Abstract
Low-coverage whole-genome sequencing followed by imputation has been proposed as a cost-effective genotyping approach for disease and population genetics studies. However, its competitiveness against SNP arrays is undermined because current imputation methods are computationally expensive and unable to leverage large reference panels. Here, we describe a method, GLIMPSE, for phasing and imputation of low-coverage sequencing datasets from modern reference panels. We demonstrate its remarkable performance across different coverages and human populations. GLIMPSE achieves imputation of a genome for less than US$1 in computational cost, considerably outperforming other methods and improving imputation accuracy over the full allele frequency range. As a proof of concept, we show that 1× coverage enables effective gene expression association studies and outperforms dense SNP arrays in rare variant burden tests. Overall, this study illustrates the promising potential of low-coverage imputation and suggests a paradigm shift in the design of future genomic studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
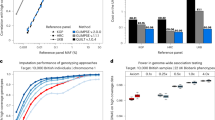
The 1000 Genomes Project phase 3 dataset sequenced at high coverage by the New York Genome Center is available on the European Nucleotide Archive under accession no. PRJEB31736. The publicly available subset of the HRC dataset is available from the European Genome-phenome Archive at the European Bioinformatics Institute (EBI) under accession no. EGAS00001001710. The Genome in A Bottle data for sample NA12878 is available at the National Center for Biotechnology Information ftp website: ftp://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data/NA12878. The subset of the 1000 Genomes samples genotyped on Affymetrix6.0 is available at ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/release/20130502/supporting/hd_genotype_chip/. GnomAD v.3 is available at https://gnomad.broadinstitute.org/downloads. The list of positions used to simulate the SNP arrays is available at https://www.well.ox.ac.uk/~wrayner/strand/. The RNA-seq data are part of the Geuvadis study and are available at the EBI ArrayExpress under accession code no. E-GEUV-1. The ENCODE project was accessed using accession nos. integration_data_jan2011 for the lymphoblastoid cell line-specific protein binding sites, ENCSR000EJD for the DNase-hypersensitive sites and ENCSR000AKC for locations with H3K27ac histone modifications. The results shown in Fig. 3a,b are a subset of the configurations tested. A full view of the results in available at the GLIMPSE website (European population: https://odelaneau.github.io/GLIMPSE/rsquare_eur.html, African-American population: https://odelaneau.github.io/GLIMPSE/rsquare_asw.html). The full raw data used to generate Fig. 3a,b and the benchmark shown on the website are available at the GLIMPSE repository (https://github.com/odelaneau/GLIMPSE/tree/master/docs/data/rsquare). Source data are provided with this paper.
Code availability
GLIMPSE is available from https://github.com/odelaneau/GLIMPSE and https://odelaneau.github.io/GLIMPSE/.
Change history
20 January 2021
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Brody, J. A. et al. Analysis commons, a team approach to discovery in a big-data environment for genetic epidemiology. Nat. Genet. 49, 1560–1563 (2017).
Alex Buerkle, C. & Gompert, Z. Population genomics based on low coverage sequencing: how low should we go? Mol. Ecol. 22, 3028–3035 (2013).
Le, S. Q. & Durbin, R. SNP detection and genotyping from low-coverage sequencing data on multiple diploid samples. Genome Res. 21, 952–960 (2011).
Pasaniuc, B. et al. Extremely low-coverage sequencing and imputation increases power for genome-wide association studies. Nat. Genet. 44, 631–635 (2012).
Cai, N. et al. Sparse whole-genome sequencing identifies two loci for major depressive disorder. Nature 523, 588–591 (2015).
Gilly, A. et al. Very low-depth sequencing in a founder population identifies a cardioprotective APOC3 signal missed by genome-wide imputation. Hum. Mol. Genet. 25, 2360–2365 (2016).
Gilly, A. et al. Very low-depth whole-genome sequencing in complex trait association studies. Bioinformatics 35, 2555–2561 (2019).
Browning, B. L. & Yu, Z. Simultaneous genotype calling and haplotype phasing improves genotype accuracy and reduces false-positive associations for genome-wide association studies. Am. J. Hum. Genet. 85, 847–861 (2009).
Spiliopoulou, A., Colombo, M., Orchard, P., Agakov, F. & McKeigue, P. GeneImp: fast imputation to large reference panels using genotype likelihoods from ultralow coverage sequencing. Genetics 206, 91–104 (2017).
Wasik, K. et al. Comparing low-pass sequencing and genotyping for trait mapping in pharmacogenetics. Preprint at bioRxiv https://doi.org/10.1101/632141 (2019).
Davies, R. W., Flint, J., Myers, S. & Mott, R. Rapid genotype imputation from sequence without reference panels. Nat. Genet. 48, 965–969 (2016).
Li, N. & Stephens, M. Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165, 2213–2233 (2003).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Delaneau, O., Zagury, J.-F., Robinson, M. R., Marchini, J. L. & Dermitzakis, E. T. Accurate, scalable and integrative haplotype estimation. Nat. Commun. 10, 5436 (2019).
Durbin, R. Efficient haplotype matching and storage using the positional Burrows–Wheeler transform (PBWT). Bioinformatics 30, 1266–1272 (2014).
Rubinacci, S., Delaneau, O. & Marchini, J. Genotype imputation using the Positional Burrows Wheeler Transform. PLoS Genet. 16, e1009049 (2020).
Browning, B. L. & Browning, S. R. Genotype imputation with millions of reference samples. Am. J. Hum. Genet. 98, 116–126 (2016).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Pritchard, J. K. & Przeworski, M. Linkage disequilibrium in humans: models and data. Am. J. Hum. Genet. 69, 1–14 (2001).
Zook, J. M. et al. Integrating human sequence data sets provides a resource of benchmark SNP and indel genotype calls. Nat. Biotechnol. 32, 246–251 (2014).
Huang, J. et al. Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat. Commun. 6, 8111 (2015).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Delaneau, O. et al. Chromatin three-dimensional interactions mediate genetic effects on gene expression. Science 364, eaat8266 (2019).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Brown, A. A. et al. Predicting causal variants affecting expression by using whole-genome sequencing and RNA-seq from multiple human tissues. Nat. Genet. 49, 1747–1751 (2017).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Preprint at bioRxiv https://doi.org/10.1101/563866 (2019).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Delaneau, O. et al. A complete tool set for molecular QTL discovery and analysis. Nat. Commun. 8, 15452 (2017).
Dale, R. K., Pedersen, B. S. & Quinlan, A. R. Pybedtools: a flexible Python library for manipulating genomic datasets and annotations. Bioinformatics 27, 3423–3424 (2011).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Acknowledgements
This work was funded by a Swiss National Science Foundation project grant no. PP00P3_176977. The New York Genome Center 1000 Genomes data were generated at the New York Genome Center with funds provided by a National Human Genome Research Institute grant no. 3UM1HG008901–03S1. We thank S. Carmi for useful comments on the preprint version of the manuscript.
Author information
Authors and Affiliations
Contributions
S.R., D.M.R. and O.D. designed the study, performed the experiments and drafted the paper. S.R. and O.D. developed the algorithm and wrote the software. S.R., R.J.H. and O.D. created the website. O.D. supervised the project. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Garrett Hellenthal, Sam Morris and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Read count distribution of downsampled sequencing data.
The y-axis shows the fractions of genotypes covered by 0 to 11 sequencing reads across multiple downsampled coverages from 0.1x to 4.0x. The color bars show the observed fractions in the downsampled data while the black dots and lines show the expected fractions assuming coverage is Poisson distributed.
Extended Data Fig. 2 Phasing performance of subsets of EUR and ASW samples.
Performance of the GLIMPSE (blue line) and SHAPEIT4 (black line) phasing algorithms. SHAPEIT4 has been run to rephase the genotype calls produced by GLIMPSE as it can only handle hard called genotypes. Validation genotypes were generated using an Affymetrix 6.0 SNP array. Validation haplotypes were derived thanks to additional samples being genotyped allowing to form multiple duos and trios.
Extended Data Fig. 3 Genotype discordance.
Genotype discordance stratified by minor-allele-frequency for the 1x coverage European population dataset on chromosome 1. The reference panel used are a subset of 5,000 samples (solid lines), and 20,000 samples from the HRC (dashed lines). The genotype discordance is shown for (A) all genotypes, and split between (B) major/major, (C) major/minor or (D) minor/minor genotypes in the validation dataset.
Extended Data Fig. 4 Zoomed-in genotype discordance for MAF > 1%.
Genotype discordance stratified by minor-allele-frequency (MAF > 1%) for the 1x coverage European population dataset on chromosome 1. The reference panels used are a subset of 5,000 samples (solid lines), and 20,000 samples from the HRC (dashed lines). The genotype discordance is shown for (A) all genotypes, and split between (B) major/major, (C) major/minor or (D) minor/minor genotypes in the validation dataset.
Extended Data Fig. 5 Non-reference discordance.
Non-reference discordance (NRD) stratified by non-reference allele frequency for the 1x coverage European population dataset on chromosome 1. The reference panels used are a subset of 5,000 samples (solid lines), and 20,000 samples from the HRC (dashed lines). (A.) Non-reference allele frequency > 0.01%; (B.) Non-reference allele frequency > 1%. The NRD is calculated as \(\left( {e_{rr} + e_{ra} + e_{aa}} \right)/\left( {m_{ra} + m_{aa} + e_{rr} + e_{ra} + e_{aa}} \right)\), where err, era and eaa are the counts of the mismatches for the homozygous reference, heterozygous and homozygous alternative genotypes, while mra and maa are the counts of the matches at heterozygous and homozygous alternative genotypes.
Extended Data Fig. 6 Calibration of genotype posteriors for 1.0x coverage.
(A.) Calibration of genotype posterior probabilities of different imputation methods for 1.0x coverage European dataset on chromosome 1. The reference panels used are a subset of 5,000 samples (solid lines), and 20,000 samples from the HRC (dashed lines). Imputed genotypes are binned according to the posterior probability distribution (x-axis) and plotted against the percentage of concordance against high coverage data (y-axis). (B.) Number of genotypes per probability bin.
Extended Data Fig. 7 Running time of imputation methods.
Running time of low-coverage sequencing imputation methods for the European population chromosome 1 dataset. We only ran GENEIMP on 1x coverage data. For BEAGLE and GENEIMP we only show reference panel size up to 5,000 samples due to time limits. The vertical axis is on a log scale.
Extended Data Fig. 8 Memory usage of imputation methods.
Memory usage of low-coverage sequencing methods for the European population chromosome 1 dataset. We only ran GENEIMP on 1x coverage data. For BEAGLE and GENEIMP we only show reference panel size up to 5,000 samples due to time limits. LOIMPUTE imputes a single sample at the time, therefore the reported memory usage is for a single sample, while we report the memory usage for the full cohort of 503 individuals for all other methods. The vertical axis is on a log scale.
Extended Data Fig. 9 Lead eQTL overlap and association p-value mean absolute error.
(A) Overlap between lead eQTLs identified in high-coverage and each low-coverage and SNP array dataset. eQTL mapping was performed independently for each dataset (FDR 5%; MAF > = 1%). eGenes in which the lead eQTL p-value was tied with another variant’s p-value (for example due to perfect linkage disequilibrium) were excluded, as the choice of variant for being the lead eQTL in these cases is arbitrary. The total number genes assessed after filtering was 5037. (B) Mean absolute error between -log10 p-values of association obtained for high-coverage lead eQTLs and those obtained in each dataset for the same set of variants. All high coverage lead eQTLs (that is a variant for each of the 16894 genes) were considered here, regardless of significance level. The scatterplots detail the -log10 p-values used to calculate the mean absolute errors for several relevant low-coverages and SNP arrays.
Supplementary information
Supplementary Information
Supplementary Note, Figs. 1–15, and Tables 1 and 2
Source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Rights and permissions
About this article
Cite this article
Rubinacci, S., Ribeiro, D.M., Hofmeister, R.J. et al. Efficient phasing and imputation of low-coverage sequencing data using large reference panels. Nat Genet 53, 120–126 (2021). https://doi.org/10.1038/s41588-020-00756-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00756-0
This article is cited by
-
A cautionary tale of low-pass sequencing and imputation with respect to haplotype accuracy
Genetics Selection Evolution (2024)
-
The hazards of genotype imputation when mapping disease susceptibility variants
Genome Biology (2024)
-
Assessing the impact of post-mortem damage and contamination on imputation performance in ancient DNA
Scientific Reports (2024)
-
Accurate detection of identity-by-descent segments in human ancient DNA
Nature Genetics (2024)
-
A cost-effective sequencing method for genetic studies combining high-depth whole exome and low-depth whole genome
npj Genomic Medicine (2024)