Abstract
Single-nucleus assay for transposase-accessible chromatin using sequencing (snATAC-seq) creates new opportunities to dissect cell type–specific mechanisms of complex diseases. Since pancreatic islets are central to type 2 diabetes (T2D), we profiled 15,298 islet cells by using combinatorial barcoding snATAC-seq and identified 12 clusters, including multiple alpha, beta and delta cell states. We cataloged 228,873 accessible chromatin sites and identified transcription factors underlying lineage- and state-specific regulation. We observed state-specific enrichment of fasting glucose and T2D genome-wide association studies for beta cells and enrichment for other endocrine cell types. At T2D signals localized to islet-accessible chromatin, we prioritized variants with predicted regulatory function and co-accessibility with target genes. A causal T2D variant rs231361 at the KCNQ1 locus had predicted effects on a beta cell enhancer co-accessible with INS and genome editing in embryonic stem cell–derived beta cells affected INS levels. Together our findings demonstrate the power of single-cell epigenomics for interpreting complex disease genetics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw sequencing data have been deposited into the National Center for Biotechnology Information Gene Expression Omnibus with accession numbers GSE160472, GSE160473 and GSE163610. Processed data files and annotations for snATAC-seq are available through the Diabetes Epigenome Atlas (https://www.diabetesepigenome.org/). All other data are either found in the article or available upon request to the corresponding author. Source data are provided with this paper.
Code availability
The code for processing and clustering the snATAC-seq datasets is available at https://github.com/kjgaulton/pipelines/tree/master/islet_snATAC_pipeline.
References
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Litzenburger, U. M. et al. Single-cell epigenomic variability reveals functional cancer heterogeneity. Genome Biol. 18, 15 (2017).
Buenrostro, J. D. et al. Integrated single-cell analysis maps the continuous regulatory landscape of human hematopoietic differentiation. Cell 173, 1535–1548.e16 (2018).
Ulirsch, J. C. et al. Interrogation of human hematopoiesis at single-cell and single-variant resolution. Nat. Genet. 51, 683–693 (2019).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell 71, 858–871.e8 (2018).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Wood, A. R. et al. A genome-wide association study of IVGTT-based measures of first-phase insulin secretion refines the underlying physiology of type 2 diabetes variants. Diabetes 66, 2296–2309 (2017).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Manning, A. K. et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Scott, R. A. et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat. Genet. 44, 991–1005 (2012).
Thurner, M. et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at type 2 diabetes susceptibility loci. eLife 7, e31977 (2018).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet. 42, 255–259 (2010).
Gaulton, K. J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat. Genet. 46, 136–143 (2014).
van der Meulen, T. et al. Urocortin3 mediates somatostatin-dependent negative feedback control of insulin secretion. Nat. Med. 21, 769–776 (2015).
Caicedo, A. Paracrine and autocrine interactions in the human islet: more than meets the eye. Semin. Cell Dev. Biol. 24, 11–21 (2013).
DiGruccio, M. R. et al. Comprehensive alpha, beta and delta cell transcriptomes reveal that ghrelin selectively activates delta cells and promotes somatostatin release from pancreatic islets. Mol. Metab. 5, 449–458 (2016).
Dorrell, C. et al. Human islets contain four distinct subtypes of β cells. Nat. Commun. 7, 11756 (2016).
Xin, Y. et al. Pseudotime ordering of single human β-cells reveals states of insulin production and unfolded protein response. Diabetes 67, 1783–1794 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Bader, E. et al. Identification of proliferative and mature β-cells in the islets of Langerhans. Nature 535, 430–434 (2016).
Liu, J. S. E. & Hebrok, M. All mixed up: defining roles for β-cell subtypes in mature islets. Genes Dev. 31, 228–240 (2017).
Greenwald, W. W. et al. Pancreatic islet chromatin accessibility and conformation reveals distal enhancer networks of type 2 diabetes risk. Nat. Commun. 10, 2078 (2019).
Khetan, S. et al. Type 2 diabetes-associated genetic variants regulate chromatin accessibility in human islets. Diabetes 67, 2466–2477 (2018).
Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc. Natl Acad. Sci. USA 114, 2301–2306 (2017).
Baron, M. et al. A single-cell transcriptomic map of the human and mouse pancreas reveals inter- and intra-cell population structure. Cell Syst. 3, 346–360.e4 (2016).
Galvagni, F. et al. CD93 and dystroglycan cooperation in human endothelial cell adhesion and migration adhesion and migration. Oncotarget 7, 10090–10103 (2016).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D1284 (2018).
Wilson, M. E., Scheel, D. & German, M. S. Gene expression cascades in pancreatic development. Mech. Dev. 120, 65–80 (2003).
Conrad, E. et al. The MAFB transcription factor impacts islet α-cell function in rodents and represents a unique signature of primate islet β-cells. Am. J. Physiol. Endocrinol. Metab. 310, E91–E102 (2016).
Katoh, M. C. MafB is critical for glucagon production and secretion in mouse pancreatic α cells in vivo. Mol. Cell. Biol. 38, e00504-17 (2018).
Nishimura, W., Takahashi, S. & Yasuda, K. MafA is critical for maintenance of the mature beta cell phenotype in mice. Diabetologia 58, 566–574 (2015).
Ozato, K., Tailor, P. & Kubota, T. The interferon regulatory factor family in host defense: mechanism of action. J. Biol. Chem. 282, 20065–20069 (2007).
De Val, S. & Black, B. L. Transcriptional control of endothelial cell development. Dev. Cell 16, 180–195 (2009).
Segerstolpe, Å. et al. Single-cell transcriptome profiling of human pancreatic islets in health and type 2 diabetes. Cell Metab. 24, 593–607 (2016).
Lawlor, N. et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type-specific expression changes in type 2 diabetes. Genome Res. 27, 208–222 (2017).
Mawla, A. M. & Huising, M. O. Navigating the depths and avoiding the shallows of pancreatic islet cell transcriptomes. Diabetes 68, 1380–1393 (2019).
Camunas-Soler, J. et al. Patch-seq links single-cell transcriptomes to human islet dysfunction in diabetes. Cell Metab. 31, 1017–1031.e4 (2020).
Trapnell, C. et al. Pseudo-temporal ordering of individual cells reveals dynamics and regulators of cell fate decisions. Nat. Biotechnol. 32, 381–386 (2014).
Parker, S. C. J. et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc. Natl Acad. Sci. USA 110, 17921–17926 (2013).
Aylward, A., Chiou, J., Okino, M.-L., Kadakia, N. & Gaulton, K. J. Shared genetic risk contributes to type 1 and type 2 diabetes etiology. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddy314 (2018).
Strawbridge, R. J. et al. Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes. Diabetes 60, 2624–2634 (2011).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Saxena, R. et al. Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nat. Genet. 42, 142–148 (2010).
Wheeler, E. et al. Impact of common genetic determinants of hemoglobin A1c on type 2 diabetes risk and diagnosis in ancestrally diverse populations: a transethnic genome-wide meta-analysis. PLoS Med. 14, e1002383 (2017).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Bentham, J. et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 47, 1457–1464 (2015).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet. 49, 1385–1391 (2017).
Nielsen, J. B. et al. Biobank-driven genomic discovery yields new insight into atrial fibrillation biology. Nat. Genet. 50, 1234–1239 (2018).
Cordell, H. J. et al. International genome-wide meta-analysis identifies new primary biliary cirrhosis risk loci and targetable pathogenic pathways. Nat. Commun. 6, 8019 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Scott, R. A. et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes 66, 2888–2902 (2017).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Lee, D. et al. A method to predict the impact of regulatory variants from DNA sequence. Nat. Genet. 47, 955–961 (2015).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Wingender, E., Schoeps, T., Haubrock, M. & Dönitz, J. TFClass: a classification of human transcription factors and their rodent orthologs. Nucleic Acids Res. 43, D97–D102 (2015).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Schmitt, A. D., Hu, M. & Ren, B. Genome-wide mapping and analysis of chromosome architecture. Nat. Rev. Mol. Cell Biol. 17, 743–755 (2016).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Miguel-Escalada, I. et al. Human pancreatic islet three-dimensional chromatin architecture provides insights into the genetics of type 2 diabetes. Nat. Genet. 51, 1137–1148 (2019).
Jian, X. & Felsenfeld, G. Insulin promoter in human pancreatic β cells contacts diabetes susceptibility loci and regulates genes affecting insulin metabolism. Proc. Natl Acad. Sci. USA 115, E4633–E4641 (2018).
Lawlor, N. et al. Multiomic profiling identifies cis-regulatory networks underlying human pancreatic β cell identity and function. Cell Rep. 26, 788–801.e6 (2019).
Rezania, A. et al. Reversal of diabetes with insulin-producing cells derived in vitro from human pluripotent stem cells. Nat. Biotechnol. 32, 1121–1133 (2014).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Fogarty, M. P., Cannon, M. E., Vadlamudi, S., Gaulton, K. J. & Mohlke, K. L. Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS Genet. 10, e1004633 (2014).
Rusu, V. et al. Type 2 diabetes variants disrupt function of SLC16A11 through two distinct mechanisms. Cell 170, 199–212.e20 (2017).
Carrat, G. R. et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am. J. Hum. Genet. 100, 238–256 (2017).
Claussnitzer, M. et al. Leveraging cross-species transcription factor binding site patterns: from diabetes risk loci to disease mechanisms. Cell 156, 343–358 (2014).
Roman, T. S. et al. A type 2 diabetes-associated functional regulatory variant in a pancreatic islet enhancer at the ADCY5 locus. Diabetes 66, 2521–2530 (2017).
Kycia, I. et al. A common type 2 diabetes risk variant potentiates activity of an evolutionarily conserved islet stretch enhancer and increases C2CD4A and C2CD4B expression. Am. J. Hum. Genet. 102, 620–635 (2018).
Avrahami, D., Klochendler, A., Dor, Y. & Glaser, B. Beta cell heterogeneity: an evolving concept. Diabetologia 60, 1363–1369 (2017).
Modi, H. et al. Ins2 gene bursting activity defines a mature β-cell state. Preprint at bioRxiv https://doi.org/10.1101/702589 (2019).
Farack, L. et al. Transcriptional heterogeneity of beta cells in the intact pancreas. Dev. Cell 48, 115–125.e4 (2019).
Rai, V. et al. Single-cell ATAC-seq in human pancreatic islets and deep learning upscaling of rare cells reveals cell-specific type 2 diabetes regulatory signatures. Mol. Metab. 32, 109–121 (2020).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Arda, H. E. et al. A chromatin basis for cell lineage and disease risk in the human pancreas. Cell Syst. 7, 310–322.e4 (2018).
Ackermann, A. M., Wang, Z., Schug, J., Naji, A. & Kaestner, K. H. Integration of ATAC-seq and RNA-seq identifies human alpha cell and beta cell signature genes. Mol. Metab. 5, 233–244 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Kerpedjiev, P. et al. HiGlass: web-based visual exploration and analysis of genome interaction maps. Genome Biol. 19, 125 (2018).
Raviram, R. et al. 4C-ker: a method to reproducibly identify genome-wide interactions captured by 4C-seq experiments. PLoS Comput. Biol. 12, e1004780 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Velazco-Cruz, L. et al. Acquisition of dynamic function in human stem cell-derived β cells. Stem Cell Rep. 12, 351–365 (2019).
Acknowledgements
This work was supported by NIH grant nos. R01DK114650 and U01DK105554 (sub-award) to K.G., grant nos. R01DK068471 and U01DK105541 to M.S. and grant no. U01DK120429 to K.G. and M.S. and by the University of California, San Diego School of Medicine to the Center for Epigenomics. We thank the QB3 Macrolab at University of California, Berkeley for the purification of the Tn5 transposase. We thank K. Jepsen, the University of California, San Diego Institute for Genomic Medicine Genomics Center and S. Kuan for sequencing and B. Li for bioinformatics support. Data from the UK Biobank was accessed under application no. 24058. We thank I. Matta for the preparation of the RNA-seq libraries.
Author information
Authors and Affiliations
Contributions
K.J.G., D.U.G. and M. Sander conceived and supervised the research. J.C. performed the analyses of the single-cell and genetic data. C.Z., M. Schlichting and J.W. performed the hESC experiments. Z.C. performed the analyses of the single-cell and Hi-C data. J.Y.H. performed the combinatorial barcoding single-cell assays and genotyping. M.M. performed the 10x single-cell assays. R.M. performed the Hi-C experiments. S.H., A.D. and M.-L.O. performed the reporter experiments. Y.Q. performed the analyses of the 4C data. Y. Sui performed the analyses of the hESC data. Y. Sun and P.K. developed and processed the data for the epigenome database. R.F. contributed to the analyses of the single-cell data. S.P. contributed to the development of the single-cell assays. K.J.G., D.U.G., M. Sander, J.C., C.Z. and Z.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.J.G. does consulting work for Genentech and holds stock in Vertex Pharmaceuticals; neither is related to the work in this study. The other authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control metrics and aggregate comparison to bulk islet ATAC.
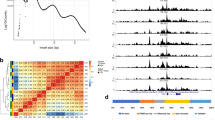
a, Insert size distribution for aggregate reads from each snATAC-seq experiment. b, Aggregated read coverage from each snATAC-seq experiment in a ± 2 kb window around individual promoters (top) and averaged across all promoters (bottom). c, Spearman correlation between normalized read coverage within a merged set of peaks from 3 aggregated islet snATAC-seq, 42 bulk islet ATAC-seq, and 4 bulk pancreas ATAC-seq datasets. Names of samples are from the original sources of the data. d, Binned log10 read depth distribution for each experiment.
Extended Data Fig. 2 Flowchart of the snATAC-seq data processing pipeline.
a, Flowchart summarizing key steps of the snATAC-seq processing pipeline, including the various steps where cells were filtered out. Samples were first processed individually. All samples were then combined using a batch correction method. Clusters corresponding to cells from low quality cells, including those with low read depth in highly variable windows and low fraction of reads in peaks were then removed. After re-clustering, iterative subclustering of the main clusters at high resolution was used to identify and remove doublet subclusters. The final clusters are not driven by potential confounders such as donor of origin. Boxplot center lines, limits, and whiskers represent median, quartiles, and 1.5 IQR respectively.
Extended Data Fig. 3 Analysis of islet single cell gene expression data.
a, log10 transformed read depth or (b) total number of genes expressed compared with number of marker genes expressed per cell from scRNA-seq data. Boxplot center lines, limits, and whiskers represent median, quartiles, and 1.5 IQR respectively. Cells expressing more than one marker gene (defined by mixture models) were marked as doublets and filtered out. c, Clusters of islet cells from single cell RNA-seq data plotted on UMAP coordinates. quies. stellate, quiescent stellate. activ. stellate, activated stellate. d, Selected marker gene log2(expression) for each cluster plotted on UMAP coordinates. e, Row-normalized t-statistics of marker gene specificity showing the most specific genes (t-statistic>20) for each cluster.
Extended Data Fig. 4 Comparison of motif enrichment between alpha and gamma cells.
Differential enrichment of motifs between alpha cell open chromatin regions and gamma cell open chromatin regions as measured by a 2-sided T-test, with FDR calculated by the Benjamini-Hochberg procedure. Examples are highlighted of motifs enriched in alpha cells and gamma cells, respectively (MAFG, HOXA9). UMAP plots show enrichment z-scores for the indicated motifs in alpha and gamma cells. Violin plots below show the distribution of enrichment z-scores across alpha or gamma cells, where the lines represent median and quartiles.
Extended Data Fig. 5 Differentially accessible promoters across pseudo-states.
a, Pseudo-state (trajectory) values for alpha cells plotted on UMAP coordinates (left) and percentage of cells with GCG promoter accessibility decreases across 10 bins along the alpha (α) cell trajectory (right). b, Pseudo-state (trajectory) values for beta (β) cells plotted on UMAP coordinates (left) and percentage of cells with INS promoter accessibility decreases across 10 bins along the beta cell trajectory (right). c, Pseudo-state (trajectory) values for delta (δ) cells plotted on UMAP coordinates (left) and percentage of cells with SST promoter accessibility decreases across 10 bins along the beta cell trajectory (right). d, Heatmaps showing promoters with dynamic accessibility across trajectories for alpha (top), beta (middle) and delta (bottom) cell trajectories. Gene promoters are clustered into 4 groups for each trajectory with k-medoids clustering. Enriched gene ontology for each k-medoid cluster (left) and selected genes present in at least one enriched gene ontology.
Extended Data Fig. 6 Single cell GWAS enrichment and correlation with TF motifs.
a, Single cell GWAS enrichment z-scores for Major depressive disorder and Systemic lupus erythematosus projected onto UMAP coordinates (left panels), z-score enrichment distribution per cell type and state (middle panels) and z-score enrichment distribution split into 10 bins based on beta cell trajectory values (right panels). Boxplot center lines, limits, and whiskers represent median, quartiles, and 1.5 IQR respectively. b, Correlation between single cell GWAS enrichment z-scores for Type 2 Diabetes and chromVAR TF motif enrichment z-scores across either all cells (left) or beta cells (right). Inset scatterplots highlight the top correlated motifs in either direction. c, Variants mapping directly in sequence motifs positively correlated with T2D risk in beta cells are enriched for T2D association, whereas variants mapping in motifs negatively correlated with T2D risk in beta cells show no such enrichment. Values represent effect size and SE.
Extended Data Fig. 7 Single cell co-accessibility analyses in islet cell types.
a, Distance-matched odds that delta cell co-accessibility links overlap islet pcHi-C chromatin loops at different co-accessibility threshold bins in 0.05 intervals demonstrate that co-accessible links are enriched for chromatin interactions. b, Same analysis as in (a) but with alpha cell co-accessibility. c, Same analysis as in (a) but with beta cell co-accessibility and Hi-C loops. d, Same analysis as in (a) but with delta cell co-accessibility and Hi-C loops. e, Same analysis as in (a) but with alpha cell co-accessibility and Hi-C loops. f, Number of distal sites linked to each promoter peak for alpha, beta, and delta cells. g, Number of promoters linked to each distal site for alpha, beta, and delta cells.
Extended Data Fig. 8 Cell type-specific and shared co-accessible sites.
a, An example of co-accessibility anchored at the promoter for the delta cell identity TF HHEX. Co-accessibility for beta, delta, and alpha cells are shown compared to high-confidence pcHi-C loops from ensemble islets. Genome browser plots scale: 0-10. b, An example of co-accessibility anchored at the promoter for the alpha cell identity TF ARX. c, An example of shared co-accessibility anchored at the promoter for the shared islet identity TF NEUROD1.
Extended Data Fig. 9 3D chromatin interactions at the T2D-associated KCNQ1 locus.
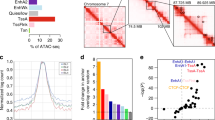
Top panels show Hi-C contact matrices from hESC-derived beta cells, visualized at 25 kb resolution. Region shown is chr11:500,00-4,500,000, hg19. Black arrows indicate putative interaction point of INS TSS and KCNQ1 enhancer. Genome browser plot below shows a zoomed view of chr11:1,750,000-3,250,000. Data from 4C-seq anchored on the INS promoter in EndoC-βH1 cells (Jian & Felsenfeld72) is shown, as analyzed with the 4C-ker package. Normalized read counts are shown in black from 3 biological replicates. Significant interactions from INS promoter are shown as arcs below read counts tracks. Interactions calls from data pooled across 3 replicates are shown here. The region containing the KCNQ1 enhancer was called as a significant interaction region with INS promotor independently in each 4C replicate. Virtual 4C plots in green show log(normalized Hi-C interaction frequency) from INS promoter.
Extended Data Fig. 10 Genome editing of the KCNQ1 locus in hESCs.
a, Schematic of the workflow and (b) Sanger sequencing for KCNQ1 enhancer deletion in three independent hESC clones. c, Representative figures of flow cytometry analysis for NKX6-1 and INS comparing control and KCNQ1ΔEnh cells (left). Quantification of the percentage of NKX6-1+/INS+ cells in beta cell stage cultures from control (n = 6; 2 clones × 3 differentiations) and KCNQ1∆Enh (n = 9; 3 clones × 3 differentiations) cells (right). Values represent mean and SEM. ns, not significant by two-sided Student’s T-test without adjustment for multiple comparisons. d, Schematic of the workflow and (e) Sanger sequencing for two independent KCNQ1G/G clones and three KCNQ1A/A clones. f, Representative figures of flow cytometry analysis for NKX6-1 and INS comparing KCNQ1G/G and KCNQ1A/A clones (left). of the percentage of NKX6-1+/INS+ cells in beta cell stage cultures from KCNQ1G/G (n = 6; 2 clones × 3 differentiations) and KCNQ1A/A (n = 9; 3 clones × 3 differentiations) cells (right). ns, not significant by two-sided Student’s T-test without adjustment for multiple comparisons. Values represent mean and SEM.
Supplementary information
Supplementary Information
Supplementary Methods and Figs. 1–10
Supplementary Tables
Supplementary Tables 1–12 and Data 1–6
Source data
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Chiou, J., Zeng, C., Cheng, Z. et al. Single-cell chromatin accessibility identifies pancreatic islet cell type– and state-specific regulatory programs of diabetes risk. Nat Genet 53, 455–466 (2021). https://doi.org/10.1038/s41588-021-00823-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00823-0
This article is cited by
-
Cell-fate conversion of intestinal cells in adult Drosophila midgut by depleting a single transcription factor
Nature Communications (2024)
-
Chromosome 20p11.2 deletions cause congenital hyperinsulinism via the loss of FOXA2 or its regulatory elements
European Journal of Human Genetics (2024)
-
ExplaiNN: interpretable and transparent neural networks for genomics
Genome Biology (2023)
-
Chromatin accessibility differences between alpha, beta, and delta cells identifies common and cell type-specific enhancers
BMC Genomics (2023)
-
Effect of tissue-grouped regulatory variants associated to type 2 diabetes in related secondary outcomes
Scientific Reports (2023)