Abstract
De novo DNA methylation (DNAme) in mammalian germ cells is dependent on DNMT3A and DNMT3L. However, oocytes and spermatozoa show distinct patterns of DNAme. In mouse oocytes, de novo DNAme requires the lysine methyltransferase (KMTase) SETD2, which deposits H3K36me3. We show here that SETD2 is dispensable for de novo DNAme in the male germline. Instead, the lysine methyltransferase NSD1, which broadly deposits H3K36me2 in euchromatic regions, plays a critical role in de novo DNAme in prospermatogonia, including at imprinted genes. However, males deficient in germline NSD1 show a more severe defect in spermatogenesis than Dnmt3l−/− males. Notably, unlike DNMT3L, NSD1 safeguards a subset of genes against H3K27me3-associated transcriptional silencing. In contrast, H3K36me2 in oocytes is predominantly dependent on SETD2 and coincides with H3K36me3. Furthermore, females with NSD1-deficient oocytes are fertile. Thus, the sexually dimorphic pattern of DNAme in mature mouse gametes is orchestrated by distinct profiles of H3K36 methylation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Baylin, S. B. & Jones, P. A. Epigenetic determinants of cancer. Cold Spring Harb. Perspect. Biol. 8, a019505 (2016).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Velasco, G. & Francastel, C. Genetics meets DNA methylation in rare diseases. Clin. Genet. 95, 210–220 (2018).
Ley, T. J. et al. DNMT3A mutations in acute myeloid leukemia. N. Engl. J. Med. 363, 2424–2433 (2010).
Tatton-Brown, K. et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat. Genet. 46, 385–388 (2014).
Tlemsani, C. et al. SETD2 and DNMT3A screen in the Sotos-like syndrome French cohort. J. Med. Genet. 53, 743–751 (2016).
Choufani, S. et al. NSD1 mutations generate a genome-wide DNA methylation signature. Nat. Commun. 6, 10207 (2015).
Brennan, K. et al. NSD1 inactivation defines an immune cold, DNA hypomethylated subtype in squamous cell carcinoma. Sci. Rep. 7, 17064 (2017).
Papillon-Cavanagh, S. et al. Impaired H3K36 methylation defines a subset of head and neck squamous cell carcinomas. Nat. Genet. 49, 180–185 (2017).
Dhayalan, A. et al. The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J. Biol. Chem. 285, 26114–26120 (2010).
Dukatz, M. et al. H3K36me2/3 binding and DNA binding of the DNA methyltransferase DNMT3A PWWP domain both contribute to its chromatin interaction. J. Mol. Biol. 431, 5063–5074 (2019).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Weinberg, D. N. et al. The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature 573, 281–286 (2019).
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat. Rev. Genet. 9, 129–140 (2008).
Kurimoto, K. & Saitou, M. Germ cell reprogramming. Curr. Top. Dev. Biol. 135, 91–125 (2019).
Kaneda, M. et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429, 900–903 (2004).
Kato, Y. et al. Role of the Dnmt3 family in de novo methylation of imprinted and repetitive sequences during male germ cell development in the mouse. Hum. Mol. Genet. 16, 2272–2280 (2007).
Shirane, K. et al. Mouse oocyte methylomes at base resolution reveal genome-wide accumulation of non-CpG methylation and role of DNA methyltransferases. PLoS Genet. 9, e1003439 (2013).
Kobayashi, H. et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet. 8, e1002440 (2012).
Aravin, A. A., Sachidanandam, R., Girard, A., Fejes-Toth, K. & Hannon, G. J. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007).
Aravin, A. A. et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 31, 785–799 (2008).
Kuramochi-Miyagawa, S. et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 22, 908–917 (2008).
Barau, J. et al. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity. Science 354, 909–912 (2016).
Stewart, K. R. et al. Dynamic changes in histone modifications precede de novo DNA methylation in oocytes. Genes Dev. 29, 2449–2462 (2015).
Veselovska, L. et al. Deep sequencing and de novo assembly of the mouse oocyte transcriptome define the contribution of transcription to the DNA methylation landscape. Genome Biol. 16, 209 (2015).
Brind’Amour, J. et al. LTR retrotransposons transcribed in oocytes drive species-specific and heritable changes in DNA methylation. Nat. Commun. 9, 3331 (2018).
Xu, Q. et al. SETD2 regulates the maternal epigenome, genomic imprinting and embryonic development. Nat. Genet. 51, 844–856 (2019).
Bogutz, A. B. et al. Evolution of imprinting via lineage-specific insertion of retroviral promoters. Nat. Commun. 10, 5674 (2019).
Morselli, M. et al. In vivo targeting of de novo DNA methylation by histone modifications in yeast and mouse. eLife 4, e06205 (2015).
Erkek, S. et al. Molecular determinants of nucleosome retention at CpG-rich sequences in mouse spermatozoa. Nat. Struct. Mol. Biol. 20, 868–875 (2013).
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012).
Kobayashi, H. et al. High-resolution DNA methylome analysis of primordial germ cells identifies gender-specific reprogramming in mice. Genome Res. 23, 616–627 (2013).
Sangrithi, M. N. et al. Non-canonical and sexually dimorphic X dosage compensation states in the mouse and human germline. Dev. Cell 40, 289–301 (2017).
Brind’Amour, J. et al. An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat. Commun. 6, 6033 (2015).
Szabó, P. E., Hübner, K., Schöler, H. & Mann, J. R. Allele-specific expression of imprinted genes in mouse migratory primordial germ cells. Mech. Dev. 115, 157–160 (2002).
Lomelí, H., Mejía, V. R., Gertsenstein, M., Lobe, C. G. & Nagy, A. Targeted insertion of Cre recombinase into the TNAP gene: excision in primordial germ cells. Genesis 26, 116–117 (2000).
Park, I. Y. et al. Dual chromatin and cytoskeletal remodeling by SETD2. Cell 166, 950–962 (2016).
Zuo, X. et al. The histone methyltransferase SETD2 is required for expression of acrosin-binding protein 1 and protamines and essential for spermiogenesis in mice. J. Biol. Chem. 293, 9188–9197 (2018).
Edmunds, J. W., Mahadevan, L. C. & Clayton, A. L. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J. 27, 406–420 (2008).
Miura, F., Enomoto, Y., Dairiki, R. & Ito, T. Amplification-free whole-genome bisulfite sequencing by post-bisulfite adaptor tagging. Nucleic Acids Res. 40, e136 (2012).
Miura, F. et al. Highly efficient single-stranded DNA ligation technique improves low-input whole-genome bisulfite sequencing by post-bisulfite adaptor tagging. Nucleic Acids Res. 47, e85 (2019).
Kubo, N. et al. DNA methylation and gene expression dynamics during spermatogonial stem cell differentiation in the early postnatal mouse testis. BMC Genom. 16, 624 (2015).
Ooi, S. K. T. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007).
Jia, D., Jurkowska, R. Z., Zhang, X., Jeltsch, A. & Cheng, X. Structure of Dnmt3a bound to Dnmt3L suggests a model for de novo DNA methylation. Nature 449, 248–251 (2007).
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Streubel, G. et al. The H3K36me2 methyltransferase Nsd1 demarcates PRC2-mediated H3K27me2 and H3K27me3 domains in embryonic stem cells. Mol. Cell 70, 371–379 (2018).
Rayasam, G. V. et al. NSD1 is essential for early post‐implantation development and has a catalytically active SET domain. EMBO J. 22, 3153–3163 (2003).
Grzybowski, A. T., Chen, Z. & Ruthenburg, A. J. Calibrating ChIP-Seq with nucleosomal internal standards to measure histone modification density genome wide. Mol. Cell 58, 886–899 (2015).
Shah, R. N. et al. Examining the roles of H3K4 methylation states with systematically characterized antibodies. Mol. Cell 72, 162–177 (2018).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 39, 457–466 (2007).
Illingworth, R. S. et al. Orphan CpG islands identify numerous conserved promoters in the mammalian genome. PLoS Genet. 6, e1001134 (2010).
Singh, P. et al. De novo DNA methylation in the male germ line occurs by default but is excluded at sites of H3K4 methylation. Cell Rep. 4, 205–219 (2013).
Watanabe, T. et al. Role for piRNAs and noncoding RNA in de novo DNA methylation of the imprinted mouse Rasgrf1 locus. Science 332, 848–852 (2011).
Tomizawa, S.-I. et al. Dynamic stage-specific changes in imprinted differentially methylated regions during early mammalian development and prevalence of non-CpG methylation in oocytes. Development 138, 811–820 (2011).
Wang, L. et al. Programming and inheritance of parental DNA methylomes in mammals. Cell 157, 979–991 (2014).
Takahashi, N. et al. ZNF445 is a primary regulator of genomic imprinting. Genes Dev. 33, 49–54 (2019).
Bourc’his, D., Xu, G. L., Lin, C. S., Bollman, B. & Bestor, T. H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Bourc’his, D. & Bestor, T. H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 (2004).
Hata, K., Kusumi, M., Yokomine, T., Li, E. & Sasaki, H. Meiotic and epigenetic aberrations in Dnmt3L-deficient male germ cells. Mol. Reprod. Dev. 73, 116–122 (2006).
Inoue, K., Ichiyanagi, K., Fukuda, K., Glinka, M. & Sasaki, H. Switching of dominant retrotransposon silencing strategies from posttranscriptional to transcriptional mechanisms during male germ-cell development in mice. PLoS Genet. 13, e1006926 (2017).
Fang, D. et al. The histone H3.3K36M mutation reprograms the epigenome of chondroblastomas. Science 352, 1344–1348 (2016).
King, A. D. et al. Reversible regulation of promoter and enhancer histone landscape by DNA methylation in mouse embryonic stem cells. Cell Rep. 17, 289–302 (2016).
Lu, C. et al. Histone H3K36 mutations promote sarcomagenesis through altered histone methylation landscape. Science 352, 844–849 (2016).
Meng, X. et al. Regulation of cell fate decision of undifferentiated spermatogonia by GDNF. Science 287, 1489–1493 (2000).
Buageaw, A. et al. GDNF family receptor alpha1 phenotype of spermatogonial stem cells in immature mouse testes. Biol. Reprod. 73, 1011–1016 (2005).
Naughton, C. K., Jain, S., Strickland, A. M., Gupta, A. & Milbrandt, J. Glial cell-line derived neurotrophic factor-mediated RET signaling regulates spermatogonial stem cell fate. Biol. Reprod. 74, 314–321 (2006).
Jijiwa, M. et al. GDNF‐mediated signaling via RET tyrosine 1062 is essential for maintenance of spermatogonial stem cells. Genes Cells 13, 365–374 (2008).
Dahl, J. A. et al. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016).
Stroud, H. et al. Early-life gene expression in neurons modulates lasting epigenetic states. Cell 171, 1151–1164 (2017).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Hagarman, J. A., Motley, M. P., Kristjansdottir, K. & Soloway, P. D. Coordinate regulation of DNA methylation and H3K27me3 in mouse embryonic stem cells. PLoS ONE 8, e53880 (2013).
Reddington, J. P. et al. Redistribution of H3K27me3 upon DNA hypomethylation results in de-repression of Polycomb target genes. Genome Biol. 14, R25 (2013).
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010).
Zheng, Y. et al. Total kinetic analysis reveals how combinatorial methylation patterns are established on lysines 27 and 36 of histone H3. Proc. Natl Acad. Sci. USA 109, 13549–13554 (2012).
Yuan, W. et al. H3K36 methylation antagonizes PRC2-mediated H3K27 methylation. J. Biol. Chem. 286, 7983–7989 (2011).
Schmitges, F. W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol. Cell 42, 330–341 (2011).
Heyn, P. et al. Gain-of-function DNMT3A mutations cause microcephalic dwarfism and hypermethylation of Polycomb-regulated regions. Nat. Genet. 51, 96–105 (2019).
Sendžikaitė, G., Hanna, C. W., Stewart-Morgan, K. R., Ivanova, E. & Kelsey, G. A DNMT3A PWWP mutation leads to methylation of bivalent chromatin and growth retardation in mice. Nat. Commun. 10, 1884 (2019).
Yamanaka, S. et al. Broad heterochromatic domains open in gonocyte development prior to de novo DNA methylation. Dev. Cell 51, 21–34 (2019).
Zhou, W. et al. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat. Genet. 50, 591–602 (2018).
Strogantsev, R. et al. Allele-specific binding of ZFP57 in the epigenetic regulation of imprinted and non-imprinted monoallelic expression. Genome Biol. 16, 112 (2015).
Quenneville, S. et al. In embryonic stem cells, ZFP57/KAP1 recognize a methylated hexanucleotide to affect chromatin and DNA methylation of imprinting control regions. Mol. Cell 44, 361–372 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
de Vries, W. N. et al. Expression of Cre recombinase in mouse oocytes: a means to study maternal effect genes. Genesis 26, 110–112 (2000).
Shirane, K. et al. Global landscape and regulatory principles of DNA methylation reprogramming for germ cell specification by mouse pluripotent stem cells. Dev. Cell 39, 87–103 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Younesy, H., Möller, T., Lorincz, M. C., Karimi, M. M. & Jones, S. J. VisRseq: R-based visual framework for analysis of sequencing data. BMC Bioinform. 16, S2 (2015).
Hanna, C. W. et al. MLL2 conveys transcription-independent H3K4 trimethylation in oocytes. Nat. Struct. Mol. Biol. 25, 73–82 (2018).
Taudt, A., Nguyen, M. A., Heinig, M., Johannes, F. & Colomé-Tatché, M. chromstaR: tracking combinatorial chromatin state dynamics in space and time. Preprint at bioRxiv https://doi.org/10.1101/038612 (2016).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Liu, S. et al. Setdb1 is required for germline development and silencing of H3K9me3-marked endogenous retroviruses in primordial germ cells. Genes Dev. 28, 2041–2055 (2014).
Ramirez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Hahne, F. & Ivanek, R. Visualizing genomic data using gviz and bioconductor. Methods Mol. Biol. 1418, 335–351 (2016).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012).
Wang, H.-Q., Tuominen, L. K. & Tsai, C.-J. SLIM: a sliding linear model for estimating the proportion of true null hypotheses in datasets with dependence structures. Bioinformatics 27, 225–231 (2010).
Acknowledgements
We thank P. Chambon (University of Strasbourg) for the Nsd1 conditional mouse line, C. L. Walker (Baylor College of Medicine) for the Setd2 conditional mouse line and J. Trasler (McGill University) for the Dnmt3l KO mouse line; A. Bogutz, J. Brind’Amour, J. Richard Albert, K. Mochizuki and S. Janssen for helpful discussions and technical assistance; W. Xie (Tsinghua University) and L. Lefebvre (UBC) for critical reading of the manuscript; and R. Vander Werff and T. Stach (BRC-seq, UBC) for deep sequencing and A. Johnson and J. Wong (ubcFLOW, UBC) for cell sorting. This research was partially supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED (Japan Agency for Medical Research and Development) under grant number JP20am0101103 (support number 1080). M.C.L. is supported by CIHR grants PJT-153049 and PJT-166170. K.S. was a recipient of a Uehara Memorial Foundation postdoctoral fellowship and a CIHR Banting postdoctoral fellowship, BPF-156568.
Author information
Authors and Affiliations
Contributions
K.S. and M.C.L. conceived and designed the project and wrote the manuscript and all authors reviewed the manuscript. F.M. and T.I. performed the WGBS experiments. K.S. performed all other experiments and analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Setd2 deletion in male PGCs leads to hypogonadism in adults.
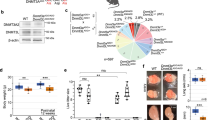
a, Schematic representation of the dynamics of DNA methylation (DNAme) during female (red) and male (blue) germline development. Developing germ cells analyzed in this study are shown in bold. Key events for germline development are also shown. Note that in both sexes, de novo DNAme occurs in non-dividing quiescent cells. PGC: primordial germ cell; SG: spermatogonium; E: embryonic day; P: postnatal day. b, Dynamics of Dnmt3l and Dnmt3a expression across key stages of male germ cell development. FPKM (fragments per kilobase of exon model per million reads mapped) values were extracted from RNA-seq data from a previous study34. The window of de novo DNAme during prenatal male germline development is shaded in yellow. c, Breeding scheme to obtain germline-specific Setd2 knockout (KO) prospermatogonia (PSG) and sorting strategy for purification of PSG using OCT3/4-EGFP36. Potential genotypes of embryos derived from this cross are indicated and a microscopy image of GFP expression in E16.5 testis is shown (scale bar, 500 μm). Note that OCT3/4-EGFP is expressed exclusively in PSG in the testis. Dead cells were excluded by gating out PI positive cells during FACS. Sorted PSG (3.9% of total testis cells) were processed as previously described for ULI-N-ChIP-seq35 and PBAT41,42. d, Gross morphology of 12-week-old testis isolated from WT control (left) or Setd2 KO mice (right). Image is representative of three litters. Scale bar, 5 mm. e, Hematoxylin and eosin (HE) staining of 12-week-old testis (top) and epididymis (bottom) isolated from WT control (left) or Setd2 KO mice (right). Histological analyses shown are representative of multiple imaged sections from a single experiment. Scale bar, 50 μm.
Extended Data Fig. 2 Genome-wide relationship between de novo DNAme in prenatal male germ cells and histone H3 methylation states.
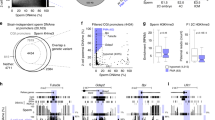
a, Contour scatter plots with outliers showing the relationship between enrichment (RPKM) of the indicated histone marks over 10-kb genomic bins and the difference in the % of DNAme between E16.5 PSG and E13.5 mPGCs33 in the same regions. The % of genomic coverage for each histone mark in E16.5 PSG is shown in each plot. R indicates Spearman’s rank correlation coefficient. b, Genome browser representation of a region on chromosome 3 showing % DNAme in E13.5 mPGCs and E16.5 PSG33 as well as RPM values of H3K4me3, H3K27me3, H3K9me3, H3K36me2 and H3K36me3 in E16.5 PSG. c, Violin plots showing % DNAme over H3K9me3 domains of <10 kb (total DNAme genomic coverage, 0.1 Gb) or >20 kb (total DNAme genomic coverage, 0.2 Gb) in E13.5 mPGCs, E16.5 PSG33 and P0 PSG43 (n = 1). The % of the genome in E16.5 PSG showing H3K9me3 enrichment in domains of <10 kb or >20 kb is shown in each plot. Box (25-75th percentile), median (inside the box) and whisker (1.5x interquartile range from the box) are included in the violin plots. d, Violin plots showing the distribution of enrichment of H3K36me2 over 10-kb genomic bins (left) or H3K9me3 domains of <10 kb (middle) or >20 kb (right) in E16.5 PSG (n = 1). Note that H3K36me2 is particularly depleted in regions of broad H3K9me3. Box, median and whisker are included in the violin plots as shown in c. e, Stacked bar chart showing % total of unique or repeat regions in the genome (total) versus H3K9me3 domains of <10 kb or >20 kb. Repeats were identified by RepeatMasker annotation.
Extended Data Fig. 3 Expression of H3K36 KMTases and the role of H3K36me2 during male germline development.
a, Dynamics of expression of H3K36 KMTases across prenatal male germ cell development. FPKM values were extracted from RNA-seq data from a previous study34. The window of global de novo DNAme is shaded in yellow. b, Breeding scheme to obtain germline-specific Nsd1 KO PSG and sorting strategy for PSG purification, as in Extended Data Fig. 1c. A microscopy image of GFP expression in E16.5 testis is shown (scale bar, 500 μm) as well as a representative FACS plot for control testis, with PSG representing 3.1% of total testis cells in this experiment. c, Tracks showing coverages and junctions of RNA-seq reads over the Nsd1 locus in Nsd1 HET and KO E16.5 PSG. Note that the exon between the two loxP sites (shaded in yellow) is efficiently excised (>99%) in the KO. d, Bar graph showing the % genomic coverage of H3K36me2 and H3K36me3 in Nsd1 HET versus KO E16.5 PSG. e, Scatterplot showing the correlation of H3K9me3 (RPKM) over 10-kb genomic bins in Nsd1 HET versus KO E16.5 PSG. f, Scatterplot showing H3K9me3 versus H3K36me2 enrichment (RPKM over 10-kb genomic bins) in Nsd1 KO E16.5 PSG.
Extended Data Fig. 4 NSD1 is required for de novo DNAme of a subset of CpG islands hypermethylated in sperm.
a, Histogram showing the frequency distribution of CpG island (CGI) % DNAme (mean across each annotated CGI) in mature sperm. WGBS data from sperm43 was re-processed for this analysis. b, Stacked bar chart showing relative % of CGIs in promoter, gene body or intergenic regions for all annotated CGIs52 in the genome (total) versus CGIs hypermethylated (>70%) in sperm. c, Parallel coordinate plot showing % DNAme over all CGIs in the indicated cell types during male germline development. Color coded according to methylation state in sperm. d, Scatterplot showing the relationship between % DNAme in Nsd1 HET versus KO P0 PSG for 50,000 randomly selected 10 kb genomic bins versus at CGIs hypermethylated in WT sperm. e, Heatmaps showing the levels of DNAme in Nsd1 HET or KO P0 PSG and sperm43 as well as H3K4me3 and H3K36me2 in Nsd1 HET or KO P0 PSG over CGIs and flanking (+/-0.5 kb) regions. The data is sorted based on H3K4me3 in Nsd1 KO P0 PSG. f, Genome browser representation of % DNAme and RPM values of H3K4me3 and H3K36me2 (spike-in normalized) in the indicated cells at representative CGIs (hatched boxes) and their flanking regions (left: non-promoter CGI, right: promoter CGI). WGBS data from E13.5 mPGCs33 was re-processed. g, Scatterplot showing the relationship between fold-change of gene expression and the difference in % DNAme between Nsd1 HET and KO P0 PSG at hypermethylated CGI promoters (as defined in panel b). Genes showing a > 20% difference in DNAme and a 2-fold difference in expression are labeled.
Extended Data Fig. 5 Comparison of germ cell phenotypes in Nsd1 KO versus Dnmt3l KO male mice.
a, Gross morphology of testes isolated from Nsd1 HET and KO (top) as well as Dnmt3l HET and KO (bottom) mice at P10, P14, P21 and 3 months. Images shown for 3 month testes are representative of three independent experiments. Scale bar, 5 mm. b, HE staining of sections of testis and epididymis isolated from 3-month-old Nsd1 HET and KO as well as Dnmt3l HET and KO mice. Histological analyses shown are representative of multiple imaged sections from a single experiment. Note the complete absence of spermatogonia in Nsd1 KO testis. Scale bar, 50 μm. c, Gross morphology of perinatal testes isolated from Oct3/4-EGFP Nsd1 HET and KO (top) as well as Dnmt3l HET and KO mice (bottom). Note that OCT3/4-EGFP is expressed exclusively in PSG in the testis. Scale bar, 500 μm.
Extended Data Fig. 6 Silencing of TEs in male germ cells is dependent upon DNMT3L but not NSD1.
a, Scatterplots showing the expression (agglomerated RNA-seq reads) of LTR (top, n = 539) or L1 (bottom, n = 145) families in Nsd1 HET versus KO P0 PSG (left panels) as well as Dnmt3l HET versus KO (right panels). A color gradient of H3K4me3 RPKM values in Nsd1 HET E16.5 PSG is also shown. Only TE families with >100 genomic copies are plotted and TE families showing a 2-fold increase in expression in the Dnmt3l KO are labelled. b, Immunofluorescence (IF) analysis of LINE1 ORF1 expression in 3-week testis isolated from HET control and Nsd1 KO or Dnmt3l KO mice. Testis were co-stained with the germ cell specific marker TRA98. IF analyses shown are representative of multiple imaged sections from a single experiment. Scale bar, 25 μm. c, Scatterplots showing the correlation of mean % DNAme of LTR (top) or L1 families (bottom) in Nsd1 HET versus KO P0 PSG. A color gradient of H3K36me2 RPKM values in Nsd1 HET E16.5 PSG is also shown. Only TE families with >100 genomic copies are plotted. d, Scatterplots showing the correlation of mean % DNAme of LTR or L1 families in Nsd1 HET versus KO P0 PSG as well as Dnmt3l or Dnmt3c HET versus KO P10 male germ cells24. RPKM values for H3K4me3 in Nsd1 HET E16.5 PSG is shown via color gradient in all plots. Only TEs with >100 genomic copies are plotted.
Extended Data Fig. 7 Effects of Nsd1 or Dnmt3l deletion on H3K36me2, H3K27me3 and gene expression in E16.5 PSG.
a, Pie chart showing the relative prevalence of promoter, intergenic or gene body regions enriched for both H3K36me2 and H3K36me3 in E16.5 PSG. Peaks were annotated using ChIPseeker96. b, Scatterplot showing H3K36me2 versus H3K27me3 enrichment (RPKM) over 10-kb genomic bins in Nsd1 HET E16.5 PSG. 50,000 randomly selected data points are plotted. The color gradient shows the difference in % DNAme between Nsd1 control HET versus KO E16.5 PSG. c, Scatterplot showing the correlation of H3K36me2 (RPKM) over 10-kb genomic bins in Dnmt3l HET versus KO E16.5 PSG. d, Scatterplot showing the correlation of H3K27me3 (RPKM) over 10-kb genomic bins in Dnmt3l HET versus KO E16.5 PSG. e, Contour scatter plot and the outliers showing the relationship between log2 fold-change of H3K27me3 over 10-kb genomic bins in Nsd1 HET versus KO and Dnmt3l HET versus KO E16.5 PSG. f, Scatterplot showing the fold-change of gene expression in E16.5 Nsd1 HET versus KO PSG. Only genes showing total count >10 in Nsd1 HET and KO are plotted. Genes showing a > 2-fold difference in expression with p < 0.05 (Wald test by DESeq284) in HET and KO groups (n = 2) were scored as up- or down-regulated and are highlighted as indicated. g, Scatterplot showing the fold-change of gene expression in E16.5 Dnmt3l HET versus KO PSG. Only genes showing total count >10 in Dnmt3l HET and KO are plotted. Genes showing a > 2-fold difference in expression with p < 0.05 (Wald test by DESeq284) in HET and KO groups (n = 2) were scored as up- or down-regulated and are highlighted as indicated.
Extended Data Fig. 8 Aberrant repression of PRC2 target genes in Nsd1 KO PSG.
a, Violin plots showing the distribution of expression levels (log2 RPKM + 0.01) (n = 2) of all genes in Nsd1 HET E16.5 PSG (left) versus genes downregulated in the Nsd1 KO. b, Scatterplot showing the relationship between H3K36me2 (RPKM) and H3K36me3 over gene body regions of all expressed (FPKM > 1) genes in control E16.5 PSG (grey) or genes downregulated in Nsd1 KO E16.5 PSG (blue). c, Scatterplot showing the relationship between the enrichment of H3K4me3 and H3K27me3 at all genic TSSs (±1 kb) in E16.5 PSG. The expression levels of associated genes in control E16.5 PSG is indicated via color gradient. Genes with H3K4me3 RPKM > 2 and H3K27me3 RPKM > 0.5 in E16.5 PSG were defined as bivalent (hatched boxes). d, Venn diagram showing the overlap of genes downregulated in Nsd1 KO E16.5 PSG with bivalent expressed genes in Nsd1 HET E16.5 PSG. e, Genome browser representation of % DNAme and RPM values of H3K36me2, H3K4me3 and H3K27me3 at the Gfra1 locus in the indicated cells. H3K36me2 and H3K27me3 levels are normalized by spike-in. Published WGBS data from E13.5 mPGCs33 was reprocessed. f, Bar graph showing the expression of Gfra1 in Nsd1 KO and Dnmt3l KO E16.5 and P0 PSG relative to their matching controls. g, Bar graph showing the expression of key transcription factors acting downstream of the GDNF/GFRa1 signaling pathway in Nsd1 KO and Dnmt3l KO E16.5 and P0 PSG relative to their matching controls.
Extended Data Fig. 9 SETD2 is required for intragenic H3K36me2 and H3K36me3 in oocytes.
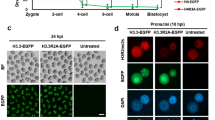
a, Violin plots showing % DNAme over 10-kb genomic bins (left, total genomic coverage, 2.5 Gb), H3K36me2 enriched regions (middle, genomic coverage, 0.8 Gb) or H3K36me3 enriched regions (right, genomic coverage, 0.6 Gb) in P0-3 NGOs, P12 GOs and FGOs (n = 1). The % genomic coverage of H3K36me2 (middle) and H3K36me3 (right) in P12 GOs is shown in each plot. Box (25-75th percentile), median (inside the box) and whisker (1.5x interquartile range from the box) are included in the violin plots. NGO: non-growing oocyte; GO: growing oocyte; FGO: fully-grown oocyte. Publicly available WGBS data from P0-3 NGOs, FGOs19 and P12 GOs69 were reprocessed. b, Scatterplot showing H3K36me2 versus H3K36me3 enrichment (RPKM) over 10-kb genomic bins in P12 GOs. 50,000 randomly selected data points are plotted and the color gradient shows the difference in % DNAme between P12 GOs and NGOs. c, Heatmaps showing the levels of DNAme, H3K36me2, H3K36me3, H3K4me3 and H3K27me3 in gene bodies and their flanking 3 kb regions in control or Setd2 KO FGOs. Genes with expression RPKM values >1 in E16.5 PSGs and/or FGOs26,27 were defined as expressed. Only forward strand genes are plotted and the heatmaps are sorted by the level of H3K36me3 in Setd2 control FGOs. TSS: transcription start site; TTS: transcription termination site. Publicly available ChIP-seq data from H3K4me3 and H3K27me394 as well as WGBS data from Setd2 control and KO FGOs28 were reprocessed. d, Bar graph showing the % genomic coverage of H3K36me2 and H3K36me3 in control or Setd2 KO FGOs. Note that the majority of H3K36me2 is dependent upon SETD2. e, Pie chart showing the relative prevalence of promoter, intergenic or gene body regions with residual H3K36me2 enrichment in Setd2 KO FGOs. f, Tracks showing coverages and junctions of RNA-seq reads over the Nsd1 locus in oocytes26 and PSG. Note the predominant form of Nsd1 initiates from an LTR in oocytes. g, Relative junction read count of the canonical exon 8-exon 9 junction versus the alternative LTR-exon 9 junction in GOs and FGOs. h, Domain structure of canonical NSD1 versus the predicted oocyte-specific, LTR-initiated isoform. Note that the oocyte-specific NSD1 lacks the amino-terminal PWWP domain of canonical NSD1.
Supplementary information
Supplementary Information
Supplementary Note
Supplementary Tables
Supplementary Tables 1–9
Rights and permissions
About this article
Cite this article
Shirane, K., Miura, F., Ito, T. et al. NSD1-deposited H3K36me2 directs de novo methylation in the mouse male germline and counteracts Polycomb-associated silencing. Nat Genet 52, 1088–1098 (2020). https://doi.org/10.1038/s41588-020-0689-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0689-z
This article is cited by
-
Imprinting at the KBTBD6 locus involves species-specific maternal methylation and monoallelic expression in livestock animals
Journal of Animal Science and Biotechnology (2023)
-
Characterization of H3K9me3 and DNA methylation co-marked CpG-rich regions during mouse development
BMC Genomics (2023)
-
Epigenetic markers in the embryonal germ cell development and spermatogenesis
Basic and Clinical Andrology (2023)
-
Emerging evidence that the mammalian sperm epigenome serves as a template for embryo development
Nature Communications (2023)
-
Mechanisms and function of de novo DNA methylation in placental development reveals an essential role for DNMT3B
Nature Communications (2023)