Abstract
Metastasis is the primary cause of cancer-related deaths, but the natural history, clonal evolution and impact of treatment are poorly understood. We analyzed whole-exome sequencing (WES) data from 457 paired primary tumor and metastatic samples from 136 patients with breast, colorectal and lung cancer, including untreated (n = 99) and treated (n = 100) metastases. Treated metastases often harbored private ‘driver’ mutations, whereas untreated metastases did not, suggesting that treatment promotes clonal evolution. Polyclonal seeding was common in untreated lymph node metastases (n = 17 out of 29, 59%) and distant metastases (n = 20 out of 70, 29%), but less frequent in treated distant metastases (n = 9 out of 94, 10%). The low number of metastasis-private clonal mutations is consistent with early metastatic seeding, which we estimated occurred 2–4 years before diagnosis across these cancers. Furthermore, these data suggest that the natural course of metastasis is selectively relaxed relative to early tumorigenesis and that metastasis-private mutations are not drivers of cancer spread but instead associated with drug resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The exome sequencing data for colorectal cancer patients sequenced in-house have been deposited at the European Genotype Phenotype Archive under accession number EGAS00001003573. The accession numbers for the public datasets are listed in Supplementary Table 1.
Code availability
Code used for genomic data analysis are available from https://github.com/cancersysbio/pan-metastasis
References
Talmadge, J. E., Wolman, S. R. & Fidler, I. J. Evidence for the clonal origin of spontaneous metastases. Science 217, 361–3 (1982).
Yamamoto, N. et al. Determination of clonality of metastasis by cell-specific color-coded fluorescent-protein imaging. Cancer Res. 63, 7785–7790 (2003).
Liu, W. et al. Copy number analysis indicates monoclonal origin of lethal metastatic prostate cancer. Nat. Med. 15, 559–565 (2009).
Huang, Y. et al. Multilayered molecular profiling supported the monoclonal origin of metastatic renal cell carcinoma. Int. J. Cancer 135, 78–87 (2014).
Gundem, G. et al. The evolutionary history of lethal metastatic prostate cancer. Nature 520, 353–357 (2015).
Maddipati, R. & Stanger, B. Z. Pancreatic cancer metastases harbor evidence of polyclonality. Cancer Discov. 5, 1086–1097 (2015).
Cheung, K. J. et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc. Natl Acad. Sci. USA 113, E854–E863 (2016).
Hunter, K. W., Amin, R., Deasy, S., Ha, N.-H. & Wakefield, L. Genetic insights into the morass of metastatic heterogeneity. Nat. Rev. Cancer 18, 211–223 (2018).
Klein, C. A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 9, 302–312 (2009).
Naxerova, K. & Jain, R. K. Using tumour phylogenetics to identify the roots of metastasis in humans. Nat. Rev. Clin. Oncol. 12, 258–272 (2015).
Robinson, D. R. et al. Integrative clinical genomics of metastatic cancer. Nature 548, 297–303 (2017).
Bertucci, F. et al. Genomic characterization of metastatic breast cancers. Nature 569, 560–564 (2019).
Priestley, P. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Zhao, Z.-M. et al. Early and multiple origins of metastatic lineages within primary tumors. Proc. Natl Acad. Sci. USA 113, 2140–2145 (2016).
Macintyre, G. et al. How subclonal modeling is changing the metastatic paradigm. Clin. Cancer Res. 23, 630–635 (2017).
Hu, Z. et al. Quantitative evidence for early metastatic seeding in colorectal cancer. Nat. Genet. 51, 1113–1122 (2019).
Leung, M. L. et al. Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res. 27, 1287–1299 (2017).
Turajlic, S. & Swanton, C. Metastasis as an evolutionary process. Science 352, 169–175 (2016).
Lee, S. Y. et al. Comparative genomic analysis of primary and synchronous metastatic colorectal cancers. PLoS ONE 9, e90459 (2014).
Kim, T.-M. et al. Subclonal genomic architectures of primary and metastatic colorectal cancer based on intratumoral genetic heterogeneity. Clin. Cancer Res. 21, 4461–4472 (2015).
Lim, B. et al. Genome-wide mutation profiles of colorectal tumors and associated liver metastases at the exome and transcriptome levels. Oncotarget 6, 22179–22190 (2015).
Uchi, R. et al. Integrated multiregional analysis proposing a new model of colorectal cancer evolution. PLoS Genet. 12, e1005778 (2016).
Brastianos, P. K. et al. Genomic characterization of brain metastases reveals branched evolution and potential therapeutic targets. Cancer Discov. 5, 1164–1177 (2015).
Um, S.-W. et al. Molecular evolution patterns in metastatic lymph nodes reflect the differential treatment response of advanced primary lung cancer. Cancer Res. 76, 6568–6576 (2016).
Chung, W. et al. Single-cell RNA-seq enables comprehensive tumour and immune cell profiling in primary breast cancer. Nat. Commun. 8, 15081 (2017).
Ng, C. K. Y. et al. Genetic heterogeneity in therapy-naïve synchronous primary breast cancers and their metastases. Clin. Cancer Res. 23, 4402–4415 (2017).
Razavi, P. et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 34, 427-438.e6 (2018).
Siegel, M. B. et al. Integrated RNA and DNA sequencing reveals early drivers of metastatic breast cancer. J. Clin. Invest. 128, 1371–1383 (2018).
Ullah, I. et al. Evolutionary history of metastatic breast cancer reveals minimal seeding from axillary lymph nodes. J. Clin. Invest. 128, 1355–1370 (2018).
Sun, R. et al. Between-region genetic divergence reflects the mode and tempo of tumor evolution. Nat. Genet. 49, 1015–1024 (2017).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 45, 1127–1133 (2013).
Rueda, O. M. et al. Dynamics of breast-cancer relapse reveal late-recurring ER-positive genomic subgroups. Nature 567, 399–404 (2019).
Christensen, S. et al. 5-Fluorouracil treatment induces characteristic T>G mutations in human cancer. Nat. Commun. 10, 4571 (2019).
Pich, O. et al. The mutational footprints of cancer therapies. Nat. Genet. 51, 1732–1740 (2019).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Rogers, M. F. et al. FATHMM-XF: accurate prediction of pathogenic point mutations via extended features. Bioinformatics 34, 511–513 (2018).
Tokheim, C. & Karchin, R. CHASMplus reveals the scope of somatic missense mutations driving human cancers. Cell Syst. 9, 9–23.e8 (2019).
Patel, S. A. & Vanharanta, S. Epigenetic determinants of metastasis. Mol. Oncol. 11, 79–96 (2017).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Makohon-Moore, A. P. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual patients with pancreatic cancer. Nat. Genet. 49, 358–366 (2017).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015).
Kang, H. et al. Many private mutations originate from the first few divisions of a human colorectal adenoma. J. Pathol. 237, 355–62 (2015).
Williams, M. J. et al. Quantification of subclonal selection in cancer from bulk sequencing data. Nat. Genet. 50, 895–903 (2018).
Caswell-Jin, J. L. et al. Clonal replacement and heterogeneity in breast tumors treated with neoadjuvant HER2-targeted therapy. Nat. Commun. 10, 657 (2019).
Williams, M. J., Werner, B., Barnes, C. P., Graham, T. A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Benzekry, S. et al. Classical mathematical models for description and prediction of experimental tumor growth. PLoS Comput. Biol. 10, e1003800 (2014).
Stein, R. G. et al. The impact of breast cancer biological subtyping on tumor size assessment by ultrasound and mammography—a retrospective multicenter cohort study of 6543 primary breast cancer patients. BMC Cancer 16, 459 (2016).
Cavalli, F., Kaye, S. B., Hansen, H. H., Armitage, J. O. & Piccart-Gebhart, M. J. Textbook of Medical Oncology 4th edn (Routledge, 2009).
Harper, K. L. et al. Mechanism of early dissemination and metastasis in Her2+ mammary cancer. Nature 540, 588–592 (2016).
Hosseini, H. et al. Early dissemination seeds metastasis in breast cancer. Nature 540, 552–558 (2016).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Ha, G. et al. TITAN: inference of copy number architectures in clonal cell populations from tumor whole-genome sequence data. Genome Res. 24, 1881–1893 (2014).
Ha, G. et al. Integrative analysis of genome-wide loss of heterozygosity and monoallelic expression at nucleotide resolution reveals disrupted pathways in triple-negative breast cancer. Genome Res. 22, 1995–2007 (2012).
Li, B. & Li, J. Z. A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data. Genome Biol. 15, 473 (2014).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Pan, H. et al. 20-year risks of breast-cancer recurrence after stopping endocrine therapy at 5 years. N. Engl. J. Med. 377, 1836–1846 (2017).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 174, 1034–1035 (2018).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Díaz-Gay, M. et al. Mutational Signatures in Cancer (MuSiCa): a web application to implement mutational signatures analysis in cancer samples. BMC Bioinformatics 19, 224 (2018).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–21 (2013).
Felsenstein, J. PHYLIP-phylogeny inference package (ver.3.2). Cladistics 5, 164–166 (1989).
Bozic, I. et al. Accumulation of driver and passenger mutations during tumor progression. Proc. Natl Acad. Sci. USA 107, 18545–18550 (2010).
Acknowledgements
We thank H. Xu, K. McNamara, E. Kotler, J. Caswell-Jin and other members of Curtis laboratory for valuable discussions. We thank J. Wang and Q. Mu for providing the scripts for the ternary plot. C.C. is supported by the National Institutes of Health (NIH) through the NIH Director’s Pioneer Award (DP1-CA238296), the American Association for Cancer Research–Triple Negative Breast Cancer Foundation–Carol’s Crusade for Cure Foundation Career Development Award for Metastatic Triple Negative Breast Cancer Research, grant no. 16-20-43-CURT) and the Emerson Collective. Z.H. is supported by an Innovative Genomics Initiative Postdoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
Z.H. and C.C. conceived and designed the study. Z.H. performed all computational analyses. Z.L. reviewed the published studies, extracted and analyzed the clinical data. Z.M. processed the in-house clinical samples and generated the genomic data. Z.H. and C.C. wrote the manuscript, which was reviewed by all authors.
Corresponding author
Ethics declarations
Competing interests
C.C. is a scientific advisor to GRAIL and reports stock options, as well as consulting for GRAIL and Genentech. Z.H., Z.L. and Z.M. have no conflicts of interest to report.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sankey diagram of patient cohorts with paired primary tumors and metastases.
In total, 136 primary tumors and 199 matched metastases from colorectal, lung and breast cancers were included. Treatment status is indicated.
Extended Data Fig. 2 Concordance of mutation burden in paired primary tumors (P) and metastasis (M).
Concordance amongst a, Clonal sSNVs; b, Subclonal SSNVs and c, sCNAs are indicated. Spearman’s correlation (ρ) is reported. Line indicates the linear regression and gray shading indicates the 95% confidence interval (CI) of the regression. The mean mutation burden across samples is reported for samples with multi-region sequencing data.
Extended Data Fig. 3 The ratio of nonsynonymous to synonymous mutations, dN/dS.
The dN/dS ratios of missense mutations (left panel) or nonsense mutations (right panel) relative to synonymous mutations are shown (on log2 scale). The dN/dS ratios for putative driver genes and passengers were computed separately. The driver gene list was obtained by merging TCGA pan-cancer drivers and COSMIC Cancer Gene Census (Methods). Circles and vertical lines correspond to the mean and 95% CI of the dN/dS ratio, respectively.
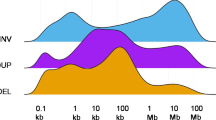
Extended Data Fig. 4 The frequency of somatic copy number alterations (sCNAs) for primary tumors and metastases across three cancer types.
The frequency of amplifications or deletions across 1 Mb genomic bins is shown for primary tumors and metastases.
Extended Data Fig. 5 The frequency of somatic copy number alterations (sCNAs) in putative driver genes in paired primary tumors (P) and metastases (M).
Left panel, amplifications (AMP) where oncogenes with an increased frequency (\(\ge\)15%) in the metastasis (M) versus primary (P) are labeled. Right panel, deletions (DEL) where tumor suppressor genes with increased frequency (\(\ge\)15%) in the metastasis (M) versus primary (P) are labeled.
Extended Data Fig. 6 Schematic illustration of a 3-D spatial-agent based model of tumor growth and metastasis.
Tumor growth is simulated via the expansion of deme subpopulations (mimicking the glandular structures often found in epithelial tumors and metastases) within a defined 3-D cubic lattice according to explicit rules dictated by spatial constraints, where cells within each deme are well-mixed and grow via a stochastic branching (birth-death) process (Methods). To model monoclonal seeding, a single cell at the tumor periphery was randomly sampled as the metastasis founder cell. To model polyclonal seeding, a cluster of cells (n = 10) was randomly sampled from the whole tumor in order to maximize the clonal diversity within the metastasis founder cells. Metastatic growth follows the same spatial-constraints as the primary and starts from the metastasis founder cell or cell cluster. The final sizes of both the primary tumor and metastasis is ~109 cells (~2×105 demes). Clonal selection is modeled by assuming a constant beneficial mutation rate that alters the cell birth/death probability according to the selection coefficient (denoted s). By simulating the acquisition of random mutations (neutral or beneficial), tracing the mutational genealogy of each cell as the tumor expands and subsequently spatially sampling (~106 cells in each sample) and sequencing the ‘final’ virtual tumor as is done experimentally after resection or biopsy, we obtain the variant allele frequencies (VAF) and cancer cell fraction (CCF) in both primary tumor and metastasis.
Extended Data Fig. 7 Lm, Lp and Ws values in tumors simulated under monoclonal versus polyclonal seeding.
The number of sSNVs in each of the three categories (M-private clonal or Lm, P-private clonal or Lp, P/M shared subclonal or Ws) in the simulated data generated by modeling monoclonal seeding or polyclonal seeding within an agent-based model (Methods) where one sample (~106 cells) was biopsied from each primary tumor and metastasis. We employed a mutation rate u = 0.6 per cell division in exonic regions (corresponding to 10-8 per site per cell division in the 60 Mb diploid coding regions). In order to account for varying scenarios of tumor growth dynamics, selection and timing of metastatic dissemination, the birth probability b of founding cells, selection coefficient s and primary tumor size at dissemination Nd was randomly sampled from a uniform distribution, b~U(0.55, 0.65), log10(s)~U(-3,-1) and log10(Nd)~U(4,8), respectively. A total of n = 500 virtual P/M pairs were simulated under monoclonal seeding and polyclonal seeding by randomly sampling these three parameters. Bar, median; box, 25th to 75th percentile (interquartile range, IQR); vertical line, data within 1.5 times the IQR.
Extended Data Fig. 8 Jaccard similarity index (JSI) values in lymph node and distant metastases and the percentage of polyclonal seeding across metastatic sites.
a, Lymph node metastases (LNM; n = 35) showed significantly higher JSI than distant metastases (n = 164). Among distant metastases, untreated metastasis showed higher JSI than treated metastasis although this was not statistically significant. However, using a cutoff of JSI = 0.3 to classify polyclonal (JSI\(\ge\)0.3) versus monoclonal seeding (JSI < 0.3), untreated distant metastases showed a significantly higher percentage of polyclonal seeding than treated distant metastases (Fig. 2e). P-value, Wilcoxon Rank-Sum Test (two-sided). Bar, median; box, 25th to 75th percentile (interquartile range, IQR); vertical line, data within 1.5 times the IQR. b, The percentage of polyclonal seeding among all LNM (lymph node metastasis), LiM (liver metastasis), BM (brain metastasis) and LuM (lung metastasis) (left panel) and stratified by treatment (right panel). P-value, Fisher’s exact test (two sided).
Extended Data Fig. 9 A mathematical method to quantify the chronology of metastatic seeding, ts.
a, Schematic of the parameters used to quantify metastatic timing ts (number of years prior to primary tumor diagnosis). We assume metastatic spread occurs at tm following the emergence of malignant founder of primary carcinoma (denoted t = 0). Let T be the time from emergence of malignant founder to diagnosis of the primary tumor, thus ts = T-tm. Let Lp and Lm be the number of private clonal sSNVs in a bulk sample from primary tumor and metastasis, respectively. Lp represents the number of sSNVs that occurred from emergence of the primary tumor founder to the most recent common ancestor (PMRCA) of cell lineages in a bulk sample. This time span is denoted as tp. Similarly, Lm denotes the number of SSNVs occurred from the emergence of primary tumor founder to the MRCA in a bulk sample from the metastasis (denoted MMRCA). Lm includes the number of M-private clonal mutations that occur: (i) within the primary tumor (Lm1) and (ii) after cells have disseminated from the primary tumor (Lm2), thus Lm = Lm1 + Lm2. b, Estimation of α by simulating an agent-based model of tumor evolution (Methods). The mean α and standard deviation from 1000 simulated tumors are shown.
Extended Data Fig. 10 Later metastatic seeding is associated with higher genomic divergence in matched primary tumors.
a, The number of primary (P)-private clonal sSNVs and metastasis (M)-private clonal sSNVs in synchronous (distant and monoclonal, n = 41) and metachronous (distant and monoclonal, n = 80) metastases, respectively. b, The number of P-to-M altered sCNAs in synchronous and metachronous metastases, respectively; P-values, two-sided Wilcoxon Rank-Sum Test. Bar, median; box, 25th to 75th percentile (interquartile range, IQR); vertical line, data within 1.5 times the IQR.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13, Note, and Tables 8 and 9
Supplementary Tables
Supplementary Table 1. Whole-exome sequencing (WES) datasets in paired primary tumors and metastases in three cancer types. Supplementary Table 2. Patient information of metastatic cancers in three cancer types (n = 136). Supplementary Table 3. Sample information of metastatic cancers in three cancer types (n = 457). Supplementary Table 4. Mutational counts in 199 primary tumor/metastasis pairs in three cancer types. Supplementary Table 5. Candidate driver genes from TCGA and COSMIC in colorectal, lung and breast cancer, respectively. Supplementary Table 6. Functional driver sSNVs/indels in the three cancer types. Supplementary Table 7. Enriched GO terms for M-private clonal (n = 152) or subclonal (135) functional sSNVs/indels.
Rights and permissions
About this article
Cite this article
Hu, Z., Li, Z., Ma, Z. et al. Multi-cancer analysis of clonality and the timing of systemic spread in paired primary tumors and metastases. Nat Genet 52, 701–708 (2020). https://doi.org/10.1038/s41588-020-0628-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0628-z
This article is cited by
-
Prediction of tumor-reactive T cell receptors from scRNA-seq data for personalized T cell therapy
Nature Biotechnology (2024)
-
How much do we know about the metastatic process?
Clinical & Experimental Metastasis (2024)
-
The evolution of metastatic upper tract urothelial carcinoma through genomic-transcriptomic and single-cell protein markers analysis
Nature Communications (2024)
-
State-dependent evolutionary models reveal modes of solid tumour growth
Nature Ecology & Evolution (2023)
-
Spatial biology of cancer evolution
Nature Reviews Genetics (2023)