Abstract
Active enhancers are frequently transcribed, yet the regulatory role of enhancer transcription remains debated. Here, we depleted the RNA polymerase II pausing and elongation factor Spt5 in activated mouse B cells and found that approximately 50% of enhancer–gene pairs showed co-regulated transcription, consistent with a potential functional requirement for enhancer transcription. In particular, Spt5 depletion led to loss of super-enhancer–promoter physical interaction and gene expression at the immunoglobulin heavy-chain locus (Igh), abrogating antibody class switch recombination. This defect correlated strictly with loss of enhancer transcription but did not affect acetylation of histone H3 at lysine 27, chromatin accessibility and occupancy of Mediator and cohesin at the enhancer. Strikingly, CRISPRa-mediated rescue of enhancer transcription in Spt5-depleted cells restored Igh gene expression. Our work suggests that Spt5-mediated enhancer transcription underlies the physical and functional interaction between a subset of active enhancers and their target promoters.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Reiter, F., Wienerroither, S. & Stark, A. Combinatorial function of transcription factors and cofactors. Curr. Opin. Genet. Dev. 43, 73–81 (2017).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Core, L. J. et al. Defining the status of RNA polymerase at promoters. Cell Rep. 2, 1025–1035 (2012).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Core, L. J. et al. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 46, 1311–1320 (2014).
Henriques, T. et al. Widespread transcriptional pausing and elongation control at enhancers. Genes Dev. 32, 26–41 (2018).
Lai, F. et al. Activating RNAs associate with mediator to enhance chromatin architecture and transcription. Nature 494, 497–501 (2013).
Schaukowitch, K. et al. Enhancer RNA facilitates NELF release from immediate early genes. Mol. Cell 56, 29–42 (2014).
Bose, D. A. et al. RNA binding to CBP stimulates histone acetylation and transcription. Cell 168, 135–149.e22 (2017).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Mikhaylichenko, O. et al. The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription. Genes Dev. 32, 42–57 (2018).
Young, R. S., Kumar, Y., Bickmore, W. A. & Taylor, M. S. Bidirectional transcription initiation marks accessible chromatin and is not specific to enhancers. Genome Biol. 18, 242 (2017).
Paralkar, V. R. et al. Unlinking an lncRNA from its associated cis element. Mol. Cell 62, 104–110 (2016).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Lloret-Llinares, M. et al. The RNA exosome contributes to gene expression regulation during stem cell differentiation. Nucleic Acids Res. 46, 11502–11513 (2018).
Gu, B. et al. Transcription-coupled changes in nuclear mobility of mammalian cis-regulatory elements. Science 359, 1050–1055 (2018).
Chen, H. et al. Dynamic interplay between enhancer–promoter topology and gene activity. Nat. Genet. 50, 1296–1303 (2018).
Alexander, J. M. et al. Live-cell imaging reveals enhancer-dependent Sox2 transcription in the absence of enhancer proximity. eLife 8, e41769 (2019).
Benabdallah, N. S. et al. Decreased enhancer–promoter proximity accompanying enhancer activation. Mol. Cell 76, 473–484.e7 (2019).
Yamaguchi, Y. et al. Structure and function of the human transcription elongation factor DSIF. J. Biol. Chem. 274, 8085–8092 (1999).
Ivanov, D., Kwak, Y. T., Guo, J. & Gaynor, R. B. Domains in the SPT5 protein that modulate its transcriptional regulatory properties. Mol. Cell. Biol. 20, 2970–2983 (2000).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Fitz, J., Neumann, T. & Pavri, R. Regulation of RNA polymerase II processivity by Spt5 is restricted to a narrow window during elongation. EMBO J. 37, e97965 (2018).
Kieffer-Kwon, K. R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Gressel, S. et al. CDK9-dependent RNA polymerase II pausing controls transcription initiation. eLife 6, e29736 (2017).
Shao, W. & Zeitlinger, J. Paused RNA polymerase II inhibits new transcriptional initiation. Nat. Genet. 49, 1045–1051 (2017).
Pinaud, E. et al. The IgH locus 3′ regulatory region: pulling the strings from behind. Adv. Immunol. 110, 27–70 (2011).
Vincent-Fabert, C. et al. Genomic deletion of the whole IgH 3′ regulatory region (hs3a, hs1,2, hs3b, and hs4) dramatically affects class switch recombination and Ig secretion to all isotypes. Blood 116, 1895–1898 (2010).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Chaudhuri, J. & Alt, F. W. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat. Rev. Immunol. 4, 541–552 (2004).
Stavnezer, J. & Schrader, C. E. IgH chain class switch recombination: mechanism and regulation. J. Immunol. 193, 5370–5378 (2014).
Wuerffel, R. et al. S–S synapsis during class switch recombination is promoted by distantly located transcriptional elements and activation-induced deaminase. Immunity 27, 711–722 (2007).
Manis, J. P. et al. Class switching in B cells lacking 3′ immunoglobulin heavy chain enhancers. J. Exp. Med. 188, 1421–1431 (1998).
Manis, J. P., Michaelson, J. S., Birshtein, B. K. & Alt, F. W. Elucidation of a downstream boundary of the 3′ IgH regulatory region. Mol. Immunol. 39, 753–760 (2003).
Bebin, A. G. et al. In vivo redundant function of the 3′ IgH regulatory element HS3b in the mouse. J. Immunol. 184, 3710–3717 (2010).
Pinaud, E. et al. Localization of the 3′ IgH locus elements that effect long-distance regulation of class switch recombination. Immunity 15, 187–199 (2001).
Cogne, M. et al. A class switch control region at the 3′ end of the immunoglobulin heavy chain locus. Cell 77, 737–747 (1994).
Saintamand, A. et al. Elucidation of IgH 3′ region regulatory role during class switch recombination via germline deletion. Nat. Commun. 6, 7084 (2015).
Pavri, R. et al. Activation-induced cytidine deaminase targets DNA at sites of RNA polymerase II stalling by interaction with Spt5. Cell 143, 122–133 (2010).
Thomas-Claudepierre, A. S. et al. Mediator facilitates transcriptional activation and dynamic long-range contacts at the IgH locus during class switch recombination. J. Exp. Med. 213, 303–312 (2016).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Thomas-Claudepierre, A. S. et al. The cohesin complex regulates immunoglobulin class switch recombination. J. Exp. Med. 210, 2495–2502 (2013).
Gunal-Sadik, G. et al. Stage-specific binding profiles of cohesin in resting and activated B lymphocytes suggest a role for cohesin in immunoglobulin class switching and maturation. PLoS ONE 9, e111748 (2014).
Seitan, V. C. et al. A role for cohesin in T-cell-receptor rearrangement and thymocyte differentiation. Nature 476, 467–471 (2011).
Merkenschlager, M. & Odom, D. T. CTCF and cohesin: linking gene regulatory elements with their targets. Cell 152, 1285–1297 (2013).
Trimarchi, T. et al. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell 158, 593–606 (2014).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
He, M., Cortizas, E. M., Verdun, R. E. & Severinson, E. Cyclin-dependent kinases regulate Ig class switching by controlling access of AID to the switch region. J. Immunol. 194, 4231–4239 (2015).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Titov, D. V. et al. XPB, a subunit of TFIIH, is a target of the natural product triptolide. Nat. Chem. Biol. 7, 182–188 (2011).
Chao, S. H. & Price, D. H. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J. Biol. Chem. 276, 31793–31799 (2001).
Jonkers, I., Kwak, H. & Lis, J. T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Wang, Y., Lu, J. J., He, L. & Yu, Q. Triptolide (TPL) inhibits global transcription by inducing proteasome-dependent degradation of RNA polymerase II (Pol II). PLoS ONE 6, e23993 (2011).
Steurer, B. et al. Live-cell analysis of endogenous GFP-RPB1 uncovers rapid turnover of initiating and promoter-paused RNA Polymerase II. Proc. Natl Acad. Sci. USA 115, E4368–E4376 (2018).
Wada, T. et al. DSIF, a novel transcription elongation factor that regulates RNA polymerase II processivity, is composed of human Spt4 and Spt5 homologs. Genes Dev. 12, 343–356 (1998).
Nair, S. J. et al. Phase separation of ligand-activated enhancers licenses cooperative chromosomal enhancer assembly. Nat. Struct. Mol. Biol. 26, 193–203 (2019).
Melo, C. A. et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 49, 524–535 (2013).
Mousavi, K. et al. eRNAs promote transcription by establishing chromatin accessibility at defined genomic loci. Mol. Cell 51, 606–617 (2013).
Kaikkonen, M. U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Wang, D. et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474, 390–394 (2011).
Rennie, S. et al. Transcription start site analysis reveals widespread divergent transcription in D. melanogaster and core promoter-encoded enhancer activities. Nucleic Acids Res. 46, 5455–5469 (2018).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178.e20 (2018).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Braikia, F. Z. et al. Inducible CTCF insulator delays the IgH 3′ regulatory region-mediated activation of germline promoters and alters class switching. Proc. Natl Acad. Sci. USA 114, 6092–6097 (2017).
Marina-Zarate, E., Perez-Garcia, A. & Ramiro, A. R. CCCTC-binding factor locks premature IgH germline transcription and restrains class switch recombination. Front. Immunol. 8, 1076 (2017).
Pefanis, E. et al. RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity. Cell 161, 774–789 (2015).
Lim, J. et al. Nuclear proximity of Mtr4 to RNA exosome restricts DNA mutational asymmetry. Cell 169, 523–537.e15 (2017).
Germier, T. et al. Real-time imaging of a single gene reveals transcription-initiated local confinement. Biophys. J. 113, 1383–1394 (2017).
McBride, K. M. et al. Regulation of hypermutation by activation-induced cytidine deaminase phosphorylation. Proc. Natl Acad. Sci. USA 103, 8798–8803 (2006).
Core, L. J. & Lis, J. T. Transcription regulation through promoter-proximal pausing of RNA polymerase II. Science 319, 1791–1792 (2008).
Schoeberl, U. E., Kurth, H. M., Noto, T. & Mochizuki, K. Biased transcription and selective degradation of small RNAs shape the pattern of DNA elimination in Tetrahymena. Genes Dev. 26, 1729–1742 (2012).
Mahat, D. B. et al. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-Seq). Nat. Protoc. 11, 1455–1476 (2016).
Kwak, H., Fuda, N. J., Core, L. J. & Lis, J. T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Feldman, S. et al. Constraints contributed by chromatin looping limit recombination targeting during Ig class switch recombination. J. Immunol. 194, 2380–2389 (2015).
Baek, S., Sung, M. H. & Hager, G. L. Quantitative analysis of genome-wide chromatin remodeling. Methods Mol. Biol. 833, 433–441 (2012).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Acknowledgements
We are grateful to the Vienna Biocenter Core Facilities for the next-generation sequencing, and to the IMP/IMBA core facilities, especially Comparitive Medicine, BioOptics, Molecular Biology Service and Protein Chemistry. We thank K. Uzunova for the Tn5 enzyme preparation, K. Mochizuki for the GRO-Seq reagents, and C. Umkehrer and A. Obenauf for the dual sgRNA vector. We thank A. Stark, M. Busslinger, C. Bücker and A. Andersen (Life Science Editors) for critical reading of the manuscript. This work was funded by Boehringer Ingelheim and the Austrian Research Promotion Agency (headquarter grant FFG-834223). U.E.S. is supported by the L’Oréal For Women in Science Austria fellowship (2015) and Austrian Science Fund (FWF T 795-B30).
Author information
Authors and Affiliations
Contributions
J.F. designed and performed the experiments and co-wrote the manuscript. T.N. performed all of the bioinformatic analyses. M.S., E.-M.W., A.C.G., A.A. and U.E.S. designed and performed the experiments. R.P. conceived of the project, designed and performed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Targeting strategy for generating Spt5-depleted (Spt5dep) mice and characterization of Spt5dep primary B cells.
a, Scheme for obtaining Spt5dep B cells (described in detail in Fitz et al. 23). Briefly, the targeting vector (Sanger Consortium) was used to generate knock-in mice, which were bred to Flp-expressing followed by EIIA-Cre-expressing mice to generate Supt5hF/+ and Supt5h-/+ genotypes, respectively. Supt5hF/F mice were crossed with Supt5h-/+Rosa26 Cre-ERT2/Cre-ERT2 mice to obtain the final genotype, Supt5hF/-Rosa26Cre-ERT2/+. b, Generation of WT (Rosa26Cre-ERT2/+) and Spt5dep (Supt5h-/-Rosa26Cre-ERT2/+) B cells. Primary B cells were isolated from spleens and cultured for 60 h with IL4 and LPS. 32 h prior to harvesting, 2 μM 4-hydroxytamoxifen (4-HT) was added to induce deletion of the floxed Supt5h exons. c, Multiplex PCR-based genotyping from genomic DNA from B cells of three Spt5dep mice at indicated times after 4-HT addition. d, Spt5 protein analysis from 10 μg nuclear extracts of WT and Spt5dep primary B cells (two mice each) treated with 4-HT for 0 h, 24 h and 32 h. Relative quantification was performed by normalization to histone H3 by setting the WT condition of each timepoint to 100%. e, Assessment of cell proliferation. Primary WT and Spt5dep B cells were stained with Cell Trace Violet upon isolation and analyzed by flow cytometry 32 h post 4-HT treatment. f, Viability curve using Trypan Blue exclusion for WT and Spt5dep primary B cells treated with 4-HT at the indicated time-points. The data represent the mean ± s.d. from eight replicates per genotype. g, Analysis of Pol II, Spt5 and histone H3 protein levels from nuclear extracts of WT and Spt5dep primary B cells treated with 4-HT for 32 h. Two replicate mice were used per genotype and a two-fold titration (3 μg and 10 μg) of nuclear extract was performed.
Extended Data Fig. 2 Replicate correlation analysis for all next-generation sequencing data used in this study.
a, Replicate correlation plots for total Pol II, Ser5P-Pol II, Ser2P-Pol II and Spt5 ChIP-seq datasets, as well as for GRO-seq, ATAC-seq and PRO-cap datasets from WT and Spt5dep primary B cells. Spearmann’s correlation coefficient (r) is indicated within the plots. b, Replicate correlation plots for mRNA-seq done in triplicate with Spearmann’s correlation coefficient indicated.
Extended Data Fig. 3 Identification of differentially expressed enhancer (DEE) groups.
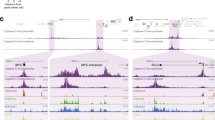
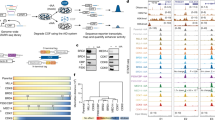
a, Heatmaps in WT activated B cells centered on extragenic DNase hypersensitive site (DHS) peaks and ordered by decreasing DHS read density. Read densities of the following datasets are shown as reads per million (RPM): DHS-seq, ChIP-seq of total Pol II, serine 5-phosphorylated Pol II (Ser5P-Pol II), serine 2-phosphorylated Pol II (Ser2P-Pol II) and Spt5 and GRO-seq (sense and anti-sense strand reads are separated). DHS-seq data is from Kieffer-Kwon et al., 2013. b, Generating differentially expressed enhancer (DEE) groups. GRO-seq fold-changes (Spt5dep/WT) at extragenic DHSs (considered as putative enhancers) were clustered into three DEE groups: up-regulated, moderately down-regulated and strongly down-regulated (shown in Extended Data Fig. 5a). DEEs were linked to genes using Pol II ChIA-PET (Kieffer-Kwon et al., 2013). DEEs with (linked) or without (unlinked) PET links were divided into H3K27ac-positive (H3K27ac-pos) enhancers, H3K27ac-negative (H3K27ac-neg) enhancers and super-enhancer (SE) classes based on histone H3 lysine 27 acetylation (H3K27ac) (Whyte et al., 79). Finally, these enhancer classes were assigned to differentially expressed genes (DEGs) based on direct PET linkages (as in Extended Data Fig. 5c, d). A detailed explanation of this workflow is provided in the Methods under the Bioinformatics section. c, Upper panel: Diagram showing the calculation of the 5’ pausing index (5’PI). Note that we do not employ the conventional PI where the whole gene body is considered because of the processivity defects within gene bodies in Spt5dep cells seen in Extended Data Fig. 5a and described previously in Spt5dep MEFs (Fitz et al., 23). Lower panel: 5’ PI of DEGs assigned to DEEs. Each bar represents DEGs linked to the indicated DEE group. Each linked DEE group is split into the three enhancer classes as described in b. Statistical significance is calculated using the Student’s t test. Asterisks indicate P < 0.05 and ns = not significant.
Extended Data Fig. 4 Spt5 depletion results in elongation defects at long genes but does not lead to a global down-regulation of mRNA synthesis.
a, GRO-seq Spt5dep /WT ratio heatmap of all expressed minus-strand genes <100 kb in length (n=2943) generated using 1025 static bins and ordered by increasing gene length. Read densities are in reads per million (RPM). The transcription start site (TSS), and transcription termination site (TTS) are indicated. b, UCSC browser snapshots at one short and one long gene in WT and Spt5dep B cells. The tracks are overlaid with blue and red representing the WT and Spt5dep samples, respectively, and the overlapping region in black. The Y axis shows normalized read counts. c, UCSC browser snapshots of a representative up-regulated DEG (top) and down-regulated DEG (bottom). The WT and Spt5dep tracks are overlaid as described in b. d, Volcano plot of mRNA-seq data from WT and Spt5dep primary B cells performed in triplicate and calculated with the DEseq2 software. Genes having an expression fold-change >2 and an adjusted P value of < 0.1 are highlighted in red and considered as up- or down-regulated genes in this study. e, Box plots showing the distribution of mRNA fold-changes (Spt5dep/WT) within the corresponding GRO-seq-based DEG classes shown in Extended Data Fig. 5c. f, RT-qPCR analysis for mRNA abundance in WT and Spt5dep cells of five up-regulated and five down-regulated genes. Drosophila S2 cells were spiked in to normalize WT and Spt5dep qPCR values to the transcripts of the Drosophila-specific housekeeping gene, Act5c. The plotted data represents the mean of eight independent WT and Spt5dep samples ± s.d. Statistical significance in e and f were calculated using the two-tailed Student’s t test. Asterisks indicate P < 0.01.
Extended Data Fig. 5 A subset of enhancer-promoter pairs shows correlated transcriptional changes upon Spt5 depletion.
a, Left panel: WT GRO-seq read densities within the DEE groups (Extended Data Fig. 3b). DEEs linked to gene promoters using Pol II ChIA-PET, as well as DEEs with no PET links (unlinked DEEs), were further divided into three enhancer classes based on H3K27ac signal. DEEs with no PET links were termed unlinked enhancers. The number of DHSs in each row is indicated below the plot. Right panel: Same as before but now showing the fold-change in GRO-seq signal upon Spt5 depletion (Spt5dep/WT). Statistical significance for both plots is calculated using the two-tailed Student’s t test. Asterisks indicate P < 0.05 and ns = not significant. b, Mediator subunit, Med12, and histone H3 lysine 4 monomethylation (H3K4me1) levels within the DEE groups displayed exactly as in a. c, Left panel: Whole gene GRO-seq composite metagene analysis of WT and Spt5dep tracks representing the three differentially expressed gene (DEG) classes (Methods). Right panel: A magnified view of the gene body region (boxed in the left panel plots). d, Each bar represents DEGs linked to the indicated DEE group shown in a. Each linked DEE group is split into the three enhancer classes as described in b. The Y axis is set to 100% and the number of DEGs per bar is indicated above the bar. The numbers in white within the bars indicate the percentage of that DEG class relative to the total DEGs in that bar. Note the relatively equal distribution of DEGs showing up-, down- or intermediate regulation within each DEE group irrespective of linkage to an H3K27ac-pos enhancer, H3K27ac-neg enhancer or SE.
Extended Data Fig. 6 Effect of Spt5 depletion on B cell activation, expression of major B cells transcription factors and IgH GLTs under various stimulation conditions.
a-b, Flow cytometry analysis of a panel of B cell activation markers in WT and Spt5dep primary B cells after 32h of 4-HT treatment. Representative histogram plots for six activation markers is shown in (A) and bar graphs of mean fluorescence intensity for all tested activation markers is shown in b. Data in b represent the mean of eight independent WT and Spt5dep samples ± s.d. No statistically significant differences were detected between WT and Spt5dep activated B cells. c, RT-qPCR analysis for Igh germline transcript abundance in WT and Spt5dep cells activated with different stimulation conditions, as mentioned above each graph. For each condition, the expressed and expected Igh transcripts were analyzed, as indicated. The data represents the mean of three independent WT and Spt5dep samples ± s.d. Statistical significance was calculated using the two tailed Student’s t test (asterisks indicate P < 0.01 and ns = not significant). d, Scatter plot of mRNA-seq data for selected B cell transcription factors and cofactors as well as general transcription factors, Pol II, Mediator and cohesin (see Supplementary Table 1 for a complete list). The data plotted represent the mean reads per kilobase per million mapped reads (RPKM) of three independent WT and Spt5dep primary B cells samples. Genes showing 2-fold significant differential expression are marked in green (up-regulated) and red (down-regulated).
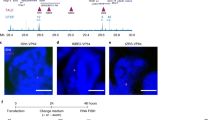
Extended Data Fig. 7 dCas9-VPR cannot stimulate Igh transcription in the absence of transcription.
a, Snapshot of the Igh locus showing the location of the uIghε sgRNA (~3 kb upstream of the Igh-ε promoter) relative to the IghH-γ1 and Igh-ε genes, and the 3’RR. b, Anti-Cas9 ChIP-qPCR at the hs4 and uIghε in cells infected with the indicated dCas9-VPR/sgRNA viruses, as described in Fig. 4a. c, RT-qPCR at hs4 and uIghε from the experiment in (B). For the uIghε, two sets of primers were used probing the region upstream and downstream of the sgRNA binding site. d, RT-qPCR for Igh-γ1 and Igh-μ transcripts in WT and Spt5dep cells infected with dCas9-VPR and uIghε sgRNA, as in Fig. 4a. In all cases, four independent experiments were performed and the two-tailed Student’s t test was used for statistical significance. Asterisks indicate P < 0.05 and ns, not significant.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, and Discussion
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Gating strategy for flow cytometry.
Rights and permissions
About this article
Cite this article
Fitz, J., Neumann, T., Steininger, M. et al. Spt5-mediated enhancer transcription directly couples enhancer activation with physical promoter interaction. Nat Genet 52, 505–515 (2020). https://doi.org/10.1038/s41588-020-0605-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0605-6
This article is cited by
-
The immunoglobulin heavy chain super enhancer controls class switch recombination in developing B cells
Scientific Reports (2024)
-
Concerted transformation of a hyper-paused transcription complex and its reinforcing protein
Nature Communications (2024)
-
Binding of small molecule inhibitors to RNA polymerase-Spt5 complex impacts RNA and DNA stability
Journal of Computer-Aided Molecular Design (2024)
-
RIF1 regulates early replication timing in murine B cells
Nature Communications (2023)
-
Using Synthetic DNA Libraries to Investigate Chromatin and Gene Regulation
Chromosoma (2023)