G., J.R.J.I. and C.C.R. The chromatin-accessibility data processing and analysis was performed by A.J.C., D.F.S.C., J.R.J.I. and C.C.R. The EMSAs were performed by H.Y., with supervision from S.O.-G. D.B.D. provided samples for genotyping through the GRID. P.D. provided ImmunoChip genotyping data through the UKBS. J.H.B. provided samples for genotyping and data from the BRI. S.L.B. Jr provided samples for genotyping through the CLEAR consortium. P.K.G. provided samples for genotyping through the NYCP project. J.D., D.D., J.M.L., S.M. and A.S.S. provided samples for genotyping and data through the SEARCH for Diabetes in Youth study (SEARCH). C.J.G. and M.A.A. provided samples for genotyping through TrialNet. R.P.K., J.C.E., M.J.R., A.K.S., J.R.O. and F.P. provided samples for genotyping through their affiliated institutions and research programs. The manuscript was written by J.R.J.I. (under supervision by J.A.T.) and C.C.R. (under supervision by S.S.R.). All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Fine mapping of the chromosome 6q22.32 region.
European (EUR, top panel) and African (AFR, middle panel) ancestry group association z-score statistics and posterior probabilities (bottom panel) from multi-ethnic fine mapping of EUR and AFR using PAINTOR. z-scores are colored by linkage disequilibrium (LD) to the lead PAINTOR-prioritized variant.
Extended Data Fig. 2 Fine mapping of the chromosome 18q22.2 region.
European (EUR, top panel) and African (AFR, middle panel) ancestry group association z-score statistics and posterior probabilities (bottom panel) from multi-ethnic fine mapping of EUR and AFR using PAINTOR. z-scores are colored by linkage disequilibrium (LD) to the lead PAINTOR-prioritized variant.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 and Note
Supplementary Tables
Supplementary Tables 1–16
Rights and permissions
About this article
Cite this article
Robertson, C.C., Inshaw, J.R.J., Onengut-Gumuscu, S. et al. Fine-mapping, trans-ancestral and genomic analyses identify causal variants, cells, genes and drug targets for type 1 diabetes. Nat Genet 53, 962–971 (2021). https://doi.org/10.1038/s41588-021-00880-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00880-5
This article is cited by
-
The immunology of type 1 diabetes
Nature Reviews Immunology (2024)
-
Priority index for critical Covid-19 identifies clinically actionable targets and drugs
Communications Biology (2024)
-
Lessons and Applications of Omics Research in Diabetes Epidemiology
Current Diabetes Reports (2024)
-
Role of the Neanderthal Genome in Genetic Susceptibility to COVID-19: 3p21.31 Locus in the Spotlight
Biochemical Genetics (2024)
-
Shared and distinct genetics of pure type 1 diabetes and type 1 diabetes with celiac disease, homology in their auto-antigens and immune dysregulation states: a study from North India
Acta Diabetologica (2024)