Abstract
Somatic mutations in driver genes may ultimately lead to the development of cancer. Understanding how somatic mutations accumulate in cancer genomes and the underlying factors that generate somatic mutations is therefore crucial for developing novel therapeutic strategies. To understand the interplay between spatial genome organization and specific mutational processes, we studied 3,000 tumor–normal-pair whole-genome datasets from 42 different human cancer types. Our analyses reveal that the change in somatic mutational load in cancer genomes is co-localized with topologically-associating-domain boundaries. Domain boundaries constitute a better proxy to track mutational load change than replication timing measurements. We show that different mutational processes lead to distinct somatic mutation distributions where certain processes generate mutations in active domains, and others generate mutations in inactive domains. Overall, the interplay between three-dimensional genome organization and active mutational processes has a substantial influence on the large-scale mutation-rate variations observed in human cancers.
Similar content being viewed by others
Main
The distribution of somatic mutations exhibits substantial variation among cancer genomes and is associated with transcription-factor-binding sites1,2, chromatin modifications3 and proximity to the nuclear periphery4. Chromatin conformation (Hi-C) studies have revealed the compartmental segregation of active and inactive chromatin regions. At a higher resolution, chromosomes are organized into topologically associating domains (TADs)5,6, which are largely cell-type invariant. Genes within the same TAD can exhibit coordinated expression and histone-modification patterns7,8,9,10. Global patterns of three-dimensional (3D) genome structure are closely related to DNA replication and transcription11. Notably, recent work demonstrated that the 3D chromatin organization creates a frame for replication timing as the changes in chromatin compartments preceded the changes in replication timing12. However, the connection between chromatin folding organization and somatic mutation rates in cancer genomes remains unclear. Diverse mutational processes lead to somatic mutations in human cancer13, and mutational signatures are used to delineate the imprints of underlying processes within cancer genomes14,15. Understanding the interplay between chromatin folding and mutational signatures is important to elucidate the mechanisms responsible for crucial DNA damage and repair processes. Here, we sought to understand the relationship between genome organization and mutational processes observed in 60.6 million somatic mutations from whole-genome sequencing datasets of 42 different histology subtypes (Supplementary Table 1).
The distribution of somatic mutations is correlated with the 3D genome organization
To evaluate the patterns of somatic mutation accumulation in relation to chromatin folding domains in this pan-cancer cohort, we first used common TAD boundaries among five different human cell types16. These boundaries represent previously identified TAD boundary profiles and exhibit 63% overlap with cancer cell line TAD boundaries (Extended Data Fig. 1a,b). In addition, we used subcompartment (domain) calls from an earlier study9, where each domain is enriched in several chromatin states and exhibits distinct transcriptional profiles. We annotated these domains based on chromatin states observed in 125 different human cell types (Extended Data Fig. 1c,d). We observed that chromatin state distribution in the majority of different cell types is consistent with our domain annotation types, suggesting there is not a strong confounding effect of using our domain annotations for specific cancer types (Extended Data Figs. 1e and 2a,b). Domains associated with active marks exhibited higher signal enrichment with the assay for transposase-accessible chromatin using sequencing (ATAC-seq; Extended Data Fig. 2c), whereas inactive domains contained higher DNA methylation levels (Extended Data Fig. 2d) in various cancer types. Overall, we observed that these domain annotations were one of the strongest single predictors of somatic mutational load that we could identify (Supplementary Table 2).
As the mutational load is correlated with the domain types (Extended Data Fig. 3a), to identify the genomic regions with marked mutational load differences, we focused on two specific domain types: transcriptionally inactive (inactive) and transcriptionally active (active) domain types that cover most (71%) of the human genome (Supplementary Table 3). We found that average mutational load increases sharply around the boundaries between transcriptionally distinct domains when compared to the boundaries separating transcriptionally similar domains (Fig. 1a and Extended Data Fig. 3a), which implies that the distribution of somatic mutations in cancer genomes corresponds to spatial chromatin organization. Late-replicating genomic regions acquire a higher mutational load17,18 (Extended Data Fig. 3b). The replication timing changes from early to late in a gradual manner11, whereas the transition in somatic mutation frequency is sharper at TAD boundaries (Fig. 1a). Notably, the change in unsmoothed replication timing between domains is lower (6.9%) than the mutational load change (20.6%) around 100 kb of the TAD boundaries. Therefore, TAD boundaries between active and inactive domains can be used as a better proxy to delineate the mutational load variation in cancer genomes (Supplementary Table 4). Of importance, mutational load changes around the TAD boundaries can be observed in various cancer types (Fig. 1b, Supplementary Fig. 1 and Extended Data Fig. 4a) suggesting that the observed distribution is not cancer-type specific. For example, melanoma and esophageal adenocarcinoma samples exhibited a similar distribution pattern, including the individual samples within the cohorts (Fig. 1b,c). To evaluate the robustness of our domain annotations, we ranked the domains in each cancer type based on their mutational load. We then compared the domain annotations in the highly mutated domains (first quantile) and the least mutated domains (fourth quantile) for each cancer type. For every cancer type in this study, the highly mutated domains were enriched in the inactive domains, whereas the least mutated domains were enriched in the active domain annotations (Extended Data Fig. 4b and Supplementary Fig. 2a), suggesting the utility of our domain annotations in this genome-wide mutational load study among various cancer types. Copy-number levels were comparable between active and inactive domains, indicating it is not a prominent confounding factor (Supplementary Fig. 2b). In addition, we observed a similar somatic mutation accumulation pattern in healthy human tissue, such as subcutaneous adipose and kidney (Supplementary Fig. 2c) tissue, suggesting that this distribution is not unique to cancer genomes. Overall, these results highlight an implicit link between regional mutation rates in human genomes and the spatial chromatin organization.
a, Average profiles of somatic mutation accumulation in 3,000 cancer samples and replication timing across 500 kb of boundaries delineating active to active, active to inactive, inactive to inactive and inactive to active domains. b, Plots show the accumulation patterns of somatic mutations in pan-cancer, melanoma and esophageal adenocarcinoma cohorts (heterochromatin: 180; inactive: 1,219; repressed: 969; active: 1,086; active-2: 593). Error bar limits are the 25th and 75th percentiles; the center line is the median. P values were derived from a Wilcoxon rank-sum test. c, Average profiles of somatic mutation accumulations in melanoma and esophageal adenocarcinoma cohorts across 500 kb of boundaries delineating inactive and active domains. Heat maps show mutation distributions in individual samples around the same genomic regions. Row maximum is represented with darker color. d, Domains are divided into transcribed and non-transcribed regions (schematic). Plots show the mutational loads in transcribed (Trx) and non-transcribed (non-Trx) regions of active, active-2 and inactive domains for the esophageal adenocarcinoma and melanoma cohorts (inactive: 1,219; active: 1,086; active-2: 593). Error bar limits are the 25th and 75th percentiles; the center line is the median. P values were derived from a Wilcoxon rank-sum test; NS, not significant. e, Schematic demonstrates the identification of genomic loci with significant mutational load change. f, Hi-C insulation score profiles for TAD boundaries, significant mutational load change sites and randomly selected genomic regions. Blue circles denote genomic loci with significant mutational load change. Black squares demonstrate TAD boundaries. g, Overlap of TAD boundaries. Significant mutational load change sites are represented with a red arrow. The empirical P value was derived from 1,000 randomly generated (gray curve) overlaps.
We then sought to understand whether the observed mutational difference between domains was due to differential transcriptional activity. We divided each domain as transcribed (including exons and introns) and non-transcribed regions (Fig. 1d). Notably, there were no significant differences in mutational load between transcribed and non-transcribed regions of the same domain type. However, it is important to note that non-transcribed regions of active domains acquired a significantly lower mutation burden when compared to transcribed regions of inactive domains (Fig. 1d and Supplementary Fig. 2d). These results suggest that the observed mutational load difference is a domain-wide effect and is not restricted to the transcriptional activity of the genome.
Lastly, if the mutational load variation is related to the 3D chromatin organization, the loci where a significant shift in mutation level was observed should overlap with the domain boundaries. To test this hypothesis, we called the loci with a significant shift in mutational distribution by identifying local variation in the second derivative of the smoothed mutational load signal (Fig. 1e). Indeed, these loci exhibited a similar interaction insulation in Hi-C matrices observed at the TAD boundaries (Fig. 1f), suggesting marked change in mutational load coincides with TAD boundaries. To assess the significance of this overlap, we randomly shuffled the same number of genomic regions and observed that genomic loci with a mutational load change were significantly (P < 0.001) overlapping with TAD boundaries (Fig. 1g). Overall, our results imply that the distribution of somatic mutations is closely related to 3D genome organization.
Distinct chromatin folding patterns and the distribution of somatic mutations
We next investigated the impact of differential chromatin folding on somatic mutation distributions. We focused on X-chromosome mutations, as the inactive (Xi) and active (Xa) X chromosomes exhibit distinct, well-studied folding structures that differ in males and females. The active X chromosome, similarly to autosomes, is organized into discrete chromatin folding domains. Strikingly, the inactive X chromosome is devoid of TADs; instead, Xi is structured into larger repressive domains19. We used allele-specific X-chromosome chromatin contact maps20 to understand the somatic mutation distributions between female and male participants with cancer on the X chromosome. Here, we did not include samples from the strongly sex-biased cancer types (breast, prostate, ovarian and uterine adenocarcinomas). Notably, female X-chromosome mutation distribution was overall more uniform when compared to the mutations in male X chromosomes, reminiscent of the TAD-less folding structure of the Xi chromosome (Fig. 2a,b and Extended Data Fig. 5a). In contrast, male X-chromosome somatic mutation distribution corresponded to domains observed within Xa, similar to mutation distribution patterns in autosomes (Fig. 2a,b). To calculate the mutational load variation along the X chromosome in both males and females, we compared the mutational load difference between a 500-kb active region (R1) and a neighboring 500-kb inactive region (R2) on the active X chromosome (Extended Data Fig. 5a). The mutational load change was 74.3% in male samples between these two regions, whereas the change was only 30.7% in female samples (Fig. 2c). This observation suggests that mutation distribution in female samples is less variable across the X chromosome, reflecting the absence of TAD-based folding structure of the inactive X chromosome. In particular, we observed in female participants that subclonal mutations on the X chromosome exhibited a more invariant distribution between R1 and R2 when compared to clonal mutations, a phenomenon implying that Xi acquires more mutations during tumor evolution than Xa (Fig. 2c and Extended Data Fig. 5b,c). We did not observe any difference between the distribution of clonal and subclonal mutations in autosomes (that is, chromosome 7; Extended Data Fig. 5d,e), indicating this difference is specific to the X chromosome. It is also important to note that hypermutation of Xi has been reported in certain cancer types21. We next sought to identify which cancer types exhibit higher mutation rates in the X chromosome in females. The biggest difference between male and female X-chromosome mutational load was observed in chronic lymphocytic leukemia samples. Female X-chromosome mutational load was significantly higher even when compared to copy-number-corrected male samples (P value < 10−6; Fig. 2d). We checked whether males with cancer/somatic X-chromosome polyploidy (non-Klinefelter syndrome) exhibited a similar mutational load to females, but found no significant gain in mutational load in males with somatic X-chromosome polyploidy, implying the observed increase is predominantly due to an inactive X chromosome (Fig. 2e). Brain cancers such as lower-grade glioma, glioblastoma (Extended Data Fig. 5f,g), pediatric brain tumors (Extended Data Fig. 5h), diffuse large B-cell lymphoma or head/neck squamous adenocarcinoma (Extended Data Fig. 5i,j) also presented significantly higher X-chromosome mutational load in females when compared to males. Consistent with previous findings21, X-chromosome copy number and the expression of Xi-specific noncoding RNA (XIST) was significantly correlated with the X-chromosome mutational load predominantly in breast, ovarian and uterine cancer samples (Extended Data Fig. 6a–d), suggesting mutational load increase is due to the inactive X chromosome.
a, Allele-specific Hi-C maps of inactive and active X chromosomes (between 115 and 155 Mb) from human retinal cells. Histograms show the distributions of somatic mutation accumulation for female and male samples. b, Clonal and subclonal mutation profiles are shown along the X chromosome (the represented part in a) in female and male participants. c, Bar plots show the degree of change in mutational load between R1 and R2 on the active X chromosome for the total, clonal and subclonal mutations. d, Distribution of X-chromosome mutational load compared to total mutation burden for males (n = 64; observed and copy-number-corrected values) and females (n = 31) in chronic lymphocytic leukemia (CLL) samples. In these and all other box plots, the center line is the median, box limits are the upper and lower quantiles and whiskers represent 1.5 × the interquartile range. P values were derived from Wilcoxon rank-sum test. e, Distribution of X-chromosome mutational load compared to total mutation burden for males and females in chronic lymphocytic leukemia and somatically acquired X-chromosome polyploidy males (non-Klinefelter syndrome). f, Mutational load difference around the SEMA6D gene between melanoma and esophageal adenocarcinoma cohorts. Normalized mutation counts are representative of melanoma and esophageal adenocarcinoma samples. Difference between the mutational load and the derivative of the difference is shown below. Highlighted regions mark significant mutational load change sites. g, Chromatin contact maps represent differential genome organization around the SEMA6D gene in melanoma (A2058) and esophageal adenocarcinoma (OE33) cell lines. Histograms represent the overall mutational load in the melanoma and esophageal adenocarcinoma cohorts. h,i, Box plots represent mutational load around the SEMA6D gene (melanoma: 219; esophageal adenocarcinoma: 127) (h) and The Cancer Genome Atlas (TCGA) normalized expression data for SEMA6D (melanoma: 448; esophageal adenocarcinoma: 182) (i).
As mutational load is correlated with the higher-order chromatin organization and differential chromatin folding leads to distinct mutational load patterns as exemplified by the inactive X chromosome, we next tested whether significant variations in mutational load between different cancer types could predict the local chromatin conformation differences between the same cancer types. To this end, we used the first derivative of the mutational load difference between melanoma and esophageal adenocarcinoma samples with a moving nonoverlapping window approach to identify changes in the mutational pattern. This analysis yielded 375 genomic loci with a significant shift in mutational load between melanoma and esophageal adenocarcinoma samples (Extended Data Fig. 6e). To test whether the predicted loci also exhibited a difference in chromatin folding, we generated 25-kb resolution Hi-C data from a melanoma (A2058) and esophageal adenocarcinoma (OE33) cell line. We observed a significant Hi-C signal change (log2 fold change > 2) around these loci compared to randomly selected genomic regions (Extended Data Fig. 6f). For example, the SEMA6D locus exhibited a distinct mutational load difference between melanoma and esophageal adenocarcinoma samples (Fig. 2f). We also observed a compartment shift in the same locus (Fig. 2g). We confirmed that this mutational load difference could be observed in individual samples of the melanoma and esophageal adenocarcinoma cohorts (Fig. 2h). The expression level of SEMA6D was also significantly higher in melanoma samples compared to the esophageal adenocarcinoma samples (Fig. 2i). Similarly, we examined genomic loci with differential mutational distributions between melanoma and esophageal adenocarcinoma samples around the SORSC1, CDH11, CPQ and PLOD2 genes and found Hi-C conformation differences in melanoma and esophageal adenocarcinoma Hi-C data (Extended Data Figs. 6g and 7a–c). Notably, in the case of CDH11 (Extended Data Fig. 7a), we observed a chromatin folding change but not the expected gene expression change between melanoma and esophageal adenocarcinoma samples suggesting that the observed mutational load difference is not only due to nucleotide excision repair (NER) with transcriptional apparatus (TC-NER)22 activity. Overall, mutational distribution patterns can be exploited to understand the local chromatin conformation structures, particularly for the cancer samples for which no Hi-C data are currently available. These results demonstrate the importance of chromatin folding for the accumulation of somatic mutations in cancer cells and highlight the plausibility of chromatin configuration significantly contributing to the regional somatic mutation variability in cancer genomes.
Patterns of mutational distributions by individual mutational signatures
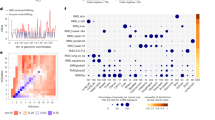
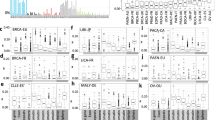
Distinct mutational processes are active and contribute to overall somatic mutation distribution in a given tumor23. Therefore, we studied how individual processes generate mutational distributions across chromatin folding domains by comparing the mutational load in active and inactive domains (Supplementary Table 5). Our analysis revealed that mutational signatures can be grouped into two distinct categories based on their accumulation patterns along chromatin folding domains (Fig. 3a). Signatures related to the known exogenous DNA damage agents such as ultraviolet (UV) light (signature 7), tobacco smoke (signature 4) or DNA-lesion-related signatures like signature 17 generated more mutations in inactive domains (Fig. 3a,b). On the other hand, mutational signatures associated with DNA damage repair deficiencies such as signature 3 associated with the failure of DNA double-strand break repair by homologous recombination (HR), exhibited a higher preference toward active domains (Fig. 3a,c). Two signatures associated with base excision repair (BER) deficiency (signatures 30 and 36) also exhibited a positive correlation with the active domains (Extended Data Fig. 8a). We observed higher activity of NER enzymes24 in active domains (Extended Data Fig. 8b); therefore, the observed pattern positively correlates with the active domains in DNA repair deficiency cases and reflects the combination of a malfunctioning repair mechanism and a damage that tends to accumulate in these regions more often. Overall, we find that DNA repair deficiencies, such as HR-, mismatch repair (MMR)- or BER-deficiency mutations, are enriched in active domains. Polymerase-proofreading deficiencies are an exception to this dichotomy. A component of the DNA polymerase ε (PoLε) deficiency (signature 10a) generated more mutations in inactive domains, which is consistent with an earlier study suggesting that Polε-related errors are biased toward late-replicating regions25. However, other PoLε-deficiency-associated components (signatures 10b and 28) generated more mutations in active domains (Extended Data Fig. 8c), which might suggest a potential link with DNA MMR deficiency26. Signatures 10b and 28 are persistent in daughter clones and single cells of Polε-deficient cancer cell lines showing continuing activity of these components in addition to the main signature 10a activity26. Notably, signature 28 mutations were predominantly T>G, similar to signature 17 mutations, but unlike the signature 17 distribution, signature 28 mutations generally occurred in the active domains (Extended Data Fig. 8d). This observation indicates that different mutational signatures with overlapping sequence context could generate unique mutational distributions. In addition, a positive correlation between mutational signature enrichment and active domain preferences was observed for mutational signatures of unknown etiology, such as signature 16. Signatures 12 and 16 were mostly observed in liver cancers with a strong transcriptional strand bias25. However, only signature 16 mutations exhibited a higher preference for active domains, with signature 12 generating more mutations in inactive domains, suggesting different underlying etiologies for these two signatures (Extended Data Fig. 8e,f). Lastly, we also noticed that significant distribution patterns in some signatures with unknown etiology such as signatures 39 and 40 generated a higher mutational load in active domains (Extended Data Fig. 8g,h). Therefore, studying the distribution of mutations induced by individual signatures within and among cancer types may help to distinguish the molecular underpinnings of different mutational processes in context.
a, Scatterplot shows the bias in mutational distribution between active and inactive domains by individual signatures. Pearson correlation was calculated for individual signatures. b,c, Dot plots demonstrate the association between the contribution of a mutation signature and mutation distribution ratio between active and inactive domains. b, For signatures exhibiting bias toward inactive domains, such as tobacco-smoke-related (SBS4) or somatic-hypermutation-related (SBS9) signatures. c, For signatures exhibiting bias toward active domains, such as DNA MMR-deficiency-related signature (SBS6) or HR-deficiency-related signature (SBS3).
Higher-order chromatin organization and DNA mismatch repair activity
To further investigate the relationship between DNA-repair-deficiency-driven mutations and chromatin folding, we analyzed MMR-deficient (MSI) samples in our cohort. We observed a flat mutation distribution pattern in various cancer types (Extended Data Fig. 9a), consistent with earlier work27. This flat inter-domain distribution was not driven by higher mutational load as we observed a low mutation burden in a colon MSI sample with a similar flat mutation distribution (Extended Data Fig. 9b). One possible explanation for this observed mutational distribution is that MSI samples have an altered 3D genome structure when compared to non-MSI genomes. To evaluate this possibility, we generated 25-kb-resolution Hi-C datasets on two DNA repair proficient (SW480 and CaCo2) and two DNA MMR-deficient (LoVo and DLD1) colon cancer cell lines. DNA repair status of these cell lines was reported by an earlier study28. As a reference for non-malignant colon tissue, we reprocessed Hi-C data from a healthy colon tissue sample29. We calculated the chromatin interactions around all TAD boundaries in MSS and MSI cancer cell lines to estimate changes in chromatin organization around the TAD boundaries (Extended Data Fig. 9c). Profiling the chromatin interactions across all TAD boundaries revealed that TAD boundary demarcation of MSI cancer cells was comparable to that of healthy colon and MSS cancer cells (Fig. 4a and Extended Data Fig. 9d). For example, chromatin interactions around a boundary between an active and an inactive domain were similar in MSS and MSI cancer cell lines despite the flat mutational distribution in MSI samples (Fig. 4b).
a, Average contact enrichment profiles across all TAD boundaries in healthy colon tissue and colon cancer cell line Hi-C data. b, Chromatin interaction maps of healthy colon tissue and colon cancer cell lines: SW480 and CaCo2: DNA repair proficient; LoVo: MSI; DLD1: MSI and PoLε deficient cell lines (histograms in d show mutation distribution in MSS and MSI colon samples around a boundary between inactive and active domains). c, Schematic of the approach used to compute the relative MMR efficiency, calculated by comparing normalized gastric MSI mutational load (aqua) with normalized gastric MSS mutational load (gray). Mutational loads were normalized based on the sum of mutations in MSS gastric samples. d, An example region represents the relative MMR efficiency and mutation distribution differences between MSS (gray) and MSI (aqua) gastric cancer samples with respect to chromatin folding. Histograms represent normalized mutation counts for MSS and MSI samples. TAD boundaries are depicted as black rectangles. Tiles show chromatin domain types (inactive: gray; active: orange). e, Aggregate plot shows the relative MMR efficiency around TAD boundaries delineating inactive domains from active domains. f, Dot plots represent the distribution of mutations in gastric MSS and MSI samples and the relative MMR efficiency in different domain types (heterochromatin: 180; inactive: 1,219; repressed: 969; active: 1,086; active-2: 593). P values were derived from a Wilcoxon rank-sum test.
Since we ruled out that MSI in and of itself was a primary effector of chromatin folding, we sought to further elucidate the underpinnings of the flat mutation distribution in MSI samples, given that the overall chromatin architecture remains unaffected. We focused on the gastric cancer samples because that cohort harbored a balanced mutational load between MSS and MSI samples compared to the colorectal samples. We estimated the efficiency of MMR proteins by computing the difference between normalized mutational load in gastric MSI and MSS samples (Fig. 4c). We observed that the difference between mutational load coincided with the TAD boundaries; the active domains acquired more mutations in MSI samples, whereas inactive domains acquired more mutations in MSS samples (Fig. 4d). The change in mutational load can be observed at the TAD boundaries delineating active domains from inactive domains (Fig. 4e), with presumptive MMR efficiency being significantly higher in active domains (Fig. 4f). This observation confirms the findings of earlier studies in yeast and humans, which suggested the MMR activity is lower in late-replicating, non-transcribed or exonic regions of the genome27,30,31. Our observation suggests that the flat inter-domain mutation distribution in MSI samples can be attributed to lack of MMR in active domains, and further, that differential MMR activity in active chromatin is a domain-wide effect, suggesting that chromatin organization plays a role in the efficiency of DNA repair enzymes.
APOBEC mutagenesis affects the inactive-X-chromosome mutation levels
In addition to the active domain bias of mutations attributed to the APOBEC family of enzymes (Fig. 3a and Extended Data Fig. 10a), these enzymes have also been implicated in the genesis of localized hypermutations (kataegis loci), based on their biochemical specificity to single-strand DNA (ssDNA; Extended Data Fig. 10b). We therefore profiled the distributions of the kataegis-like loci (C- or G-coordinated mutation clusters) with respect to 3D genome architecture. Kataegis-like loci significantly overlapped with TAD boundaries and domains with higher transcriptional activity (P values < 10−5); however, these loci were significantly depleted at inactive domains (P value < 10−5; Fig. 5a,b). Clustered APOBEC mutagenesis occurs in ssDNA during replication or in association with repair of double-strand breaks32,33. Our observation suggests that chromatin folding features could affect the distribution of persistent ssDNA prone to hypermutation. We then extended our analysis to investigate whether any mutational signature activity is correlated with higher mutational load around the TAD boundaries. Similar to the result between domain types (Fig. 3a), certain mutational signatures exhibit a higher load of mutations around TAD boundaries such as MSI signatures and Polε-deficiency (SBS10b.28) and BER-deficiency (SBS 30.36) associated signatures23 (Extended Data Fig. 10c).
a, Distribution of kataegis loci in the genome. Black arrowheads represent the overlap of kataegis loci with different domain types and TAD boundaries. Curves indicate the distribution of the expected overlap based on randomized data. b, APOBEC-related clustered mutations (kataegis) favor transcriptionally active domains. Kataegis-like clusters were identified as described previously. Example kataegis loci are shown with log10 intermutation distance on the y axis in two different breast cancer samples. Each dot represents a mutation and colors indicate different base substitutions. Colored rectangles depict individual domains within the given genomic locus (gray and blue: transcriptionally inactive domains; orange and red: transcriptionally active domains). c, Negative correlation between APOBEC-related mutation signatures (SBS2.13; log2 > 1.5) and X-chromosome mutation accumulation in all female participants (n = 584). P value was derived from Student’s t-test.
As APOBEC-related mutations are enriched in active domains, we investigated whether the rate of X-chromosome mutational load changes for the samples with higher APOBEC activity. Indeed, a negative correlation between X-chromosome mutation ratio and APOBEC-related signatures enrichments was observed in all female tumor samples across different cancer types (r2: −0.15; Fig. 5c). A subset of lung adenocarcinoma from females (12 samples) showed hypermutation on the X chromosome, but another subset (8 samples) contained similar levels of X-chromosome mutations as a subset from males (17 samples; Extended Data Fig. 10d). Indeed, signatures 2 and 13 (APOBEC related) occurred more frequently in females with a lower X-chromosome mutations ratio. Signatures 2 and 13 enrichment was negatively correlated with X-chromosome hypermutation levels in female lung adenocarcinoma samples (r2: −0.33; Extended Data Fig. 10e). Therefore, the degree of mutation accumulation in the inactive X chromosome is dependent on the active mutational processes and when a mutational process predominantly generates more mutations in active domains, the ratio of X-chromosome mutational load to overall mutation burden decreases. Taken together, our results demonstrate that, even though the underlying genomic sequence is identical, the activity of different mutational processes and chromatin configuration significantly contribute to the regional somatic mutation variability in cancer genomes.
Mutation distribution depends on the active mutational processes and can shift during tumor evolution
Mutational processes can change during tumor evolution; recent work profiled the tumor evolution in 2,658 cancer whole genomes and reported that in 40% of the samples the dominant mutational process changed from clonal to subclonal mutations34. Interestingly, the mutational processes that decrease activity during tumor evolution (clonal > subclonal) are generally affecting inactive domains (Fig. 6a), as demonstrated by the finding that mutational signature 7 was associated with UV light exposure or signature 4 was strongly correlated with smoking. We also noticed some signatures with unknown etiologies, for example, signatures 9, 12 and 17, were higher in clonal mutations than subclonal mutations. Conversely, mutational signatures generating more mutations in active domains tend to be more contributory to subclonal than clonal mutations such as MSI-related signatures or APOBEC-related signatures. Therefore, we hypothesize that there could be a link between mutational signatures biased toward the inactive domains and cell external carcinogens.
a, Bar plots show the direction of temporal changes (percentage of total samples with at least a twofold change) in mutational signatures between clonal or subclonal timing. b, Scatterplot shows the relationship between shift in APOBEC-related mutational signatures (SBS2.13) and the change in mutational distribution (ratio of active versus inactive domain loads) between clonal and subclonal mutations. c, An example of clonal to subclonal mutational spectrum change in a lung adenocarcinoma sample (SA286004). Clonal mutations were enriched in tobacco smoking signature (SBS4), whereas subclonal mutations were enriched in APOBEC-related signatures (SBS2.13). The probability bars for the six types of substitutions are displayed in different colors. The attribution of mutations obtained from the deConstructSigs algorithm (spectrum × 100) to the most characteristic signatures are shown.
Next, we investigated how the mutational distribution changed within a sample when a marked change in the activity of mutational signatures was observed during tumor evolution. For example, tobacco smoke has been postulated to indirectly activate APOBEC-related mutagenesis in certain cancer types35. Overall, the samples with higher APOBEC signatures in the subclonal mutation gained a significant level of active domain mutations (Fig. 6b, P value < 10−12), whereas samples with higher signature 4 clonal activity had higher inactive domain mutations (Supplementary Fig. 3a). For example, in a lung adenocarcinoma sample (SA286004), clonal mutations were enriched in signature 4, but subclonal mutations reflect APOBEC signatures (SBS2 and SBS13), indicating a shift of active mutational processes during somatic evolution of cancers (Fig. 6c). Indeed, there was a significant shift (P value < 10−10) in the mutational distribution where clonal mutations were significantly enriched in inactive domains but subclonal mutations exhibited a higher mutation burden in active domains (Supplementary Fig. 3b). We extended this analysis to other significantly changing mutational signatures such as UV-related signature 7. In the samples that showed a decline of the signature 7 predominance from clonal to subclonal mutations, there was a strong mutational shift from inactive domain mutations to active domain mutations (Supplementary Fig. 3c). We also observed a similar shift in mutational distribution change for signatures 9 (Supplementary Fig. 3d) and 17 (Supplementary Fig. 3e), where the decline of mutational activity from clonal to subclonal mutations coincided with the mutational burden shift from higher inactive domain load to active domains. These results highlight that mutational distributions in cancer genomes can shift during tumor evolution based on the active mutational processes and chromatin organization.
Discussion
Through integrating whole-genome somatic mutation with spatial genome organization data, we have found that the activity of various mutational processes causes distinct mutation distributions across local chromatin folding domains, as well as the genome as a whole. Our results highlight that somatic mutation accumulation is correlated with genome organization. Therefore, it may well be prudent that algorithms modeling the background mutation rate for discovering disease-relevant loci take topological chromatin organization into account. The distribution of different mutational processes could point to underlying etiologies of particular cancer types. Further investigations into the mutation distribution patterns along chromatin domains could help in identifying the biases generated by different mutational process over time, thereby augmenting our ability to elucidate the underlying etiologies for specific processes and giving deeper insights into avenues to explore prevention, early detection and therapeutic intervention. In addition, further clarity will require engineering specific repair deficiencies36 within the same DNA damage background to provide further mechanistic insights about the contribution of these components in somatic mutation accumulation patterns along chromatin domains.
The impact of chromatin folding is perhaps best illustrated in the female X-chromosome mutation distribution where there is marked interaction of mutation distribution within the distinct topology of inactive X chromosomes. The subclonal mutation rate on Xi is higher when compared to that of Xa in female tumors, with higher incidences in cancer types where mutational processes preferentially target the inactive chromatin domains. As such, these Xi observations illustrate the potential clinical insights that can be gained from exploring mutational processes and chromatin folding. Hypermutation is linked to immunotherapy response rate, likely due to higher neoantigen production37, and Xi activation is reported in human breast cancer models38. Reactivation of the Xi chromosome could potentially lead to higher neoantigen production and improved response rate to immunotherapy in female participants as the repressed gene regions in Xi potentially acquire twofold higher mutational load compared to the same regions in Xa. More broadly, thorough understanding of chromatin folding and pervasive mutation processes may present similar opportunities. Inhibiting repressive epigenetic enzymes such as EZH2 might open up immunotherapy response rate modulation in various cancer types as inactive domains with higher mutational load could lead to enhanced neoantigens expression upon EZH2 inhibition. Of note, 23% of coding mutations across the dataset of 3,000 genomes are in repressed chromatin domains, and thus may be potentially available for exploitation as neoantigens should efficient strategies for chromatin state modulation be developed.
Overall, chromatin conformation seems to contribute to the variable accumulation of somatic mutation within human cancer genomes. Whether germline mutation distribution and genetic variations among different population groups, somatic mutation accumulation throughout the aging process or mutations affecting somatic clonal proliferation, such as clonal hematopoiesis of indeterminate potential (CHIP), are correlated with the spatial genome organization remain interesting open questions. Utilizing tissue-specific chromatin folding datasets will reveal lineage-specific features of the mutational load variation among cancer types. A more thorough understanding of the 3D genome architecture in human cancers will advance our understanding of mutational processes and DNA repair activity—ultimately painting a more informative, nuanced picture of this disease.
Methods
Hi-C experiment and data analysis
In this study, we generated in situ high-resolution Hi-C data from three different colon cancer cell lines (CaCo2, LoVo and DLD1) and one melanoma cell line (A2058). Hi-C was performed using the in situ Hi-C protocol as described previously9, with 2–5 million cells per experiment digested with the MboI restriction enzyme. Hi-C libraries were sequenced on a HiSeq 4000.
In addition, we utilized Hi-C data from colon cancer (SW480) and esophageal adenocarcinoma (OE33) cell lines from our previous study (GSE116694). We also downloaded FASTQs of normal colon tissue Hi-C data from the ENCODE project data portal (Supplementary Table 1). For all these Hi-C datasets, we deployed the same processing pipeline. In brief, reads were aligned to the hg19 reference genome using BWA-MEM39, and PCR duplicates were removed with Picard. Hi-C interaction matrices were generated using in-house pipelines, and matrices were normalized using the iterative correction method40. HiCPlotter was used for visualizing normalized Hi-C interaction maps41.
To compare TAD boundary strengths in Fig. 4a, we used quantile normalization across the whole genome-wide interactions to normalize normal colon, SW480, CaCo2, LoVo and DLD1 Hi-C datasets together. For each common TAD boundary, we sliced the Hi-C interaction maps as a 32 × 32 matrix where the TAD boundary was at the center (Extended Data Fig. 9c), similar to an analysis method used previously42. Next, we calculated boundary strength as a ratio of two values in each matrix, with the first denoting within-TAD intensity (that is, the sum of intra-TAD interactions) and the second denoting between-TAD intensity (that is, the sum of interactions falling outside of TAD regions). Boundary strength was calculated as the log2 ratio of within-TAD intensity to between-TAD intensity. Individual boundary strength calculations for each Hi-C dataset are plotted in Extended Data Fig. 9d, and the aggregate plot across all TAD boundaries for each Hi-C dataset is shown in Fig. 4a.
Chromatin domain annotations
We obtained chromatin domain information (originally defined as subcompartments) for the GM12878 cell line from Rao et al.9. To cluster individual chromatin domains based on epigenetic modifications, we sought to utilize comprehensive epigenome profiling from various human cell types. To this end, we used an entropy-based approach (epilogos) to calculate the occurrence of each chromatin state enrichment for a given genomic region across all cell types profiled in the Roadmap Epigenome Consortium43 (Supplementary Table 1). We calculated the ratio of a domain genomic space covered by each chromatin state, divided by the length of the domains and generated a normalized matrix where columns are domains and rows are each 15 ChromHMM chromatin states generated by the Roadmap Epigenome Consortium. We applied hierarchical clustering to the rows to identify similar chromatin states and k-means clustering to the columns to pinpoint group domains containing similar epigenetic modifications. We performed k-means clustering with k = 2–8 clusters and selected k = 5 clusters as previous chromatin studies revealed five distinct epigenetically modified chromosomal domains and k = 5 corresponded to better visually discernible domains (Extended Data Fig. 1c). To demonstrate that these domain annotations were overall reflective of the domain annotations in various cell types, we performed a similar analysis using ChromHMM state data. We generated a matrix for each individual cell type, where we calculated the ratio of a domain genomic space covered by each state based on the 15-chromatin state model by the Roadmap Epigenome Consortium. We then sorted the domains based on our domain annotations obtained with all cell types and plotted the heat maps represented in Extended Data Figs. 1e and 2a,b. The ratios of enriched states for domain annotations from each cell type were shown as box plots beneath the heat maps.
To identify the average DNA methylation and open-chromatin levels for each domain annotation, we used TCGA 450 K array44 and ATAC-seq45 datasets (Supplementary Table 1). We calculated the average values for each domain across a cancer type. These average values are shown as box plots in Extended Data Fig. 2c,d. A similar analysis was performed for the copy-number level for each domain across the whole cohort.
Replication timing
We generated the unsmoothed weighted average Repli-seq data for the GM12878 cell line by using the following formula obtained from the University of California Santa Cruz (UCSC) genome browser: (0.917 × G1b) + (0.750 × S1) + (0.583 × S2) + (0.417 × S3) + (0.250 × S4) + (0 × G2) as plotted in Fig. 1a. We calculated the average replication timing in various cell types by downloading Repli-seq data from the UCSC genome browser ENCODE portal for the following cell types: BJ, GM06990, GM12801, GM12812, GM12813, GM12878, HeLa-S3, HepG2, HUVEC, IMR90, K562, MCF-7, NHEK and SK-N-SH. Replication timing values for smoothed wavelength transformed data were binned into 25-kb windows across the genome to discretize the data.
TAD boundary analysis
We used our common TAD boundary analyses from a recent study16. Briefly, Hi-C data for GM12878, HUVEC, IMR90, HMEC, NHEK and K562 cell lines were downloaded from the Gene Expression Omnibus (GEO; GSE63525)9. Raw intrachromosomal 25-kb-resolution values were normalized by diving with the multiplication of Knight and Ruiz normalization scores for two contacting loci. To identify the TAD boundaries in each cell line, we used an insulation-score approach with the source code obtained from https://github.com/dekkerlab/crane-nature-2015/. We used the parameters --is 1000000 --ids 200000 --im mean --bmoe 1 --nt 0.1 --v. Common TAD boundaries were identified where boundaries of all five cell types (GM12878, HUVEC, IMR90, HMEC and NHEK) occurred within two Hi-C bins or 50 kb in genomic range. These boundaries and flanking domain types16 are provided in Supplementary Table 4.
To compare the overlaps of these common boundaries with the cancer cell line TAD boundaries, we downloaded TAD boundary regions also identified via the insulation-score approach for the following cancer cell lines: ACHN, DLD1, SKMEL, A549, T47D, Caki2, H460, PRMI, PANC1, G401, SK-N-MC and SJCRH30 (Supplementary Table 1).
We calculated the insulation score by moving a window across the Hi-C matrix diagonally and calculating the sum of interaction for a given bin with up to 2-Mb flanking regions and calculated the log2 of the observed bin to the mean of interaction values within the given 2-Mb window.
Somatic mutation analysis
We used the somatic mutation calls from 3,000 tumor samples cited by the Pan-Cancer Analysis of Whole Genomes (PCAWG) mutational signatures working group in a recent publication23, which includes PCAWG samples with consensus mutation calling pipeline (Supplementary Table 1). To calculate the mutational load variation across chromatin domains, we binned the genome in 25-kb nonoverlapping windows and calculated the sum of mutations in each window. We performed this analysis by summing the mutational load in each cancer type or across the whole cohort. We plotted the enrichment patterns as line plots around TAD boundaries (Fig. 1a–c and Extended Data Fig. 3a) or calculated the total number of mutations in each domain and divided by the domain length for box plot representations (Fig. 1b and Supplementary Fig. 1).
To identify the mutational distribution in individual samples (Fig. 1c), we calculated the average mutational load for each sample across the TAD boundaries, delineating active and inactive for each sample. We then normalized this array of values to scale between 0 to 1 for each sample and plotted the result as a heat map.
To demonstrate the utility of our aggregate domain annotations from all of the Roadmap Epigenome cell types for mutational load analysis, we sorted the domains based on their mutational load in each cancer type. Next, we divided the domains into quarters based on this mutational load analysis. For domains with the least mutational load (fourth quarter) and domains with the highest mutational load (first quarter), we calculated the domain annotations. The values for domain annotations for the first and fourth quarters for each histology type are shown as bar plots (Extended Data Fig. 4b and Supplementary Fig. 2a). As an example of the appearance of mutational load distribution in the first and the fourth quarters of the melanoma cohort, we calculated the mutation accumulation in each sample for each domain. This array of values was then normalized to scale between 0 to 1 for each sample and plotted as a heat map (Extended Data Fig. 4b).
The active signatures extracted by the SigProfiler algorithm in each sample were obtained from the same publication (https://www.synapse.org/#!Synapse:syn12025148)23. We used deconstructSigs46 to delineate the contribution of mutational signatures with the default Cosmic 30 signature option. Somatic mutation data from normal human tissue were obtained from a recent publication47. MSI status of the samples was determined by the PCAWG mutational signatures working group and obtained from the synapse repository: https://www.synapse.org/#!Synapse:syn8016399. We generated P-value calculations with the Wilcoxon rank-sum test using the scipy.stats.ranksums function in SciPy, unless otherwise indicated.
Mutation timing analysis
The PCAWG tumor evolution and heterogeneity working group classified mutations as clonal (early, late or NA) or subclonal mutations34. This approach uses a hierarchical model to calculate timing parameters based on copy number and variant allele frequency data. If the mutation clonal frequency is equal to 1, those mutations are classified as clonal, whereas frequency lower than 1 is designated as subclonal. Code to determine timing of the mutations is available at https://www.github.com/gerstung-lab/MutationTime.R/.
We obtained the changes in mutational signatures during tumor evolution (clonal to subclonal) from the same publication’s Source Data Fig. 5. For the mutational shift analysis in this work, we used samples with at least twofold change (log2 ≥ 1) between clonal and subclonal mutations for a mutational signature. In order to analyze the changes in mutation distribution associated with mutational signatures during tumor evolution, we calculated the difference between clonal mutational load (active versus inactive mutation ratio) and subclonal mutational load (active versus inactive ratio) for each sample. We plotted these values with the changes in mutational signature activity in Fig. 6 and Supplementary Fig. 3.
Mutation distribution analysis
To compare the mutational load distribution generated by different mutational signatures, we calculated the ratio of mutation burden in transcriptionally inactive domains versus transcriptionally active domains. First, for each sample, we binned the mutations in 25-kb nonoverlapping windows along the genome. Next, we calculated the sum of mutations at inactive and active domains and normalized this value by the length of active and inactive domains. For evaluating the biases of the mutational load contribution for an individual signature toward active or inactive domains, we plotted this ratio of normalized mutational load between active and inactive domains and the activity of a specific mutational signature (obtained from https://www.synapse.org/#!Synapse:syn12025148). We did not apply any filter for the activity level of a specific mutational signature. The best-fitting curve for the points of each signature was calculated with the polynomial.fit function in the NumPy library. The obtained slopes and Pearson correlation statistics were then used to generate Fig. 3a.
To calculate the enrichment patterns of mutations generated by individual mutational signatures around the TAD boundaries, we first calculated the total number of mutations overlapping with common TAD boundaries in each sample. Next, we generated the expected number of mutations by shuffling the boundary regions on the same chromosome and calculating the total number of mutations at the random locations. We performed this shuffling exercise 10,000 times and calculated the median expected number for each sample. Then, we plotted the log2 fold change between observed and expected mutations around the TAD boundaries and the activity of a specific mutational signature. The best-fitting curve for the points of each signature was calculated with the polynomial.fit function in NumPy. The obtained slopes and Pearson correlation statistics were then used to generate Extended Data Fig. 10c.
To estimate the mutational load of the transcribed and non-transcribed regions inside a domain, we first merged gene-body (including introns) annotations from the RefSeq and UCSC knownGenes tables to obtain a set of transcribed regions in the genome. Then, for the active, active-2 and inactive domain annotations, we calculated the number of mutations occurring at the transcribed and non-transcribed regions. We used the mutation calls from melanoma, esophageal adenocarcinoma, hepatocellular carcinoma and lung carcinoma (adeno- and squamous- carcinoma histologies). These ratios are presented in Fig. 1d and Supplementary Fig. 2d.
To estimate the contribution of genomic correlates to the observed mutational load variation in cancer genomes, we binned the mutations in 25-kb nonoverlapping windows along the genome. We then identified the highest (first quantile) and the lowest (fourth quantile) mutational load bins. To calculate the domain annotations, we coded each domain type (heterochromatin: 1; inactive: 2; repressed: 3; active: 4; active-2: 5) and assigned each genomic bin a chromatin domain-type score. We used Roadmap Epigenome’s E059 (melanocyte) and E079 (esophagus tissue) heterochromatin H3K9me3 ChIP-seq and ChromHMM 15-chromatin states information for calculating their contribution to mutational distribution patterns in melanoma and esophageal adenocarcinoma genomes, respectively. We used linear models that were fitted by generalized least-squares estimation between each individual feature and mutation accumulation value between the first and fourth quantiles. Adjusted r2 values were calculated using the lm function in R (Supplementary Table 2). For chromatin states we reported the best individual chromatin state score.
Identifying significant changes in mutational load
To test whether the significant changes in mutational load coincide with the TAD boundaries, we sought to identify the genomic loci where there was a significant shift in mutational load patterns. We first calculated the mutational load as the sum of mutations from all the samples for this analysis. We smoothed this mutational load data with 25-kb, 50-kb, 75-kb and 100-kb genomic windows. Next, we calculated the second derivative of the smoothed mutational load data for each window size. Local maxima of the second-derivative signal were calculated with the find_peak function in SciPy. A consensus peak list was derived by identifying the overlapping peaks for 25-kb, 50-kb, 75-kb and 100-kb genomic windows. We calculated the overlap rate between this consensus list of significant mutational load change loci and the TAD boundaries if these events occurred within a 25-kb distance (Fig. 1e). To test whether the observed overlap was significant, we generated random genomic regions by keeping events on the same chromosome (using the --chrom option in shuffleBed48). We calculated the overlap between these random genomic sites and performed shuffling of the TAD boundaries 10,000 times. We computed the cumulative distribution of expected overlap, while z-scores and P values were calculated based on the observed number and obtained distribution from bootstrapping (Fig. 1f).
Estimating local chromatin conformation changes via mutational load differences
We used melanoma and esophageal adenocarcinoma samples to estimate the local chromatin conformation changes by mutational load differences. We scanned the genome with 10-Mb nonoverlapping windows. For each window, we normalized the mutational load in melanoma and esophageal adenocarcinoma genomes between 0 to 1 because of the number of mutation differences between these two cohorts. Next, we subtracted these normalized mutational loads and smoothed this difference signal with a 1-Mb genomic window. We calculated the first derivative of this smoothed mutational load difference. Local maxima and minima of the first-derivative signal were calculated with the find_peak function in SciPy using a high cutoff of 0.01. The identified loci were designed as differential chromatin folding regions between melanoma and esophageal adenocarcinoma genomes. To assess the validity of the potential genomic loci with differential chromatin conformation in melanoma and esophageal adenocarcinoma genomes, we generated 25-kb-resolution Hi-C data from melanoma (A2058) and esophageal adenocarcinoma (OE33)16 cell lines. We calculated the log2 fold change between normalized A2058 and OE33 Hi-C signals around the estimated chromatin folding difference loci. As a negative control, we randomly selected the same number of genomic loci and calculated the log2 fold change between normalized A2058 and OE33 Hi-C signals around these random genomic sites. We plotted this comparison in Extended Data Fig. 6f. The normalized gene expression data for the genes are depicted in Fig. 2i and Extended Data Figs. 6g and 7a–c. We downloaded the melanoma and esophageal adenocarcinoma cohort from TCGA and normalized RNA sequencing by expectation-maximization (RSEM) values from cBioPortal49.
APOBEC enrichment and kataegis-like cluster analysis
APOBEC enrichments were calculated using a published method49. Non-APOBEC-enriched groups represent samples with no statistically significant enrichment with the stringent tCa motif of APOBEC mutagenesis. APOBEC-enriched samples include both ‘A3A-like’ and ‘A3B-like’ groups of tumors. To remove a potentially confounding factor, we excluded melanoma samples in this analysis because UV-mediated mutation signature resembles APOBEC signatures.
Kataegis-like G- or C-strand-coordinated mutation clusters were identified across the PCAWG dataset (https://www.synapse.org/#!Synapse:syn7437313) as described in Chan et al.50. Briefly, groups of closely spaced mutations in either C or G of the top (sequenced) strand were identified, such that any pair of adjacent mutations within each group was separated by less than 10 kb and with P values calculated based on mutation density in a sample of 10−4 or less. Shuffled clustered mutations were generated by randomly assigning clusters while keeping the number of boundaries per chromosome constant. Shuffling was performed 10,000 times for the clustered mutation set. We computed the cumulative distribution of expected overlaps, while z-scores and P values were calculated based on observed number and obtained distribution from bootstrapping.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Code availability
Python scripts used in this manuscript are available at https://github.com/kcakdemir/MutationalDistribution/.
References
Perera et al. Differential DNA repair underlies mutation hotspots at active promoters in cancer genomes. Nature 532, 259–263 (2016).
Sabarinathan, R., Mularoni, L., Deu-Pons, J., Gonzalez-Perez, A. & López-Bigas, N. Nucleotide excision repair is impaired by binding of transcription factors to DNA. Nature 532, 264–267 (2016).
Makova, K. D. & Hardison, R. C. The effects of chromatin organization on variation in mutation rates in the genome. Nat. Rev. Genet. 16, 213–223 (2015).
Smith, K. S., Liu, L. L., Ganesan, S., Michor, F. & De, S. Nuclear topology modulates the mutational landscapes of cancer genomes. Nat. Struct. Mol. Biol. 24, 1000–1006 (2017).
Dekker, J. & Heard, E. Structural and functional diversity of topologically associating domains. FEBS Lett. 589, 2877–2884 (2015).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014).
Miura, H. et al. Single-cell DNA replication profiling identifies spatiotemporal developmental dynamics of chromosome organization. Nat. Genet. 51, 1356–1368 (2019).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Akdemir, K. C. et al. Disruption of chromatin folding domains by somatic genomic rearrangements in human cancer. Nat. Genet. 52, 294–305 (2020).
Stamatoyannopoulos, J. A. et al. Human mutation rate associated with DNA replication timing. Nat. Genet. 41, 393–395 (2009).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Jégu, T., Aeby, E. & Lee, J. T. The X chromosome in space. Nat. Rev. Genet. 18, 377–389 (2017).
Darrow, E. M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl Acad. Sci. USA 113, E4504–E4512 (2016).
Jäger, N. et al. Hypermutation of the inactive X chromosome is a frequent event in cancer. Cell 155, 567–581 (2013).
Kamarthapu, V. & Nudler, E. Rethinking transcription coupled DNA repair. Curr. Opin. Microbiol. 24, 15–20 (2015).
Alexandrov et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Hu, J., Adar, S., Selby, C. P., Lieb, J. D. & Sancar, A. Genome-wide analysis of human global and transcription-coupled excision repair of UV damage at single-nucleotide resolution. Genes Dev. 29, 948–960 (2015).
Haradhvala et al. Mutational strand asymmetries in cancer genomes reveal mechanisms of DNA damage and repair. Cell 164, 538–549 (2016).
Petljak et al. Characterizing mutational signatures in human cancer cell lines reveals episodic APOBEC mutagenesis. Cell 176, 1282–1294 (2019).
Supek, F. & Lehner, B. Differential DNA mismatch repair underlies mutation rate variation across the human genome. Nature 521, 81–84 (2015).
Mouradov, D. et al. Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. Cancer Res. 74, 3238–3247 (2014).
GTEx Consortium. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Lujan, S. A. et al. Heterogeneous polymerase fidelity and mismatch repair bias genome variation and composition. Genome Res. 24, 1751–1764 (2014).
Frigola, J. et al. Reduced mutation rate in exons due to differential mismatch repair. Nat. Genet. 49, 1684–1692 (2017).
Chan, K. & Gordenin, D. A. Clusters of multiple mutations: incidence and molecular mechanisms. Annu. Rev. Genet. 49, 243–267 (2015).
Sakofsky, C. J. et al. Repair of multiple simultaneous double-strand breaks causes bursts of genome-wide clustered hypermutation. PLoS Biol. 17, e3000464 (2019).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 https://doi.org/10.1038/s41586-019-1907-7 (2020).
Alexandrov, L. B. et al. Mutational signatures associated with tobacco smoking in human cancer. Science 354, 618–622 (2016).
Volkova, N. V. et al. Mutational signatures are jointly shaped by DNA damage and repair. Nat. Commun. 11, 2169 (2020).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. N. Engl. J. Med. 377, 2500–2501 (2017).
Chaligné, R. et al. The inactive X chromosome is epigenetically unstable and transcriptionally labile in breast cancer. Genome Res. 25, 488–503 (2015).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Akdemir, K. C. & Chin, L. HiCPlotter integrates genomic data with interaction matrices. Genome Biol. 16, 198 (2015).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Hutter, C. & Zenklusen, J. C. The cancer genome atlas: creating lasting value beyond its data. Cell 173, 283–285 (2018).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science 362, eaav1898 (2018).
Rosenthal, R., McGranahan, N., Herrero, J., Taylor, B. S. & Swanton, C. DeconstructSigs: delineating mutational processes in single tumors distinguishes DNA repair deficiencies and patterns of carcinoma evolution. Genome Biol. 17, 31 (2016).
Franco, I. et al. Whole-genome DNA sequencing provides an atlas of somatic mutagenesis in healthy human cells and identifies a tumor-prone cell type. Genome Biol. 20, 285 (2019).
Dale, R. K., Pedersen, B. S. & Quinlan, A. R. Pybedtools: a flexible Python library for manipulating genomic datasets and annotations. Bioinformatics 27, 3423–3424 (2011).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072 (2015).
Acknowledgements
We thank the participants and their families for contributing to this study. We thank Z. Coban-Akdemir, E. Keung, P. Polak, T. Gutschner, J. Posey and R. Jenq for their critical reading of this manuscript. We also thank our families, especially our children (special thanks to S. Akdemir from K.C.A.) for their support and understanding during the COVID-19 outbreak. This work was supported by a Cancer Prevention Research Institute of Texas award (R1205), the Welch Foundation’s Robert A. Welch Distinguished Chair Award (G-0040 to P.A.F.), the Emerson Collective Cancer Research Fund (to K.C.A.), SPORE in Brain Cancer CEP (P50CA127001 to K.C.A.), a US National Institutes of Health (NIH) Director’s Early Independence Award (DP5OD023071 to J.R.D.) and an NIH Intramural Research Program Project (Z1AES103266 to D.A.G.).
Author information
Authors and Affiliations
Contributions
K.C.A., J.R.D. and P.A.F. designed the study. K.C.A. and J.R.D. performed the computational analysis. V.T.L. and J.R.D. performed the Hi-C experiments on DLD1, CaCo2, LoVo and A2058 cell lines. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chromatin domains reflect a similar domain-wide epigenetic profile across different cell types.
a, Overlaps between common TAD boundaries in this manuscript and TAD boundaries from ENCODE cancer cell lines. Horizontal bars represent the total number of TAD boundaries per cell type. Vertical bars represent the number of intersecting boundaries between cell types. Combination matrix (below), circles indicate that denote cell types are part of the intersection for each vertical bar. b, Aggregate plots show average cell-type specific enrichment levels for Insulation score, CTCF binding sites, DNAseI hypersensitivity regions and H3K9me3 ChIP-seq enrichment levels from GM12878 cell line around common TAD boundaries. c, Classification of subcompartments (domains) based on chromatin state coverage. Heatmap shows domain-length normalized coverage of each chromatin state from Roadmap Epigenome aggregate data (rows) for each domain (columns). Chromatin states are clustered based on co-enrichments within domains. Domains are classified into five groups: heterochromatin (purple), inactive (gray), repressed (blue), active (orange), and active-2 (red) according to dominant chromatin state combinations. d, Box plots represent the enrichment of selected ChromHMM chromatin states in each domain type. These states are selected based on their highest enrichment in the heatmap plotted in Extended Data Fig. 1c. In these and all other boxplots, the center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. Heterochromatin:180, Inactive:1219:, Repressed:969, Active:1086, Active-2:593. e, Heatmaps represent domain-length normalized coverage of cell-type specific ChromHMM chromatin states within domain for representative Roadmap Epigenome cell types. Cell type’s name and the Roadmap Epigenome IDs are denoted on top of each heatmap. Domain-state matrices are not clustered but sorted based on our five domain annotations. These annotations are denoted as colored bars beneath the heatmap. Box plots below represent the domain-length normalized coverage of chromatin states for the corresponding cell type. Domain numbers are Heterochromatin:180, Inactive:1219:, Repressed:969, Active:1086, Active-2:593.
Extended Data Fig. 2 Enrichment of domain-wide epigenetic profile across different cell types.
a, b, Heatmaps represent domain-length normalized coverage of cell-type specific ChromHMM chromatin states within domain for representative Roadmap Epigenome cell types. Cell type’s name and the Roadmap Epigenome IDs are denoted on top of each heatmap. Domain-state matrices are not clustered but sorted based on our five domain annotations. These annotations are denoted as colored bars beneath the heatmap. Box plots below represent the domain-length normalized coverage of chromatin states for the corresponding cell type. In these and all other boxplots, the center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. In these and following plots domain numbers are Heterochromatin:180, Inactive:1219:, Repressed:969, Active:1086, Active-2:593. c, Dot plots show the normalized log2 ATAC-Seq signal in each domain type. Median ATAC-Seq signal was calculated across the samples of each histology-type. Name of the cancer type is denoted on top of the plot. Error bar limits are 25th and 75th percentiles; the center line is the median. d, Dot plots show the average beta value from Illumina 450 K DNA methylation array in each domain type. Average beta values were calculated across the samples of each histology-type. Name of the cancer type is denoted on top of the plot. Error bar limits are 25th and 75th percentiles; the center line is the median.
Extended Data Fig. 3 The distribution of somatic mutations in cancer genomes is correlated with the three-dimensional genome organization.
a, Mutational load is correlated with spatial chromatin organization. Average profiles of somatic mutation accumulation in 3000 cancer samples (green) across 500Kb of TAD boundaries. Flanking domain types are denoted on top of each plot. b, Dot plots show the average replication timing patterns across different domain types. Domain numbers are Heterochromatin:180, Inactive:1219:, Repressed:969, Active:1086, Active-2:593. P-value was derived from Wilcoxon rank-sum test.
Extended Data Fig. 4 The distribution of somatic mutations in different cancer histology types.
a, Average profiles of somatic mutation accumulation in different cancer histology types across 500Kb of TAD boundaries delineating inactive and active domains. Y-axis represents average mutational load in 25 kb windows across all samples of a given histology type. Name of the cancer type is denoted on top of the plot. b, Heatmap represents the mutational load for every melanoma sample in the fourth and first quartile domains. Domains are ranked based on their mutation load in the whole melanoma cohort (summing mutations from all samples). On top, colored bars denote the domain annotations. Bar plots on the left show the number of domain annotations in the fourth and first quantiles.
Extended Data Fig. 5 Unique folding of inactive X chromosome shapes the distribution of somatic mutations.
a, Hi-C map of active (Xa) X-chromosome showing an active domain (R1) between inactive domains from a male human umbilical vein cell line (HUVEC, cell line from a male donor). Below, histograms show the distributions of clonal and subclonal somatic mutation accumulation (in 25 kb non-overlapping windows) per female (red) and male (blue) samples. Mutation numbers were normalized based on the number of samples. b, c, Histograms show the distributions of b) clonal and c) subclonal somatic mutation accumulation (in 25 kb non-overlapping windows) per female (red) and male (blue) samples along the entire X chromosome. d, e, Autosomal mutation accumulation patterns are not distinct between males and females. Plots show mutational load along chromosome 7 for the clonal (d) and subclonal (e) mutations in female (red) and male (blue) patients. f–j, Distribution of X chromosome mutation load compare to total mutation burden for males (observed and copy-number corrected values) and females in f) lower grade glioma (n = 18), g) glioblastoma (n = 40), h) pediatric brain tumors (n = 253), i) diffuse large b-cell lymphoma (n = 107), j) head and neck squamous carcinoma cohorts (n = 57). The center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. P-values were derived from Wilcoxon rank-sum test.
Extended Data Fig. 6 Distinct chromatin folding patterns and the distribution of somatic mutations.
a–c, Correlation between X chromosome copy number and X chromosome mutation burden in a) breast adenocarcinoma (n = 197), b) uterine endometrial carcinoma (n = 51) and c) ovarian adenocarcinoma (n = 113). P-values were derived from Student’s t-test. d, Correlation between expression of XIST with X chromosome mutation burden in breast adenocarcinoma (n = 197). P-value was derived from Student’s t-test. e, Dot plots show the normalized mutation load difference between melanoma and esophageal-adenocarcinoma cohort around the genomic regions identified as significant mutational load loci. Same number of regions are randomly selected across the genome and normalized mutation load difference was calculated for the same samples. In these and all other boxplots, the center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. P-value was derived from Wilcoxon rank-sum test. f, Dot plots represent log2 fold-change between a melanoma (A2058) and an esophageal-adenocarcinoma (OE33) cell line Hi-C data around the genomic regions identified as significant mutational load loci. Same number of regions are randomly selected across the genome and log2 Hi-C fold-change numbers were calculated for the A2058 and OE33 cell lines. P-value was derived from Wilcoxon rank-sum test. g, Mutational load difference SORCS1 gene between melanoma and esophageal-adenocarcinoma cohorts. Normalized mutation counts represented for melanoma (green) and esophageal-adenocarcinoma (purple) samples. Difference between the mutation load and the derivative of the difference is shown below. Highlighted regions mark significant mutation load change sites. Below heatmaps represent chromatin folding organization around SORCS1 gene in a A2058 and OE33 cell lines. Histograms represent the overall mutation load in melanoma and esophageal-adenocarcinoma cohorts. Box plots represent mutation load (melanoma:219; esophageal-adenocarcinoma:127) and normalized expression levels (melanoma:448; esophageal-adenocarcinoma:182) of the SORCS1 in the melanoma and esophageal-adenocarcinoma cohorts. P-values were derived from Wilcoxon rank-sum test.
Extended Data Fig. 7 Comparing mutational load patterns between cancer types can yield local chromatin conformation differences.
a–c, Mutational load difference around a) CDH11, b CPQ, c) PLOD2 genes between melanoma and esophageal-adenocarcinoma cohorts. Normalized mutation counts represented for melanoma (green) and esophageal-adenocarcinoma (purple) samples. Difference between the mutation load and the derivative of the difference is shown below. Highlighted regions mark significant mutation load change sites. Below heatmaps represent chromatin folding organization around a) CDH11, b) CPQ, c) PLOD2 genes in a melanoma (A2058) and esophageal-adenocarcinoma (OE33) cell lines. Histograms represent the overall mutation load in our melanoma and esophageal-adenocarcinoma cohorts. Box plots represent mutation load in individual samples (melanoma:219; esophageal-adenocarcinoma:127) and normalized expression levels (melanoma:448; esophageal-adenocarcinoma:182) of the a) CDH11, b) CPQ, c) PLOD2 genes in the melanoma and esophageal-adenocarcinoma cohorts. The center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. P-values were derived from Wilcoxon rank-sum test.
Extended Data Fig. 8 Patterns of mutational distributions by individual mutational signatures.
Dot plots demonstrate the association between the contribution of mutation signatures and mutation distribution in active and inactive domains (x-axis). Signatures exhibiting a preference in the active domains, namely a, SBS30.36, c, SBS10b.28, f, SBS16, g, SBS39, h, SBS40, and were colored in orange color. On the other hand, signatures exhibiting a preference toward the inactive domains, namely c, SBS10a, d, SBS17, e, SBS12 were colored in gray color. b, Dot plots show the NER binding in different domain types. Domain numbers: Heterochromatin:180, Inactive:1219:, Repressed:969, Active:1086, Active-2:593. P-value was derived from Wilcoxon rank-sum test.
Extended Data Fig. 9 Higher order chromatin organization and DNA mismatch repair activity.
a, Average profiles of mutation loads across boundaries delineating inactive to active domains MSS and MSI samples from colorectal adenocarcinoma, uterine adenocarcinoma, gastric adenocarcinoma, stomach adenocarcinoma, glioblastoma multiforme (CNS-GBM), pancreas adenocarcinoma and liver cancer; with dashed lines for MSI samples. b, Number of mutations in colorectal adenocarcinoma samples per megabase active versus inactive domains. MSI sample is denoted with cyan color. c, Schematic describes the TAD boundary strength calculation. Briefly, for each TAD boundary we obtained a matrix around 400Kb up- and down-stream of the boundary by centering the TAD boundary in the middle. Next we calculated the log2 ratio of sum of interactions occurring within TADs and between TADs for each boundary region. d, Average TAD boundary strength profiles across all TAD boundaries (n = 2477) in healthy colon tissue and colon cancer cell line Hi-C data. The center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range.
Extended Data Fig. 10 APOBEC-mutagenesis affects the inactive X-chromosome mutation levels.
a, Dot plot demonstrates the association between the contribution of APOBEC-related mutation signatures (SBS2.13) and mutation distribution in active and inactive domains (x-axis). b, The distribution of APOBEC-signature enrichment subtypes (dark-green: A3A-like, gray: Not APOBEC-enriched) versus slope of mutations (active versus inactive domains). c, Scatter plot shows the preference of mutational signature toward TAD boundaries. Pearson correlation and the median slope calculations were described in the methods section. d, Distribution of X chromosome mutation load compared to total mutation burden for males (n:20) (observed and copy-number corrected values) and females (n:18) in lung adenocarcinoma patients. The center line is the median; box limits are the upper and lower quantiles; whiskers represent 1.5x the interquartile range. e, Dot plots show the enrichment of APOBEC signatures 2.13 and ratio of X-chromosome mutation load for female lung adenocarcinoma patients.
Supplementary information
Rights and permissions
About this article
Cite this article
Akdemir, K.C., Le, V.T., Kim, J.M. et al. Somatic mutation distributions in cancer genomes vary with three-dimensional chromatin structure. Nat Genet 52, 1178–1188 (2020). https://doi.org/10.1038/s41588-020-0708-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0708-0
This article is cited by
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Cell cycle gene alterations associate with a redistribution of mutation risk across chromosomal domains in human cancers
Nature Cancer (2024)
-
Tissue mosaicism following stem cell aging: blood as an exemplar
Nature Aging (2024)
-
Redistribution of mutation risk in cancer
Nature Cancer (2024)
-
Mutational signatures association with replication timing in normal cells reveals similarities and differences with matched cancer tissues
Scientific Reports (2023)