Abstract
Three-dimensional organization of the genome is important for transcriptional regulation1,2,3,4,5,6,7. In mammals, CTCF and the cohesin complex create submegabase structures with elevated internal chromatin contact frequencies, called topologically associating domains (TADs)8,9,10,11,12. Although TADs can contribute to transcriptional regulation, ablation of TAD organization by disrupting CTCF or the cohesin complex causes modest gene expression changes13,14,15,16. In contrast, CTCF is required for cell cycle regulation17, embryonic development and formation of various adult cell types18. To uncouple the role of CTCF in cell-state transitions and cell proliferation, we studied the effect of CTCF depletion during the conversion of human leukemic B cells into macrophages with minimal cell division. CTCF depletion disrupts TAD organization but not cell transdifferentiation. In contrast, CTCF depletion in induced macrophages impairs the full-blown upregulation of inflammatory genes after exposure to endotoxin. Our results demonstrate that CTCF-dependent genome topology is not strictly required for a functional cell-fate conversion but facilitates a rapid and efficient response to an external stimulus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The Hi-C, RNA-seq, CTCF ChIP–seq, ATAC-seq datasets generated and analyzed for the current study are available in the Gene Expression Omnibus (GEO) database under accession number GSE140528. ATAC-seq and CEBPA ChIP–seq datasets used in the current study are available in the GEO database under accession number GSE131620. The H3K27ac and H3K4me1 ChIP–seq datasets used in this study are available in the ArrayExpress database under accession number E-MTAB-9010. Source data for Fig. 2c are provided with the paper.
References
de Laat, W. & Duboule, D. Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502, 499–506 (2013).
Gorkin, D. U., Leung, D. & Ren, B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14, 762–775 (2014).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Furlong, E. E. M. & Levine, M. Developmental enhancers and chromosome topology. Science 361, 1341–1345 (2018).
Stadhouders, R., Filion, G. J. & Graf, T. Transcription factors and 3D genome conformation in cell-fate decisions. Nature 569, 345–354 (2019).
Kim, S. & Shendure, J. Mechanisms of interplay between transcription factors and the 3D genome. Mol. Cell 76, 306–319 (2019).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Hou, C., Li, L., Qin, Z. S. & Corces, V. G. Gene density, transcription, and insulators contribute to the partition of the Drosophila genome into physical domains. Mol. Cell 48, 471–484 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944.e22 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707.e14 (2017).
Heath, H. et al. CTCF regulates cell cycle progression of αβ T cells in the thymus. EMBO J. 27, 2839–2850 (2008).
Arzate-Mejía, R. G., Recillas-Targa, F. & Corces, V. G. Developing in 3D: the role of CTCF in cell differentiation. Development 145, dev137729 (2018).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Beagan, J. A. & Phillips-Cremins, J. E. On the existence and functionality of topologically associating domains. Nat. Genet. 52, 8–16 (2020).
Rapino, F. et al. C/EBPα induces highly efficient macrophage transdifferentiation of B lymphoma and leukemia cell lines and impairs their tumorigenicity. Cell Rep. 3, 1153–1163 (2013).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Stadhouders, R. et al. Transcription factors orchestrate dynamic interplay between genome topology and gene regulation during cell reprogramming. Nat. Genet. 50, 238–249 (2018).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
Ouboussad, L., Kreuz, S. & Lefevre, P. F. CTCF depletion alters chromatin structure and transcription of myeloid-specific factors. J. Mol. Cell Biol. 5, 308–322 (2013).
Nikolic, T. et al. The DNA-binding factor Ctcf critically controls gene expression in macrophages. Cell. Mol. Immunol. 11, 58–70 (2014).
Cuartero, S. et al. Control of inducible gene expression links cohesin to hematopoietic progenitor self-renewal and differentiation. Nat. Immunol. 19, 932–941 (2018).
Parelho, V. et al. Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 132, 422–433 (2008).
Wendt, K. S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Faridi, M. H. et al. CD11b activation suppresses TLR-dependent inflammation and autoimmunity in systemic lupus erythematosus. J. Clin. Invest. 127, 1271–1283 (2017).
Bell, A. C., West, A. G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Serra, F. et al. Automatic analysis and 3D-modelling of Hi-C data using TADbit reveals structural features of the fly chromatin colors. PLoS Comput. Biol. 13, e1005665 (2017).
Despang, A. et al. Functional dissection of the Sox9–Kcnj2 locus identifies nonessential and instructive roles of TAD architecture. Nat. Genet. 51, 1263–1271 (2019).
Ghavi-Helm, Y. et al. Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression. Nat. Genet. 51, 1272–1282 (2019).
Williamson, I. et al. Developmentally regulated Shh expression is robust to TAD perturbations. Development 146, dev179523 (2019).
Le Dily, F. L. et al. Distinct structural transitions of chromatin topological domains correlate with coordinated hormone-induced gene regulation. Genes Dev. 28, 2151–2162 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Ay, F. et al. Identifying multi-locus chromatin contacts in human cells using tethered multiple 3C. BMC Genomics 16, 121 (2015).
Vidal, E. et al. OneD: increasing reproducibility of Hi-C samples with abnormal karyotypes. Nucleic Acids Res. 46, e49 (2018).
Ramírez, F. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Flyamer, I. M., Illingworth, R. S. & Bickmore, W. A. Coolpup.py: versatile pile-up analysis of Hi-C data. Bioinformatics 36, 2980–2985 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. DeepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Miguel-Escalada, I. et al. Human pancreatic islet three-dimensional chromatin architecture provides insights into the genetics of type 2 diabetes. Nat. Genet. 51, 1137–1148 (2019).
Stefano, M. Di et al. Dynamic simulations of transcriptional control during cell reprogramming reveal spatial chromatin caging. Preprint at https://doi.org/10.1101/642009 (2019).
Baù, D. & Marti-Renom, M. A. Genome structure determination via 3C-based data integration by the Integrative Modeling Platform. Methods 58, 300–306 (2012).
Pettersen, E. F. et al. Chimera - a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We thank M. T. Kanemaki for the degron plasmids; R. Guigó’s laboratory, and S. Pérez-Lluch in particular, for the H3K27ac and H3K4me1 ChIP–seq, produced in the framework of the RNA-MAPS project (ERC-2011-AdG-294653-RNA-MAPS); Y. Cuartero for help with sequencing and CTCF ChIP–seq; C. Segura for help with immunofluorescence microscopy; the CRG Genomics and flow cytometry core facilities and the CRG-CNAG Sequencing Unit for sequencing; and members of T.G.’s laboratory for discussions. This work was supported by the European Research Council under the 7th Framework Programme FP7/2007-2013 (ERC Synergy Grant 4D-Genome, grant agreement 609989, to T.G. and M.A.M.-R.), the Ministerio de Educación y Ciencia (SAF.2012-37167, to T.G., and BFU2017-85926-P, to M.A.M.-R.), the AGAUR (to T.G.) and the Marató TV3 (201611) (to M.A.M.-R.). P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB860, SPP1935, EXC 2067/1-390729940), the European Research Council (advanced investigator grant TRANSREGULON, grant agreement no. 693023) and the Volkswagen Foundation. G.S. was supported by a Marie Skłodowska-Curie fellowship (H2020-MSCA-IF-2016, miRStem) and by the ‘Fundación Científica de la Asociación Española Contra el Cáncer’. T.V.T. was supported by Juan de la Cierva postdoctoral fellowship (MINECO; FJCI-2014-22946). B.B. was supported by the fellowship 2017FI_B00722 from the Secretaria d’Universitats i Recerca del Departament d’Empresa i Coneixement (Generalitat de Catalunya) and the European Social Fund (ESF). R.S. was supported by a Netherlands Organisation for Scientific Research Veni fellowship (91617114) and an Erasmus MC Fellowship. We also acknowledge support from ‘Centro de Excelencia Severo Ochoa 2013-2017’ (SEV-2012-0208), the Spanish Ministry of Science and Innovation to the EMBL partnership and the CERCA Program Generalitat de Catalunya to the CRG, as well as the support of the Spanish Ministry of Science and Innovation through the Instituto de Salud Carlos III, the Generalitat de Catalunya through Departament de Salut and Departament d’Empresa i Coneixement, and co-financing by the Spanish Ministry of Science and Innovation with funds from the European Regional Development Fund (ERDF) corresponding to the 2014-2020 Smart Growth Operating Program to CNAG.
Author information
Authors and Affiliations
Contributions
G.S., R.S. and T.G. conceived the study and wrote the manuscript with input from all coauthors. G.S. performed molecular biology, RNA-seq, ChIP–seq and in situ Hi-C experiments. T.V.T., J.C. and A.A. performed ChIP–seq and ATAC-seq. G.S., E.V., M.V.-C., J.M.-E. and B.B. performed bioinformatic analyses. G.S., E.V., M.V.-C., J.M.-E. and R.S. integrated and visualized data. G.S. and M.B. performed the CTCF-degron CRISPR targeting and S.C. performed cytokine arrays. G.S. performed the transdifferentiation experiments with help from M.B. and C.B. F.L.D., P.C., M.A.M.-R. and R.S. provided valuable advice and T.G. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
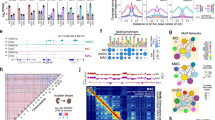
Extended Data Fig. 1 Characterization of chromatin compartmentalization and TAD dynamics during transdifferentiation.
a, Genome-wide Pearson correlation matrix between PC1 values of Hi-C samples at different time points. b, Scatter plots of PC1 values (n = 1,332 100-kb bins) showing changes relative to initial B cell genome compartmentalization for chromosome 12. c, Line chart depicting fractions of the genome assigned to A or B compartments at 10 time points during transdifferentiation. Y-axis represents the number of 100-kb bins. d, Gene ontology analysis of genes in regions switching from B to A (n = 980 genes) or A to B (n = 1,815 genes) compartments (P values, FDR corrected Fisher test). e, CTCF binding signal at TADs, normalized for TAD size in samples at various transdifferentiation time points. f, PCA of insulation score values at TAD borders during transdifferentiation (n = 4,006 TAD borders). Grey arrow depicts an averaged trajectory. g, RNA expression of genes at TAD borders gained (n = 254 genes) or lost (n = 293 genes) during transdifferentiation. All box plots depict the first and third quartiles as the lower and upper bounds of the box, with a thicker band inside the box showing the median value and whiskers representing 1.5x the interquartile range. h, Homer DNA motif analysis at ATAC-seq peaks detected at stable (n = 2,044), gained (n = 591) or lost (n = 135) TAD borders (P values are calculated using hyper-geometric statistical tests).
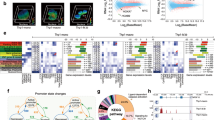
Extended Data Fig. 2 Molecular characterization of CTCF-mAID BLaER cells during transdifferentiation.
a, Heatmap of CTCF binding signal at CTCF ChIP-seq peaks detected in untreated CTCF-mAID cells. b, Browser snapshot showing CTCF binding loss upon 24 h of auxin treatment. c, Overall scaling of Hi-C contact frequency as a function of genomic distance in cells treated with DMSO or auxin. d, Venn diagram showing the overlap of TAD borders detected in B cells and in CTCF-AID B cells. e, Scatter plots comparing insulation scores at TAD borders at B cell and at CTCF-AID B cells. Lower values indicate stronger insulation. f, Top: Representative in situ Hi-C contact maps (20-kb resolution) of iMacs obtained after treatment with DMSO or auxin. Color scale represents the normalized number of contacts. Bottom: plots of the corresponding insulation scores for each bin within the 10-Mb region shown. g, Scatter plots comparing insulation scores at TAD borders at 24 hpi and at the iMac stage after DMSO or auxin treatment. Lower values indicate stronger insulation. h, Contact enrichment of interactions inside TADs versus outside TADs at the indicated time points for B cell (n = 1), DMSO- (n = 2) or auxin-treated cell (n = 2) biologically independent samples. Dots represent point estimates and bars (wide and narrow) indicate confidence intervals (50% and 95 %, respectively) for the log2 fold changes. All estimations are computed using all 9 samples in a single linear mixed model. i, Outline of strategy used to identify dynamic promoters and enhancers during transdifferentiation. Numbers of ATAC-seq peaks intersecting with TSS (promoters) and H3K4me1 peaks (enhancers) are indicated. j, H3K27ac decoration at dynamic promoters and enhancers that become either inactivated or activated. k, CEBPA binding at activated and inactivated regulatory elements (RE) in B cell and iMacs. l, RNA expression of genes associated with inactivated (n = 1,259) and activated (n = 1,421) promoters during transdifferentiation. All box plots depict the first and third quartiles as the lower and upper bounds of the box, with a thicker band inside the box showing the median value and whiskers representing 1.5x the interquartile range.
Extended Data Fig. 3 CTCF depletion does not impair transdifferentiation or long-range enhancer-promoter contact dynamics.
a, Representative flow cytometry analysis of CD19 and Mac-1 marker expression during transdifferentiation of CTCF-mAID B cells treated with DMSO or auxin. The experiment was repeated 3 times with similar results. b, Transdifferentiation kinetics of CTCF-mAID B cells (clone C1) in the presence of DMSO or auxin analysed at 0, 96 and 168 hpi by flow cytometry for CD19 and Mac-1 expression (n = 3 biologically independent samples). Centre indicates mean, error bars show standard deviation and P unpaired two-tailed t-test. c, Phagocytosis assay of iMacs analyzed by flow cytometry showing uptake of blue fluorescent beads. The experiment was repeated 3 times with similar results. d, RNA expression measured by qRT-PCR of cytokines in noninduced (NI) or 2h LPS-induced iMacs DMSO (n = 6), iMacs AUX (n = 6) or B cells (n = 3). Mean values are shown, error bars represent standard error and n represents biologically independent samples. e, Venn diagram showing the overlap of genes upregulated (left) and downregulated (right) in iMacs after transdifferentiation in the presence of DMSO or auxin based on RNA-Seq (n = 2 biologically independent samples). f, Gene ontology analysis of genes specifically upregulated (n = 419) and downregulated (n = 744) specifically in CTCF-depleted iMacs (q-value, FDR corrected Fisher exact test). g, Aggregate metaplots (10-kb resolution) depicting long range (5–10-Mb) interaction frequencies between enhancers and promoter (E-P) during transdifferentiation. Area shown is centered on enhancers or promoters ± 250-kb). h, Venn diagram showing the number of switching compartment regions (100-kb bins) during transdifferentiation in presence of DMSO or auxin. i, Expression of MAFB during transdifferentiation with or without CTCF, as measured by RNA-seq (n = 2 biologically independent samples, lines connect mean values). j, Enhancer activity at the MAFB locus during transdifferentiation. Browser snapshot showing H3K27ac ChIP-seq profiles of a 4-Mb domain surrounding the MAFB locus. The enhancer and the promoter shown in Fig. 3g are highlighted in light brown.
Extended Data Fig. 4 CTCF depletion in iMacs attenuates the acute inflammatory response to endotoxins.
a, Genome-wide aggregation of normalized Hi-C signal anchored at cohesin loops during transdifferentiation with DMSO or auxin. b, Distance distribution between enhancers and TSS of genes responsive (n = 378) or unresponsive (n = 380) to LPS (P, two-sided Wilcoxon rank-sum test). c, CTCF enrichment at promoters and enhancers d, of genes responsive (n = 378) or unresponsive (n = 378) to LPS (P, two-sided Wilcoxon rank-sum test). e, Differential gene expression between LPS-induced iMac treated with auxin or DMSO (n = 2 biologically independent samples, P-adj two-tailed likelihood ratio test followed by FDR correction). f, Gene ontology analysis of the significantly (p < 0.01) upregulated (n = 746) and downregulated (n = 694) genes in LPS-induced iMacs treated with auxin compared to DMSO (q-value, FDR-corrected Fisher exact test). g, Overlap of upregulated (top) and downregulated (bottom) genes (AUX vs DMSO) between non-induced iMacs (NI) and iMacs treated with LPS (LPS). h, LPS-upregulated genes in iMacs exposed to DMSO compared to auxin (n = 2,470 genes, P two-sided Wilcoxon rank-sum test). i, RNA expression in non-induced (NI) iMacs of genes upregulated after LPS stimulation of DMSO treated iMacs (n = 2,470). j, RNA expression of key transcription factors and receptors of the LPS signalling pathway (n = 2 biologically independent samples). k, CTCF binding at promoters of genes deregulated in LPS-induced iMacs treated with auxin as compared to DMSO. l, Micrographs show uptake of fluorescent beads (shown in red) by iMacs treated with DMSO or auxin (Scale bar represents 10 µm). The experiment was repeated 3 times with similar results. m, Quantification of phagocytosis assay. Upper panel shows percentage of cells with bead uptake; lower panel shows mean fluorescent intensity (MFI). Bars represent mean values of n = 3 biologically independent samples and error bars denote standard deviation. All box plots depict the first and third quartiles as the lower and upper bounds of the box, with a thicker band inside the box showing the median value and whiskers representing 1.5x the interquartile range.
Extended Data Fig. 5 CTCF depletion in iMacs impairs 3D chromatin organization at inflammatory response gene loci.
a, Distance distribution between promoters and their closest TAD borders of genes downregulated in auxin treated iMacs after LPS induction (as compared to iMacs exposed to DMSO) or for a random set of genes with a similar size (n = 687) (P, two-sided Wilcoxon rank-sum test). Box plots depict the first and third quartiles as the lower and upper bounds of the box, with a thicker band inside the box showing the median value and whiskers representing 1.5x the interquartile range. b, Average insulation scores of TAD borders closest to genes downregulated in auxin treated iMacs after LPS induction or closest to a random set of genes with a similar size (n = 687). Area shown is centered on boundary regions ± 250-kb (P, two-sided Wilcoxon rank-sum test). c, Hi-C maps (10-kb resolution) at the IL6 locus. Color scale represents the normalized number of contacts and the genes within the locus are indicated on the right. d, Virtual 4 C of iMacs treated with DMSO (dark blue) or auxin (light blue) using IL6 enhancer 1 (e1) as viewpoint; Browser snapshot of H3K27ac ChIP-seq signal is shown and the STEAP1B promoter is highlighted. e, Differential expression of IL6 and STEAP1B in LPS-induced iMacs treated with auxin as compared to DMSO (bars represent the mean values of n = 2 biologically independent samples). f, Distance distribution between STEAP1B promoter and IL6 enhancer regions (n = 1,000 models). Median (solid line), first and third quartile (dashed line) are indicated (P, two-sided Komogorov-Smirnov test). g, Hi-C maps (10-kb resolution) at the CCL2 locus in iMacs generated in the presence of DMSO or auxin. Color scale represents the normalized number of contacts. Genes within the locus are indicated on the right. h, Top: Differential Hi-C maps of the CCL2 locus (10-kb resolution) in iMacs generated in the presence of DMSO or auxin; CTCF ChIP-seq signal and gene positions are shown below the Hi-C map. Middle: Virtual 4 C of the CCL2 locus of iMacs treated with DMSO (dark blue) or auxin (light blue), using CCL2 promoter as viewpoint. Bottom: browser snapshots showing H3K27ac ChIP-seq and PC1 A/B compartment tracks. The CCL2 enhancer is highlighted. i, Distance distribution between CCL2 TSS and enhancer regions (n = 1,000 models). Median (solid line), first and third quartiles (dashed line) are indicated (P, two-sided Komogorov-Smirnov test). j, 3D model of the CCL2 locus in DMSO or auxin treated iMacs.
Supplementary information
Supplementary Information
Supplementary Note
Source data
Source Data Fig. 2
Unmodified western blot (Fig. 2c).
Rights and permissions
About this article
Cite this article
Stik, G., Vidal, E., Barrero, M. et al. CTCF is dispensable for immune cell transdifferentiation but facilitates an acute inflammatory response. Nat Genet 52, 655–661 (2020). https://doi.org/10.1038/s41588-020-0643-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0643-0
This article is cited by
-
Cellular reprogramming is driven by widespread rewiring of promoter-enhancer interactions
BMC Biology (2023)
-
LAST-seq: single-cell RNA sequencing by direct amplification of single-stranded RNA without prior reverse transcription and second-strand synthesis
Genome Biology (2023)
-
Three-dimensional genome organization in immune cell fate and function
Nature Reviews Immunology (2023)
-
An ancestral molecular response to nanomaterial particulates
Nature Nanotechnology (2023)
-
CTCF controls three-dimensional enhancer network underlying the inflammatory response of bone marrow-derived dendritic cells
Nature Communications (2023)