Abstract
The cohesin complex has an essential role in maintaining genome organization. However, its role in gene regulation remains largely unresolved. Here we report that the cohesin release factor WAPL creates a pool of free cohesin, in a process known as cohesin turnover, which reloads it to cell-type-specific binding sites. Paradoxically, stabilization of cohesin binding, following WAPL ablation, results in depletion of cohesin from these cell-type-specific regions, loss of gene expression and differentiation. Chromosome conformation capture experiments show that cohesin turnover is important for maintaining promoter–enhancer loops. Binding of cohesin to cell-type-specific sites is dependent on the pioneer transcription factors OCT4 (POU5F1) and SOX2, but not NANOG. We show the importance of cohesin turnover in controlling transcription and propose that a cycle of cohesin loading and off-loading, instead of static cohesin binding, mediates promoter and enhancer interactions critical for gene regulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw and processed sequencing data generated in this study are available from the Gene Expression Omnibus under accession GSE135180. Source data are provided with this paper.
Code availability
Custom R code associated with this manuscript can be found at https://github.com/deWitLab/Liu_2020_NatureGenetics.
References
Rao, S. S. P., Huntley, M. H., Durand, N. C. & Stamenova, E. K. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
de Wit, E. et al. CTCF binding polarity determines chromatin looping. Mol. Cell 60, 676–684 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, 201518552 (2015).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Hyle, J. et al. Acute depletion of CTCF directly affects MYC regulation through loss of enhancer–promoter looping. Nucleic Acids Res. 47, 6699–6713 (2019).
Hadjur, S. et al. Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature 460, 410–413 (2009).
Paliou, C. et al. Preformed chromatin topology assists transcriptional robustness of Shh during limb development. Proc. Natl Acad. Sci. USA 116, 12390–12399 (2019).
Chan, K.-L. et al. Cohesin’s DNA exit gate is distinct from its entrance gate and is regulated by acetylation. Cell 150, 961–974 (2012).
Huis in’t Veld, P. J. et al. Characterization of a DNA exit gate in the human cohesin ring. Science 346, 968–972 (2014).
Kueng, S. et al. Wapl controls the dynamic association of cohesin with chromatin. Cell 127, 955–967 (2006).
Nishiyama, T. et al. Sororin mediates sister chromatid cohesion by antagonizing Wapl. Cell 143, 737–749 (2010).
Misulovin, Z., Pherson, M., Gause, M. & Dorsett, D. Brca2, Pds5 and Wapl differentially control cohesin chromosome association and function. PLoS Genet. 14, e1007225 (2018).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Nitzsche, A. et al. RAD21 cooperates with pluripotency transcription factors in the maintenance of embryonic stem cell identity. PLoS ONE 6, e19470 (2011).
Faure, A. J. et al. Cohesin regulates tissue-specific expression by stabilizing highly occupied cis-regulatory modules. Genome Res. 22, 2163–2175 (2012).
Kojic, A. et al. Distinct roles of cohesin-SA1 and cohesin-SA2 in 3D chromosome organization. Nat. Struct. Mol. Biol. 25, 496–504 (2018).
Cuadrado, A. et al. Specific contributions of cohesin-SA1 and cohesin-SA2 to TADs and Polycomb domains in embryonic stem cells. Cell Rep. 27, 3500–3510 (2019).
Tedeschi, A. et al. Wapl is an essential regulator of chromatin structure and chromosome segregation. Nature 501, 564–568 (2013).
Haarhuis, J. H. I. et al. WAPL-mediated removal of cohesin protects against segregation errors and aneuploidy. Curr. Biol. 23, 2071–2077 (2013).
Aladjem, M. I. et al. ES cells do not activate p53-dependent stress responses and undergo p53-independent apoptosis in response to DNA damage. Curr. Biol. 8, 145–155 (1998).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Bult, C. J. et al. The Mouse Genome Database: enhancements and updates. Nucleic Acids Res. 38, D586–D592 (2010).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
de Jonge, H. J. M. et al. Evidence based selection of housekeeping genes. PLoS ONE 2, e898 (2007).
Schwalb, B. et al. TT-seq maps the human transient transcriptome. Science 352, 1225–1228 (2016).
Gregersen, L. H., Mitter, R. & Svejstrup, J. Q. Using TTchem-seq for profiling nascent transcription and measuring transcript elongation. Nat. Protoc. 15, 604–627 (2020).
van de Werken, H. J. G. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
Rhodes, J. D. P. et al. Cohesin disrupts Polycomb-dependent chromosome interactions in embryonic stem cells. Cell Rep. 30, 820–835 (2020).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Beagan, J. A. et al. YY1 and CTCF orchestrate a 3D chromatin looping switch during early neural lineage commitment. Genome Res. 27, 1139–1152 (2017).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome–nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Garel, S., Marín, F., Grosschedl, R. & Charnay, P. Ebf1 controls early cell differentiation in the embryonic striatum. Development 126, 5285–5294 (1999).
Driller, K. et al. Nuclear factor I X deficiency causes brain malformation and severe skeletal defects. Mol. Cell. Biol. 27, 3855–3867 (2007).
Cuartero, S. et al. Control of inducible gene expression links cohesin to hematopoietic progenitor self-renewal and differentiation. Nat. Immunol. 19, 932–941 (2018).
Lavagnolli, T. et al. Initiation and maintenance of pluripotency gene expression in the absence of cohesin. Genes Dev. 29, 23–38 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Bulger, M. & Groudine, M. Functional and mechanistic diversity of distal transcription enhancers. Cell 144, 327–339 (2011).
Gurumurthy, A., Shen, Y., Gunn, E. M. & Bungert, J. Phase separation and transcription regulation: are super-enhancers and locus control regions primary sites of transcription complex assembly? BioEssays 41, 1800164 (2019).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
van den Berg, D. L. C. et al. An Oct4-centered protein interaction network in embryonic stem cells. Cell Stem Cell 6, 369–381 (2010).
King, H. W. & Klose, R. J. The pioneer factor OCT4 requires the chromatin remodeller BRG1 to support gene regulatory element function in mouse embryonic stem cells. eLife 6, e22631 (2017).
Lopez-Serra, L., Kelly, G., Patel, H., Stewart, A. & Uhlmann, F. The Scc2–Scc4 complex acts in sister chromatid cohesion and transcriptional regulation by maintaining nucleosome-free regions. Nat. Genet. 46, 1147–1151 (2014).
Yan, J. et al. Transcription factor binding in human cells occurs in dense clusters formed around cohesin anchor sites. Cell 154, 801–813 (2013).
Krantz, I. D. et al. Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nat. Genet. 36, 631–635 (2004).
van Heeringen, S. J. & Veenstra, G. J. C. GimmeMotifs: a de novo motif prediction pipeline for ChIP-sequencing experiments. Bioinformatics 27, 270–271 (2011).
Fedr, R. et al. Automatic cell cloning assay for determining the clonogenic capacity of cancer and cancer stem-like cells. Cytometry A 83, 472–482 (2013).
Liu, N. Q. et al. The non-coding variant rs1800734 enhances DCLK3 expression through long-range interaction and promotes colorectal cancer progression. Nat. Commun. 8, 14418 (2017).
Geeven, G., Teunissen, H., de Laat, W. & de Wit, E. peakC: a flexible, non-parametric peak calling package for 4C and Capture-C data. Nucleic Acids Res. 46, e91 (2018).
Orlando, D. A. et al. Quantitative ChIP–Seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Liu, T. Use model-based analysis of ChIP–Seq (MACS) to analyze short reads generated by sequencing protein–DNA interactions in embryonic stem cells. Methods Mol. Biol. 1150, 81–95 (2014).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Li, H. & Wren, J. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30, 2843–2851 (2014).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Bolstad, B. M., Irizarry, R., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Gel, B. et al. regioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics 32, 289–291 (2016).
Acknowledgements
We thank the NKI Genomics Core Facility for help with sequencing, the NKI Bioimaging Facility for help with microscopy and the NKI Flow Cytometry Facility for help with single-cell sorting of genome-edited cells. We thank M. Kanemaki of the National Institute of Genetics in Japan for his suggestions on AID tagging and sharing antibody to OsTir1. We thank B. Nabet and N. Gray of the Dana Farber Cancer Institute for providing the dTAG-13 molecule. We thank R. Young of the Whitehead Institute for Biomedical Research for providing the OCT4–dTAG mESC line. Work in the de Wit laboratory is supported by ERC StG 637587 (‘HAP-PHEN’) and a Vidi grant from the Netherlands Scientific Organization (NWO, 016.16.316). N.Q.L. is supported by a Veni grant from the Netherlands Scientific Organization (NWO, 016.Veni.181.014). N.Q.L., M.M., T.v.d.B., L.B., H.T., M.M.G.A.S. and E.d.W. are part of Oncode, which is partly financed by the Dutch Cancer Society.
Author information
Authors and Affiliations
Contributions
N.Q.L. and E.d.W. conceived and designed the study. N.Q.L., M.M., L.B. and H.T. performed experiments in the laboratory of E.d.W. E.P.N. engineered OsTir1 and RAD21–AID cell lines in the laboratory of B.G.B. N.Q.L., T.v.d.B., M.M.G.A.S. and E.d.W. analyzed data. E.d.W. supervised the study. N.Q.L. and E.d.W. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generating a WAPL-AID degron line.
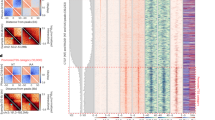
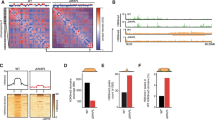
a, Homologous recombination strategy for tagging the endogenous Wapl gene with AID-eGFP. Note that the targeting construct contains a NeoR/KanR resistance gene, but that this has not been used for selecting the first clone. Middle panel shows PCR validation (primers highlighted in left panel) showing homozygous integration of the donor vector (n=1). Right panel shows Sanger sequencing results confirming in-frame tagging of AID tag. b, Left panel shows live cell imaging of WAPL in untreated (0h) and treated (24h) cells (representative images from 3 separate fields). DNA is visualized with Hoechst. Right panel show immunostaining of cohesin subunit RAD21 (representative images from 100 independent cells). DNA is labeled with DAPI. c, WAPL-AID-GFP levels were measured by FACS (gated on all cells besides debris). High temporal resolution shows rapid depletion of WAPL-AID-GFP upon IAA treatment. Median GFP intensity is quantified in bottom plots. Signals are the average of three experiments. Dots indicate mean values. Error bars indicate standard deviation. d, ChIP-seq of WAPL and CTCF before and after IAA treatment. Heatmaps show the distribution of signal for the peaks called in the untreated cells. Average signal is shown above the heatmap. ChIP-seq data is calibrated using spiked-in human HEK-293T cells. Middle two heatmaps show WAPL ChIP-seq signal derived from the human cells.
Extended Data Fig. 2 Effects of WAPL depletion on cell cycle and pluripotency of ES cells.
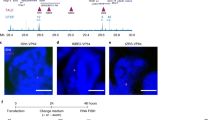
a, FACS plots showing EdU incorporation to measure the cycle phase of individual cells. All single cells are gated, and G1, S and G2 cells are then separated using the blue, orange and red gates, respectively. b, Quantification of cycle phases based on DAPI/EdU FACS analysis for various times after WAPL depletion (n=3 independent experiments). Height of bar indicates mean value. Error bars show standard deviation. Cyan dots show the value of individual measurements. c, An illustration of cell proliferation assay following WAPL depletion. d, Cell cycles measured following WAPL depletion (n=3 independent experiments). Height of bar indicates mean value. Error bars show standard deviation. Blue dots show the value of individual measurements. e, Western blot showing the protein levels of the pluripotency transcription factors SOX2, OCT4 and NANOG (cropped, n=1). f, Alkaline phosphatase (AP) staining is measured as a marker for pluripotency on the untreated (0 h), treated (96 h) and 48 h treatment followed by 48 h wash-off WAPL-AID and parental cell line (representative images from 2 independent clones). g, An illustration of single cell colony formation experiment following WAPL depletion. h, Single cell colony formation measured following WAPL depletion (n=3 independent experiments). Height of bar indicates mean value. Error bars show standard deviation. Blue dots show the value of individual measurements.
Extended Data Fig. 3 Effects of WAPL depletion on global gene expression.
a, A volcano plot shows differential genes between the untreated parental (PT) and WAPL-AID cells at a false discovery rate (FDR) cutoff of 0.05. b, Transcriptomic changes of PT cells with 96 h IAA treatment following using the same FDR cutoff as (a). c, Transcriptomic changes at different time points following WAPL depletion using the same FDR cutoff as (a, b). d, Heatmap showing the genes that are differentially expressed following IAA treatment in WAPL-AID cells. Genes are clustered according to the timepoint in which they are first observed to be differentially expressed (left panel). e, Gene Set Enrichment Analysis of RNA-seq data following WAPL depletion. Analysis was performed for Gene Ontology databases and the significant (FDR < 0.01) terms with a positive normalized enrichment score (NES) are plotted. Terms that are related to development, differentiation or morphogenesis are highlighted.
Extended Data Fig. 4 ChIP-seq heatmaps of two active histone modifications.
Left panel shows signal of the active promoter mark H3K4me3 aligned to the promoters of differentially expressed genes. Right panel shows H3K27ac alignment to enhancers within 10kb up- or downstream of the promoter of a differentially expressed gene.
Extended Data Fig. 5 Validation and functional annotation of the Regions of Dynamic Cohesin.
a, Venn diagram showing the uniquely bound and overlapping RAD21 binding sites identified by ChIP-seq in a second WAPL-AID clone. Peak calling was performed on a subset of 7 M sequencing reads for both of the samples. b, Scatter plot showing the Pearson correlation of RAD21 ChIP-seq data between the two different WAPL-AID clones. Red rectangle highlighting the correlation of the same time points. c, Alignment of RAD21 binding data from a second WAPL-AID clone on the RDCs and RSCs identified in the original WAPL-AID clone. d, Alignment of published ChIP-seq data for five different cohesin subunits generated in V6.5 mESC line (see Supplementary Table 3 for details). e, Example locus showing a region with an RDC and RSC. RAD21 binding is shown for treated and untreated cell for two WAPL-AID clones. Cohesin binding is shown for the five subunits shown in (d). f, Table containing the top 5 (by lowest FDR) categories in a GREAT analysis using the Mouse Genome Informatics (MGI) expression database for RDCs and RSCs.
Extended Data Fig. 6 Molecular features of the Regions of Dynamic Cohesin and the Regions of Stalled Cohesin.
a, Alignments of H3K27ac and MED1 ChIP-seq data on RDCs and RSCs. b, Example region showing the relationship between the factors analyzed in Fig. 1g and the position of a RSC. c, similar alignments as (a) but for SOX2, OCT4 and NANOG ChIP-seq data. d, RAD21 alignment on the transcription start sites (TSS) of the top 1000 most stably expressed housekeeping genes for treated and untreated cells in two different clones.
Extended Data Fig. 7 Global effects of WAPL depletion on genome architecture and nascent transcription.
a, Relative contact probability plot (RCP) showing the distribution of the chance of two loci contacting in the context of the 3D genome as a function of the distance between them on the linear chromosome. b, Example locus showing the formation of extended loops upon WAPL depletion. Heatmaps visualize the contact frequency matrices for the Nfe2l3 locus. Rightmost panel shows the differential contacts between the 24h treated cells and the untreated cells. c, Total number of TADs detected in control (0 h) and WAPL depleted (24 h) cells. d, TAD size from control (0 h) and WAPL depleted (24 h) depleted cells (Box plot elements: center line, median; box, inter quartile range/IQR; whiskers, 1.5 times the IQR from the median value; dots, instances lower or higher than the IQR from the median value). e, Distribution of intra- and inter-TAD interactions. f, Global effects of WAPL depletion on nascent transcription at early time points (6 and 24 h). g, Nascent transcription changes following WAPL depletion are plotted using the same gene clusters described in Extended Data Fig. 3d.
Extended Data Fig. 8 Depletion of WAPL and RAD21 reorganizes the 3D genome and affects gene expression.
a, Effects of WAPL depletion on gene expression similar to Fig. 2d but using RNA-seq. The yellow stars indicate the observed over expected enrichment of the RNA-seq data (n=2 independent experiments), and the violin plots show distribution of enrichment based on 10,000 random circular permutations (red lines: median values, orange lines: interquartile range, *: P<0.001, **: P<0.0001). b, High-resolution 4C-seq data for the Klf4 locus (viewpoint: Klf4 enhancer, overlapping with the distal RDC of the Klf4 gene). The 4C-seq data is visualized in the same way as Fig. 2e. The bottom two rows show nascent transcription of the Klf4 gene and neighboring region in the untreated (0 h) and treated (24 h) ES cells. c, Expression of the Klf4 gene was quantified by RT-qPCR (n=3 independent experiments) after 24 h of WAPL depletion and 24 h depletion followed by 24/48 h wash-off (two-sided student’s t-test without multiple testing correction, bar height: mean values, error bars: standard deviations, red dots: individual data points). d, High-resolution 4C-seq analysis of the Klf4 locus in the RAD21 depletion experiment (viewpoint: the same as (b)). The 4C-seq data is visualized in the same way as Fig. 2e. The bottom two rows show expression of the Klf4 gene and neighboring region in the untreated (0 h) and treated (24 h) ES cells. e, Global effects of RAD21 depletion on gene expression at early time points (6 and 24 h).
Extended Data Fig. 9 The role of transcription factors in the recruitment of cohesin.
a–c, heatmaps showing the binding of pluripotency transcription factor SOX2 (a), OCT4 (b) and NANOG (c) remain unchanged following WAPL depletion. d, A published OCT4-FKBP-mCherry cell lines was used. The FKBP degron can be degraded with the dTAG-13 molecule. e, A western blot shows protein levels of OCT4 after 6 and 24 hours of dTAG-13 treatment (cropped). f, g, ChIP-seq heatmaps showing MED1 binding at OCT4 binding sites and NANOG binding sites in the OCT4-FKBP (f) and NANOG-FKBP (g) lines, respectively. h, i, ChIP-seq heatmaps showing RAD21 binding and chromatin accessibility at CTCF binding sites in OCT4-FKBP (h), SOX2-FKBP (i, left) and NANOG-FKBP (i right). For OCT4-FKBP CTCF sites from V6.5 mESCs were used for alignment and for Nanog-FKBP, which are derived from E14 mESCs CTCF sites identified in WAPL-AID cells were used.
Extended Data Fig. 10 Further characterization of RAD21 binding in WAPL-AID NPCs.
a, Staining for Nestin (Green) and GFAP (red) in NPCs derived from the WAPL-AID cell line (representative images from 2 experiments and 3 separate fields per experiment). b, Staining for alkaline phosphatase in NPCs derived from the WAPL-AID cell line (n=1). c, Comparison of the differential genes between ESC and NPC identified in a published study and in the WAPL-AID line. d, Venn diagram shows overlap of RAD21 binding sites measured ChIP-seq for untreated and 24h treated WAPL-AID NPCs. Only sites that showed consistent unique or shared binding across two replicates were included. Peak calling was performed on a subset of 7 M sequencing reads for all the samples. e, MA-plot showing the log2 fold-change as a function of the number of reads in a RAD21 peak. Highlighted dots indicate significantly changed RAD21 levels based on DESeq2 with an FDR <0.01 and a fold-change of at least 2 up or down. Red is increase in RAD21, blue is decrease. f, Annotation of the motifs identified in Fig. 5d.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3
Supplementary Tables
Supplementary Tables 1–3
Source data
Source Data Fig. 1
Unprocessed western blots for Fig. 1.
Source Data Fig. 3
Unprocessed western blots for Fig. 3.
Source Data Fig. 4
Unprocessed western blots for Fig. 4.
Source Data Extended Data Fig. 2
Unprocessed western blots for Extended Data Fig. 2.
Source Data Extended Data Fig. 9
Unprocessed western blots for Extended Data Fig. 9.
Rights and permissions
About this article
Cite this article
Liu, N.Q., Maresca, M., van den Brand, T. et al. WAPL maintains a cohesin loading cycle to preserve cell-type-specific distal gene regulation. Nat Genet 53, 100–109 (2021). https://doi.org/10.1038/s41588-020-00744-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00744-4
This article is cited by
-
BRWD1 orchestrates small pre-B cell chromatin topology by converting static to dynamic cohesin
Nature Immunology (2024)
-
RAD21 is the core subunit of the cohesin complex involved in directing genome organization
Genome Biology (2023)
-
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Nature Communications (2023)
-
SARS-CoV-2 restructures host chromatin architecture
Nature Microbiology (2023)
-
Enhancer–promoter contact formation requires RNAPII and antagonizes loop extrusion
Nature Genetics (2023)